Proteomics I Mass Spectrometry Functional Genomics by Mass

Proteomics I Mass Spectrometry Functional Genomics by Mass Spectrometry optional (Andersen and Mann, 2000) FEBS Letters 480, 25 -31

Proteomics II Yeast Two Hybrid Toward a Protein-Protein Map of the Budding Yeast optional (Ito et al. , 2000) PNAS 97(3), 1143 -1147

Mass Spectrometry • Molecules to be analyzed, referred to as analytes are first ionized (usually in a vacuum), • Newly charged molecules are introduced into an electric and/or magnetic field in gas phase, • Their path through the field is a function of the mass to charge ratio m/z, • m/z of the ionized species can be used to deduce the mass of the analyte with high precision.

Biological Samples. . bringing polypeptides and nucleic acids to the gas phase usually degrades the molecules, 1988 matrix assisted laser desorption/ionization mass spectrometry MALDI-MS electrospray ionization mass spectrometry ESI-MS 2002 Nobel Prize on Chemistry, Fenn & Tanaka

Electrospray Ionization Mass Spectrometry ESI-MS • Peptides analytes, in solution, are passed through a charged needle that is kept at high electrical potential, – – the peptides are ionized, this disperses the solution into a fine spray, the solvent quickly evaporates, peptides now in gas phase, • Enter mass spectrometer for mass fingerprinting, or Peptide Sequencing.
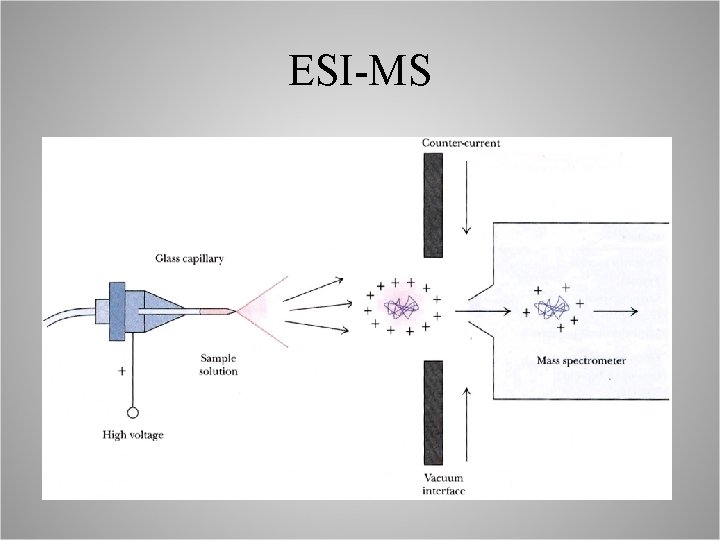
ESI-MS

Multi-protein Complexes “mud pit” analysis ? . . . i. e. nuclear pore complexes, . . . i. e. cellulose synthase complexes, . . . i. e. spindle pole apparati, . . . i. e. proteins involved in the spliceosome, …etc.

