Proteomics and posttranslational modifications proteinligand interactions protein complexes









- Slides: 25

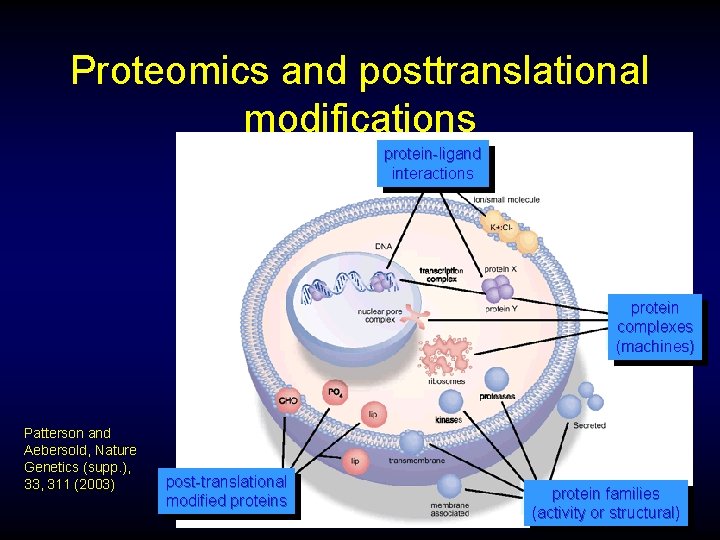
Proteomics and posttranslational modifications protein-ligand interactions protein complexes (machines) Patterson and Aebersold, Nature Genetics (supp. ), 33, 311 (2003) post-translational modified proteins protein families (activity or structural)

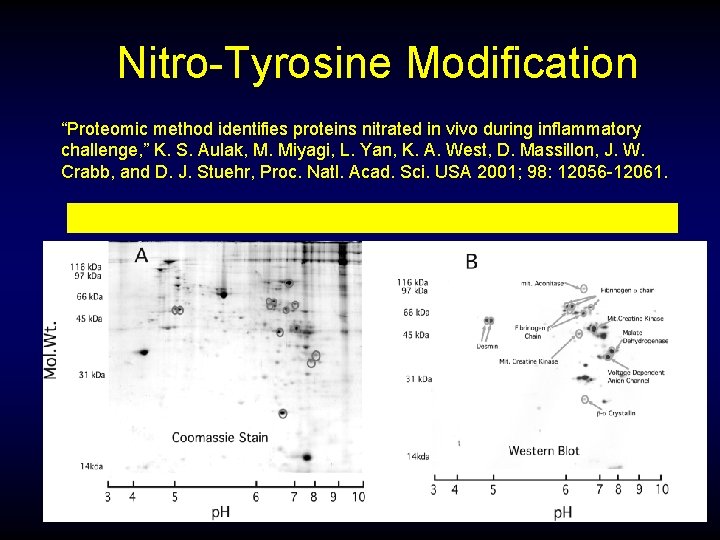
Nitro-Tyrosine Modification “Proteomic method identifies proteins nitrated in vivo during inflammatory challenge, ” K. S. Aulak, M. Miyagi, L. Yan, K. A. West, D. Massillon, J. W. Crabb, and D. J. Stuehr, Proc. Natl. Acad. Sci. USA 2001; 98: 12056 -12061. Anti-nitrotyrosine immunopositive proteins in lung of rats induced with LPS.

Phosphoprotein Stain Phospho Peppermint. Stick phosphoprotein molecular weight standards separated on a 13% SDS polyacrylamide gel. The markers contain (from largest to smallest) beta-galactosidase, bovine serum albumin (BSA), ovalbumin, betacasein, avidin and lysozyme. Ovalbumin and beta-casein are phosphorylated. The gel was stained with Pro-Q Diamond phosphoprotein gel stain (blue) followed by SYPRO Ruby protein gel stain (red). The digital images were pseudocolored.

Phosphoprotein Stain Visualization of total protein and phosphoproteins in a 2 -D gel Proteins from a Jurkat T-cell lymphoma line cell lysate were separated by 2 -D gel electrophoresis and stained with Pro-Q Diamond phosphoprotein gel stain (blue) blue followed by SYPRO Ruby protein gel stain (red). red After each dye staining, the gel was imaged and the resulting composite image was digitally pseudocolored and overlaid. T. H. Steinberg et al. , Global quantitative phosphoprotein analysis using Multiplexed Proteomics technology, Proteomics 2003, 3, 1128 -1144