From the abstract… This is the first time that the components of a multi-protein complex from an organism with sequenced genome have been characterized by mass spectrometry. The technique should be applicable to any protein complex that can be biochemically purified from an organism whose genome is known.
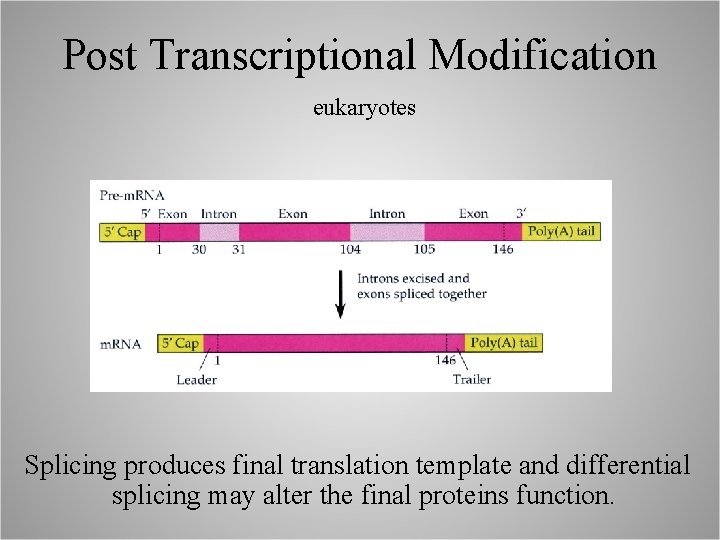
Post Transcriptional Modification eukaryotes Splicing produces final translation template and differential splicing may alter the final proteins function.
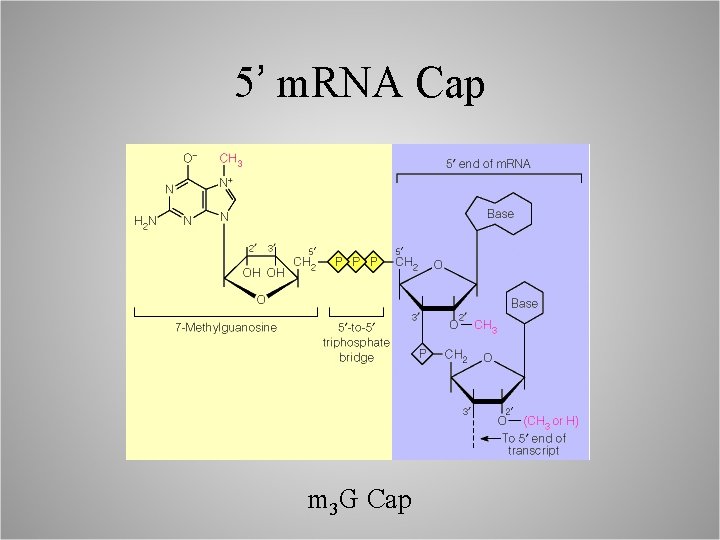
5’ m. RNA Cap m 3 G Cap
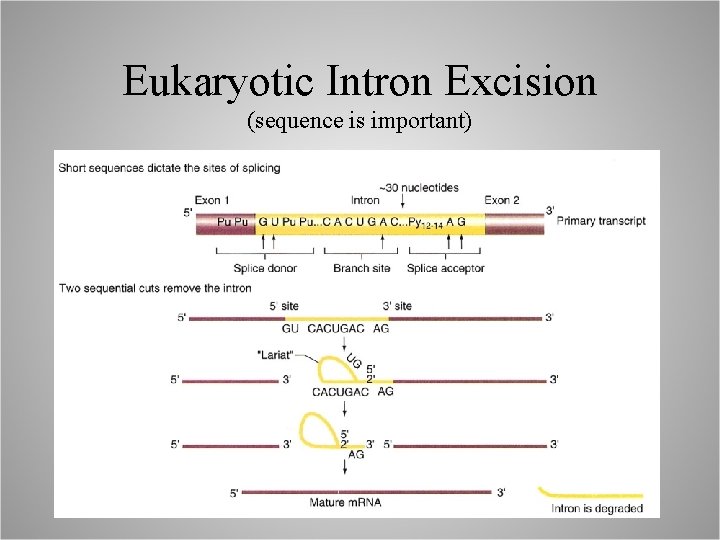
Eukaryotic Intron Excision (sequence is important)
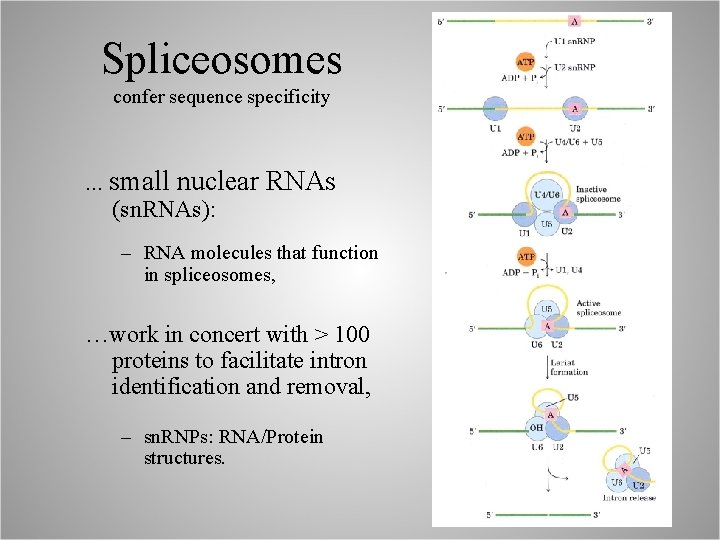
Spliceosomes confer sequence specificity . . . small nuclear RNAs (sn. RNAs): – RNA molecules that function in spliceosomes, …work in concert with > 100 proteins to facilitate intron identification and removal, – sn. RNPs: RNA/Protein structures.
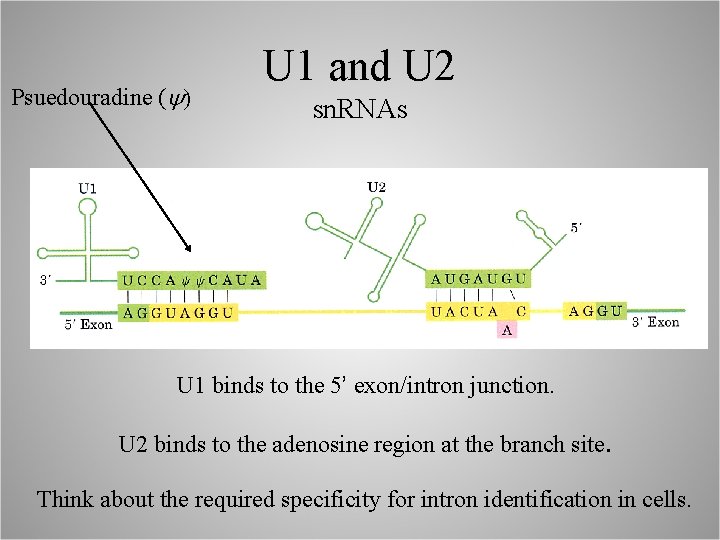
Psuedouradine (y) U 1 and U 2 sn. RNAs U 1 binds to the 5’ exon/intron junction. U 2 binds to the adenosine region at the branch site. Think about the required specificity for intron identification in cells.
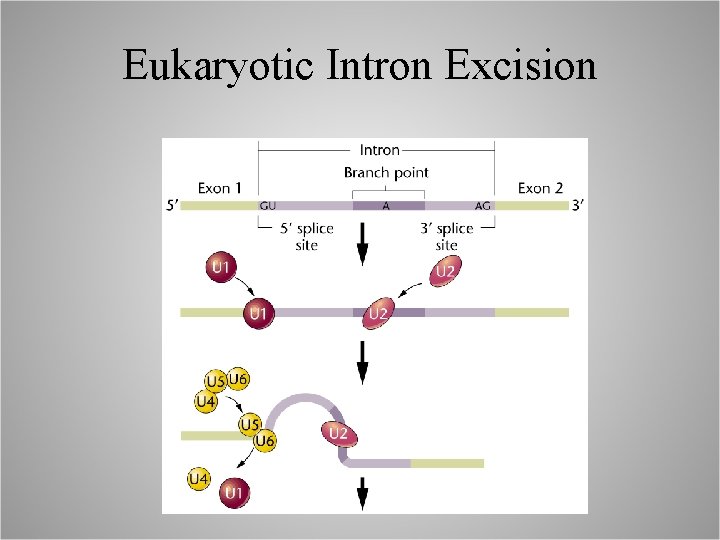
Eukaryotic Intron Excision
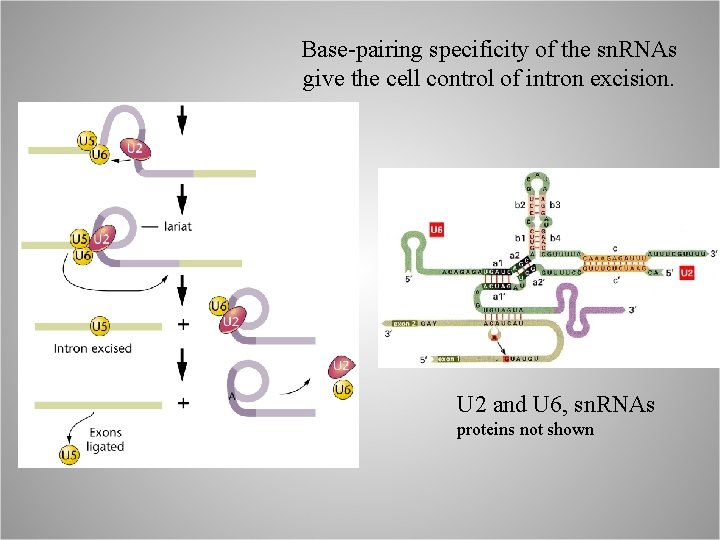
Base-pairing specificity of the sn. RNAs give the cell control of intron excision. U 2 and U 6, sn. RNAs proteins not shown
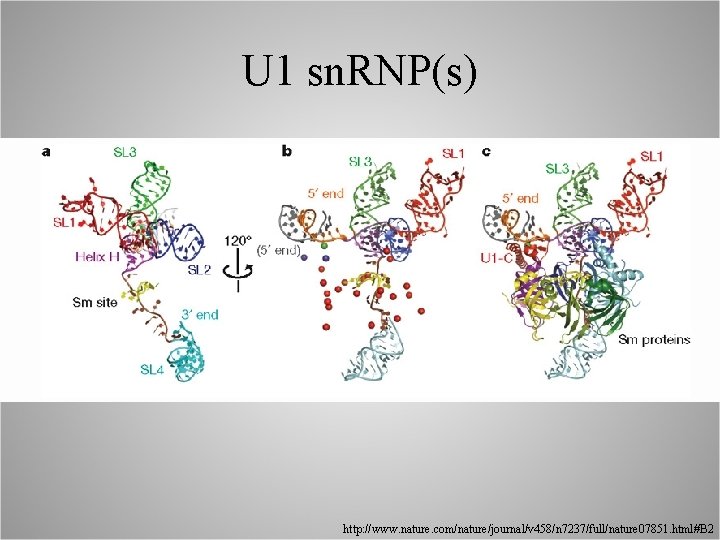
U 1 sn. RNP(s) http: //www. nature. com/nature/journal/v 458/n 7237/full/nature 07851. html#B 2