MS/MS and Phosphorylation • Detection of phosphopeptides in complex mixtures can be facilitated by neutral loss and precurson ion scanning using tandem mass spectrometers • Allow selective visualization of peptides containing phosphorylated residues • Most commonly performed with triple quadrupole mass spectrometers precursor ion transmission collision cell (chamber) mass analysis of product ions

MS/MS and Phosphorylation • Precursor ion scan – Q 1 is set to allow all the components of the mixture to enter the collision cell and undergo CAD – Q 3 is fixed at a specific mass value, so that only analytes which fragment to give a fragment ion of this specific mass will be detected – Phospho-peptide fragments by CAD to give an ion at m/z 79 (PO 3) – Set Q 3 to m/z 79: only species which fragment to give a fragment ion of 79 reach the detector and hence indicating phosphorylation detector Q 1 Q 2 collision cell Q 3

MS/MS and Phosphorylation • Neutral loss scan – Q 1 and Q 3 are scanned synchronously but with a specific m/z offset – The entire mixture is allowed to enter the collision cell, but only those species which fragment to yield a fragment with the same mass as the offset will be observed at the detector – p. Ser and p. Thr peptides readily lose phosphoric acid during CAD (98 Da) – For 2+ ion set offset at 49 • Any species which loses 49 from a doubly charged ion would be observed at the detector and be indicative of phosphorylation

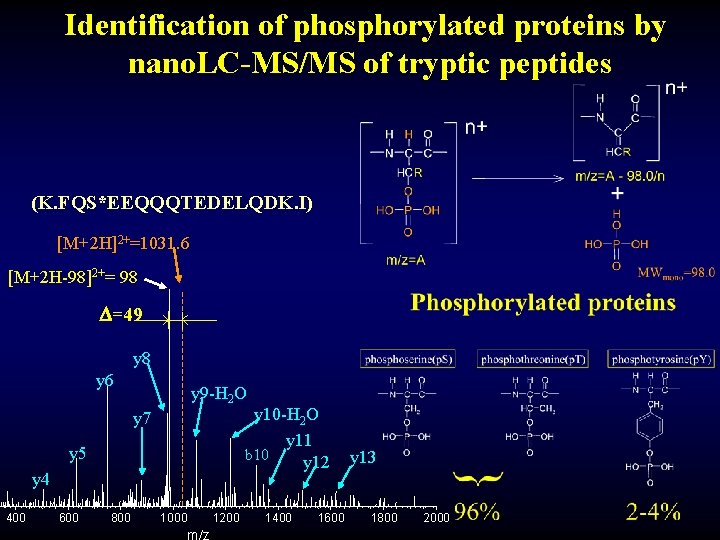
Identification of phosphorylated proteins by nano. LC-MS/MS of tryptic peptides (K. FQS*EEQQQTEDELQDK. I) [M+2 H]2+=1031. 6 [M+2 H-98]2+= 98 =49 y 8 y 6 y 9 -H 2 O y 10 -H 2 O y 11 b 10 y 12 y 7 y 5 y 4 400 600 800 1000 1200 1400 1600 y 13 1800 2000

Enrichment strategies to analyze phosphoproteins/peptides • Phospecific antibodies – Anti-p. Y quite successful – Anti-p. S and anti-p. T not as successful, but may be used (M. Grønborg, T. Z. Kristiansen, A. Stensballe, J. S. Andersen, O. Ohara, M. Mann, O. N. Jensen, and A. Pandey, “Approach for Identification of Serine/Threoninephosphorylated Proteins by Enrichment with Phospecific Antibodies. ” Mol. Cell. Proteomics 2002, 1: 517– 527. • Immobilized metal affinity chromatography (IMAC) – Negatively charged phosphate groups bind to postively charged metal ions (e. g. , Fe 3+, Ga 3+) immobilized to a chromatographic support – Limitation: non-specific binding to acidic side chains