Isolate Spliceosome Complex Anti-m 3 G Antibody Chromotography Histidine Tag Snp 1 Nickel Nitrilotriacetic Acid (Ni-NTA) Affinity Chromotography . . . RNA processing includes the capping of the 5’ end by 7 methylguanosine triphosphate, . . . Snp 1 has been identified as a U 1 specific sn. RNP, …antibodies are available for this molecule, . . . it was modified by a hexahistidine addition to the N-terminus, …all molecules with a capped RNA can be coprecititated. . the functional Snp 1, in the experiment (yeast), is carried on a plasmid, . . . the exposed his-tag binds to nickel on a chromatography column, is washed off with imidazole.
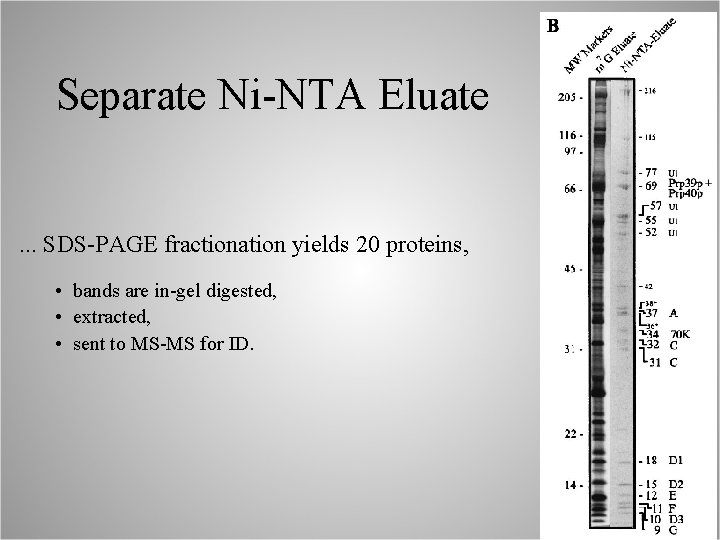