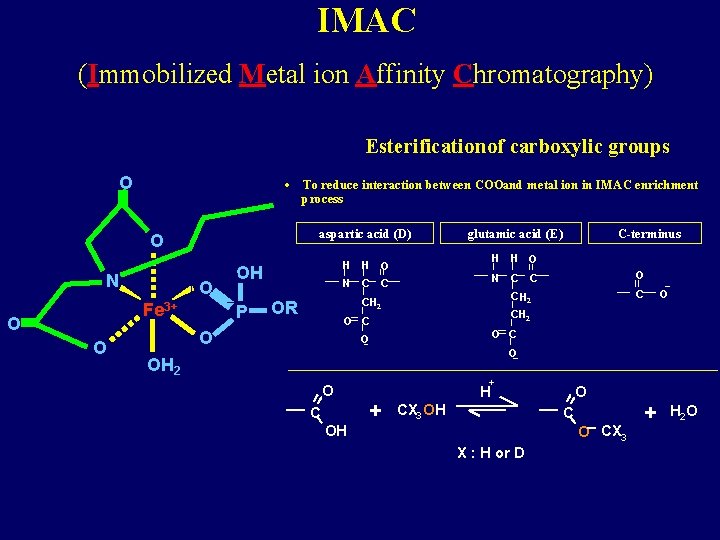
IMAC (Immobilized Metal ion Affinity Chromatography) Esterification of carboxylic groups O • aspartic acid (D) O N O Fe 3+ O O - and metal ion in IMAC enrichment To reduce interaction between COO process OH P H H O N C C glutamic acid (E) H H O N C C O CH 2 C CH 2 OR O CH 2 O C-terminus O C O O OH 2 O C OH + + O H CX 3 OH C O CX 3 X : H or D + H 2 O

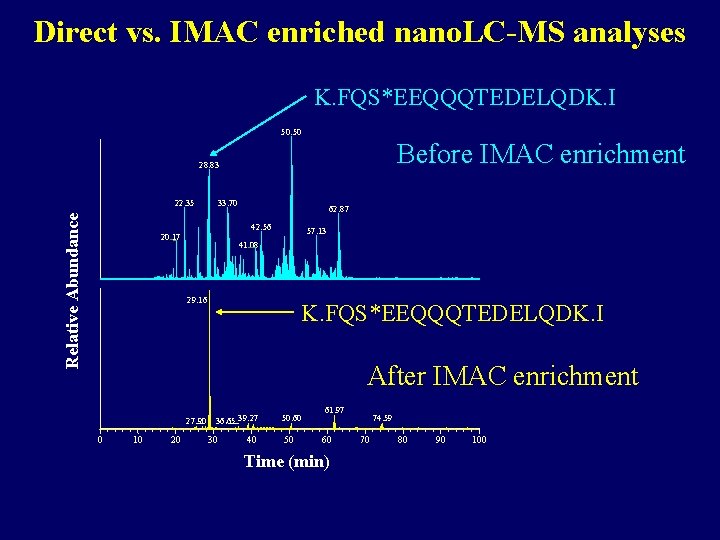
Direct vs. IMAC enriched nano. LC-MS analyses K. FQS*EEQQQTEDELQDK. I 50. 50 Before IMAC enrichment 28. 83 Relative Abundance 22. 35 33. 70 62. 87 42. 56 20. 17 57. 13 41. 08 29. 16 K. FQS*EEQQQTEDELQDK. I After IMAC enrichment 27. 90 0 10 20 36. 65 39. 27 30 40 50. 60 50 61. 97 60 Time (min) 74. 59 70 80 90 100

Enrichment strategies to analyze phosphoproteins/peptides • Chemical derivatization – Introduce affinity tag to enrich for phosphorylated molecules • e. g. , biotin binding to immobilized avidin/streptavidin

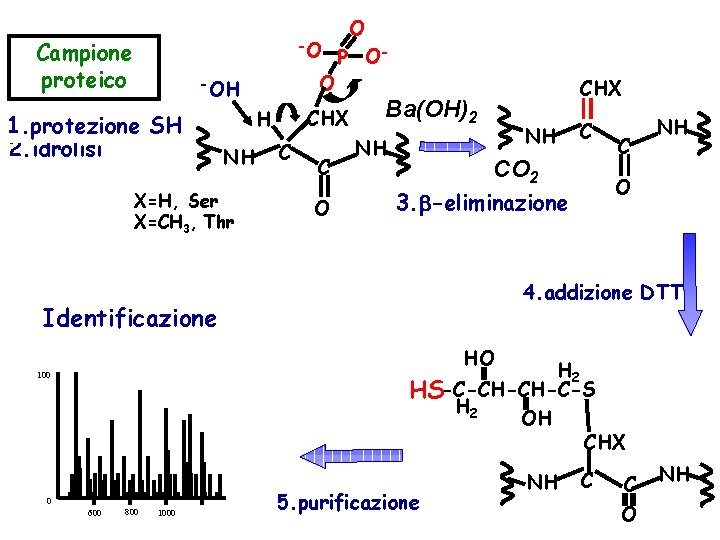
Enrichment strategies to analyze phosphoproteins/peptides • • • Oda et al. , Nature Biotech. 2001, 19, 379 for analysis of p. S and p. T Remove Cys-reactivity by oxidation with performic acid Base hydrolysis induce ß-elimination of phosphate from p. S/p. T Addition of ethanedithiol allows coupling to biotin Avidin affinity chromatography to purify phosphoproteins