Separate Ni-NTA Eluate . . . SDS-PAGE fractionation yields 20 proteins, • bands are in-gel digested, • extracted, • sent to MS-MS for ID.
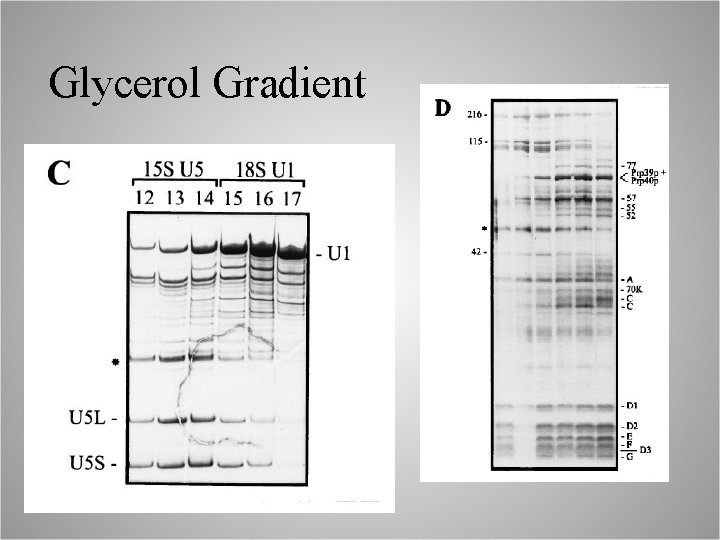
Glycerol Gradient
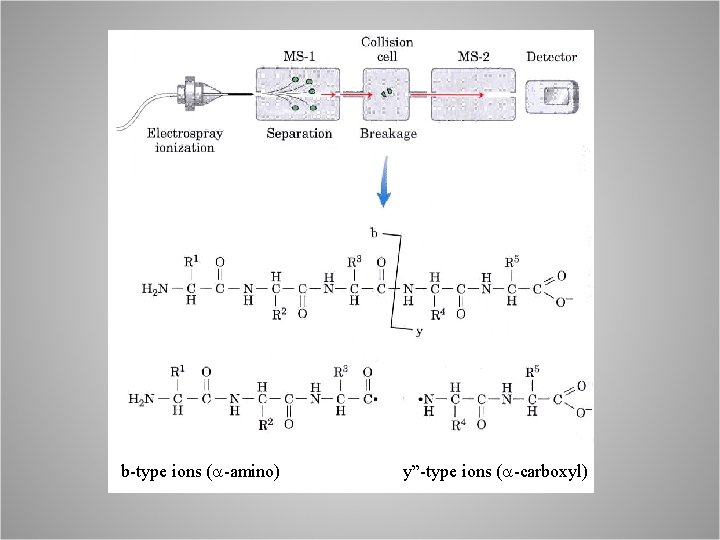
b-type ions (a-amino) y”-type ions (a-carboxyl)
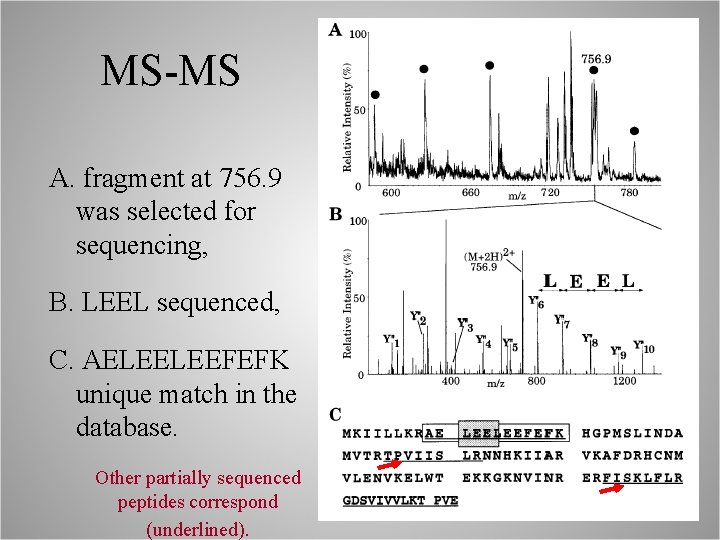
MS-MS A. fragment at 756. 9 was selected for sequencing, B. LEEL sequenced, C. AELEELEEFEFK unique match in the database. Other partially sequenced peptides correspond (underlined).
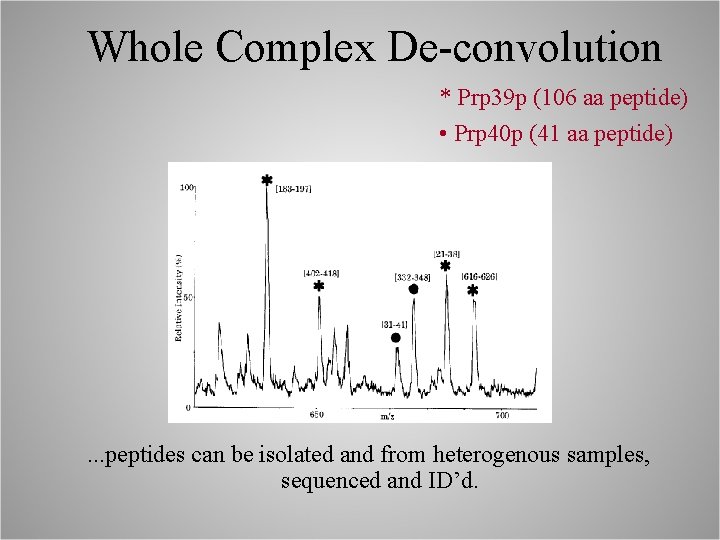
Whole Complex De-convolution * Prp 39 p (106 aa peptide) • Prp 40 p (41 aa peptide) . . . peptides can be isolated and from heterogenous samples, sequenced and ID’d.