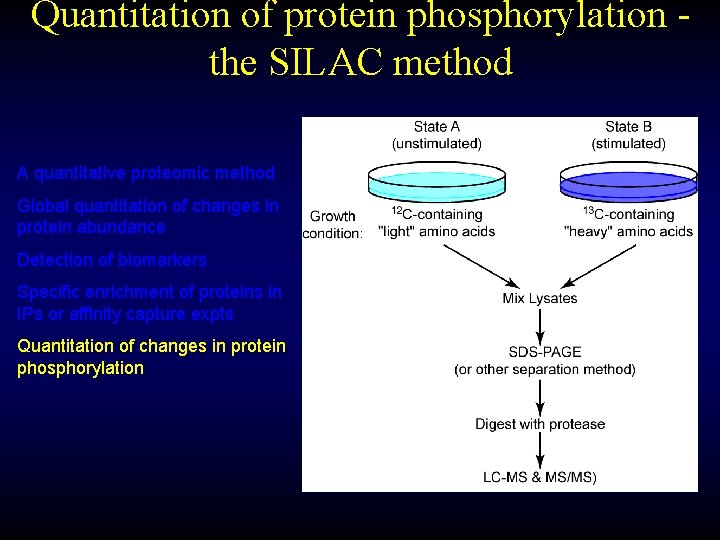
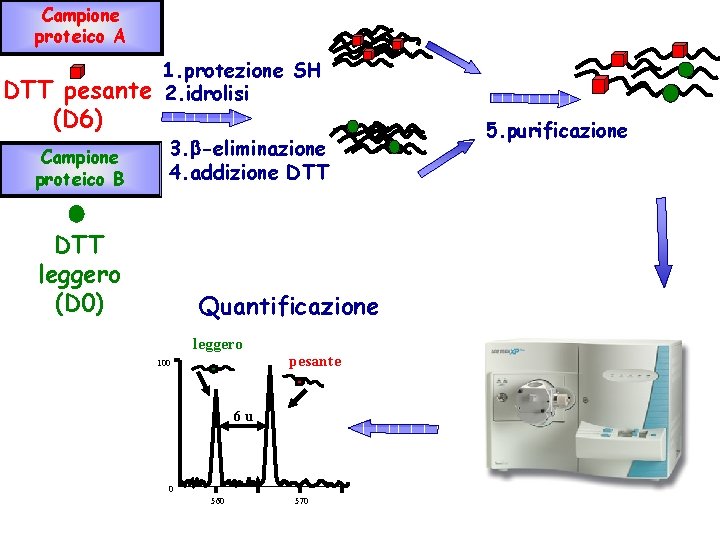
Quantitation of protein phosphorylation the SILAC method A quantitative proteomic method Global quantitation of changes in protein abundance Detection of biomarkers Specific enrichment of proteins in IPs or affinity capture expts Quantitation of changes in protein phosphorylation

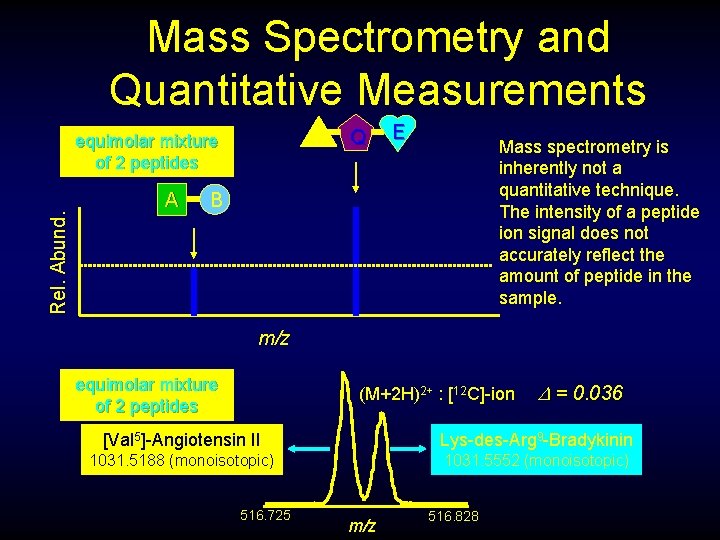
Mass Spectrometry and Quantitative Measurements H Rel. Abund. equimolar mixture of 2 peptides A Q E Mass spectrometry is inherently not a quantitative technique. The intensity of a peptide ion signal does not accurately reflect the amount of peptide in the sample. B m/z equimolar mixture of 2 peptides (M+2 H)2+ : [12 C]-ion = 0. 036 [Val 5]-Angiotensin II Lys-des-Arg 9 -Bradykinin 1031. 5188 (monoisotopic) 1031. 5552 (monoisotopic) 516. 725 m/z 516. 828