Found Players • All of the known U 1 -specific proteins found to date were identified, • An ortholog to human U 1 -C was found in yeast, that had not been previously identified, • Four novel proteins were also identified.

Establishment of Principle. . . so, in one fell swoop, years of molecular genetic and biochemical research was replicated, and. . . a U 1 -C protein, not previously detected, was identified as a component in the system, and. . . four novel proteins have been ID’d, – not necessarily components, – but excellent research leads.

Use of Principle. . . in one subsequent experiment with isolated human spliceosome, – 70 spliceosome protein spots were analyzed, – 19 novel splicing factors were identified, and. . . all 19 could be cloned directly via EST libraries, and. . . in vivo conformation of the role of these factors in splicing was obtained through co-localization studies using green fluorescence protein (GFP).
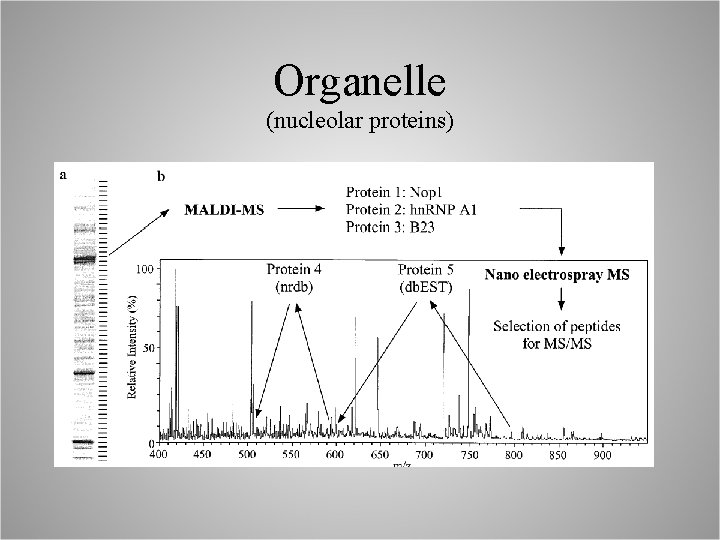
Organelle (nucleolar proteins)