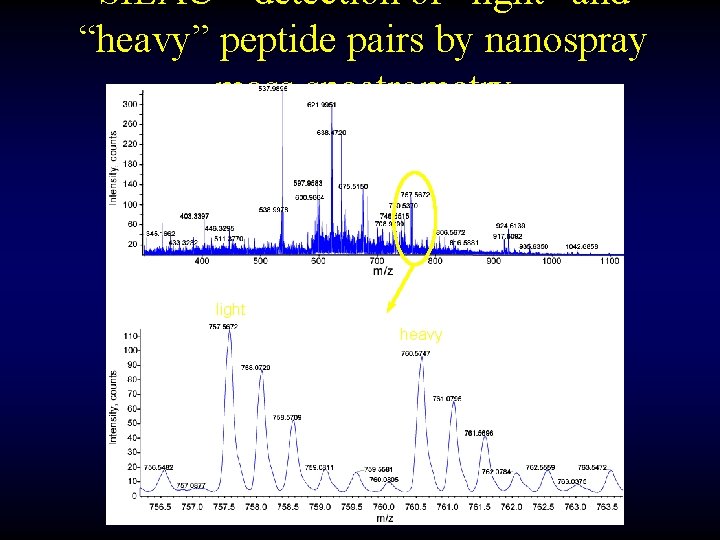
SILAC – detection of “light” and “heavy” peptide pairs by nanospray mass spectrometry light heavy

Campione proteico O -O P OO -OH 1. protezione SH 2. idrolisi CHX H NH X=H, Ser X=CH 3, Thr C C O CHX Ba(OH)2 NH NH C CO 2 C NH O 3. -eliminazione 4. addizione DTT Identificazione HO H 2 HS-C-CH-CH-C-S H 2 OH CHX 100 0 600 800 1000 5. purificazione NH C C O NH

Campione proteico A DTT pesante (D 6) Campione proteico B 1. protezione SH 2. idrolisi 3. -eliminazione 4. addizione DTT leggero (D 0) Quantificazione leggero 100 pesante 6 u 0 560 570 5. purificazione




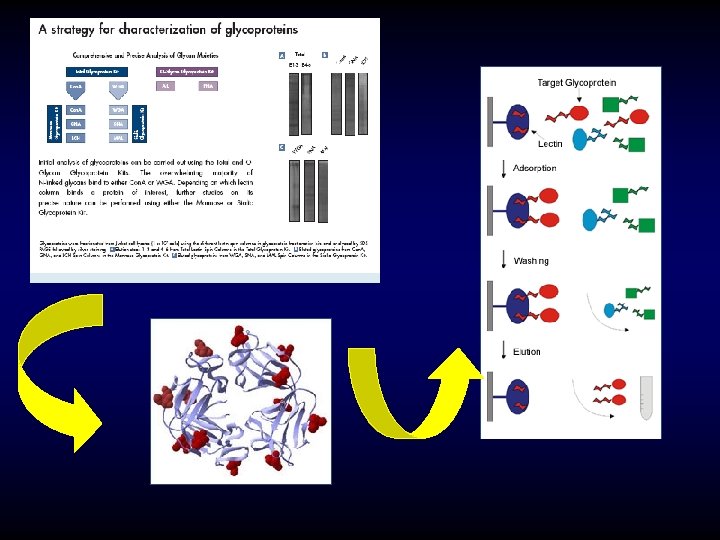
Glycoprotein Gel Stain Detection of glycoproteins and total protein on an SDS-polyacrylamide gel using the Pro-Q Fuchsia Glycoprotein Gel Stain Kit. Candy. Cane glycoprotein molecular weight standards containing alternating glycosylated and nonglycosylated proteins were electrophoresed through a 13% polyacrylamide gel. After separation, the gel was stained with SYPRO Ruby protein gel stain to detect all eight marker proteins (left). Subsequently, the gel was stained by the standard periodic acid–Schiff base (PAS) method in the Pro-Q Fuchsia Glycoprotein Gel Stain Kit to detect the glycoproteins alpha 2 macroglobulin, glucose oxidase, alpha 1 -glycoprotein and avidin. Pro-Q™ Glycoprotein Stain (DDAO phosphate) Molecular Formula: C 15 H 18 Cl 2 N 3 O 5 P (MW 422. 20)