Proteomics II Spliceosome Analysis Yeast Two-Hybrid Nature Genetics, 1997, 16 pp 277 - 282
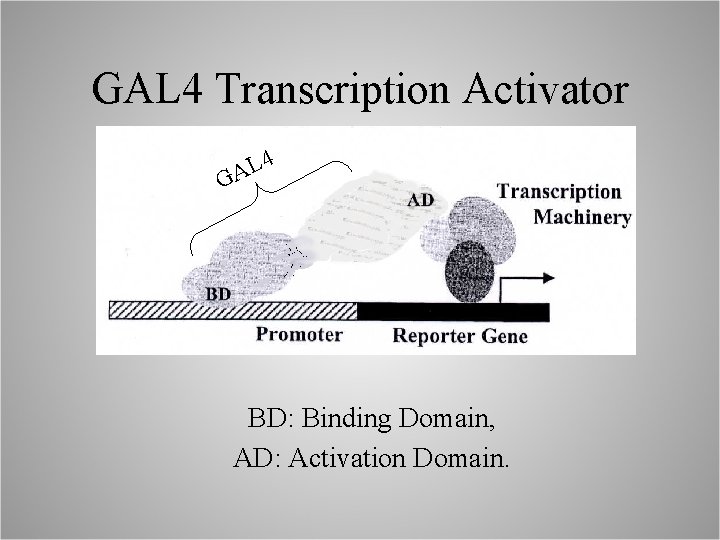
GAL 4 Transcription Activator 4 L A G BD: Binding Domain, AD: Activation Domain.
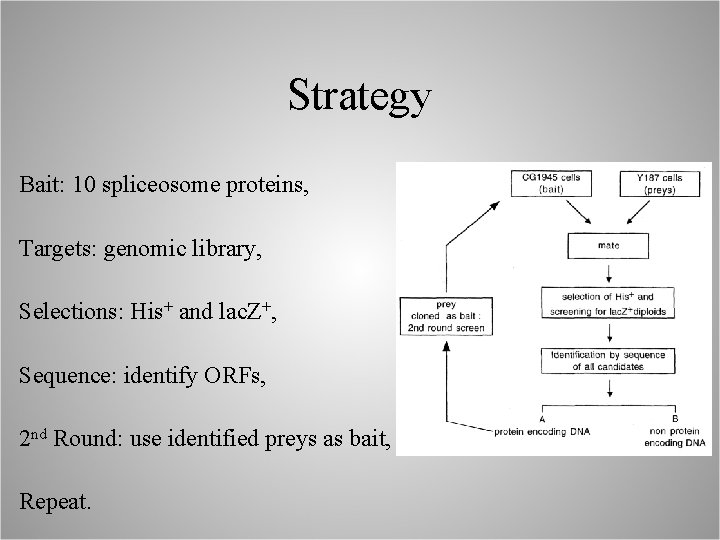
Strategy Bait: 10 spliceosome proteins, Targets: genomic library, Selections: His+ and lac. Z+, Sequence: identify ORFs, 2 nd Round: use identified preys as bait, Repeat.
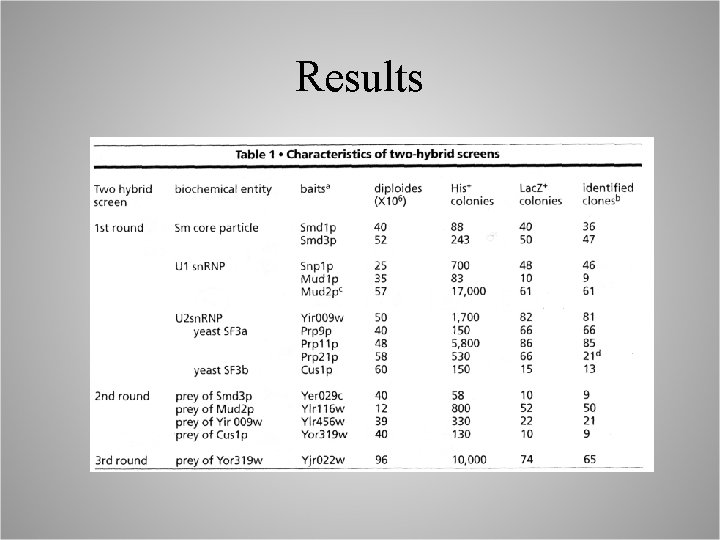
Results


Monday • Group 1, • Gut Metagenomics.
- Slides: 32