Proteomics and mass spectrometry Manimalha Balasubramani Outline Mass

Proteomics and mass spectrometry Manimalha Balasubramani

Outline Mass spectrometers Protein identification Quantitative proteomics Protein-protein interactions
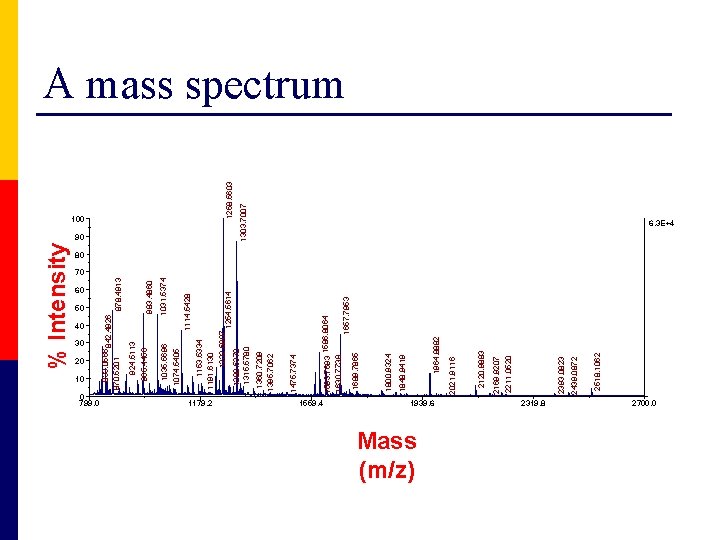
% Intensity 60 50 40 30 20 10 1179. 2 1475. 7374 1254. 5614 1559. 4 1939. 6 Mass (m/z) 2319. 8 2518. 1062 2439. 0872 2393. 0823 2211. 0520 2169. 9207 2120. 9883 2021. 9116 1964. 8882 1848. 9419 1800. 9324 1258. 5603 1303. 7007 90 1360. 7209 1395. 7062 1280. 5370 1315. 5780 1031. 5374 983. 4860 1114. 5428 1232. 5907 1153. 5334 1191. 6130 1035. 5696 1074. 5405 924. 5113 965. 4456 100 1593. 7693 1586. 8064 1630. 7738 1657. 7953 1689. 7865 0 799. 0 833. 0566 842. 4926 870. 5201 878. 4913 A mass spectrum 6. 3 E+4 80 70 2700. 0

Basically measures mass Adapted from google
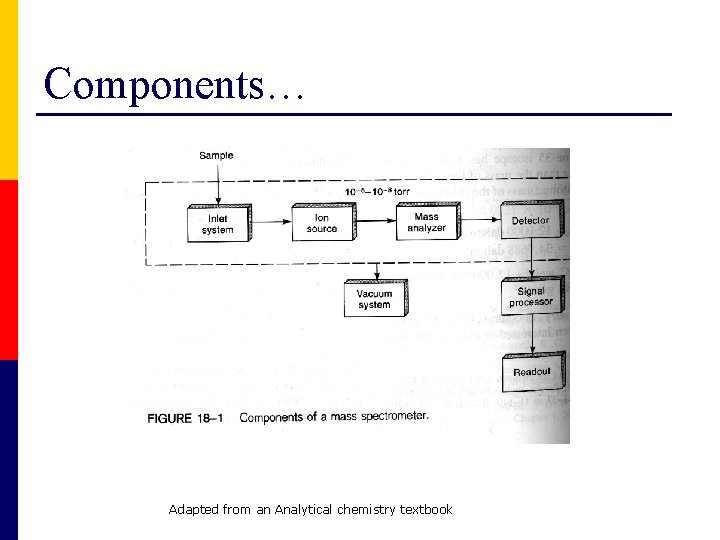
Components… Adapted from an Analytical chemistry textbook

Ionization process MALDI ESI Matrix Assisted Laser Desorption Ionization Electro. Spray Ionization Nobel prize in Chemistry, 2002
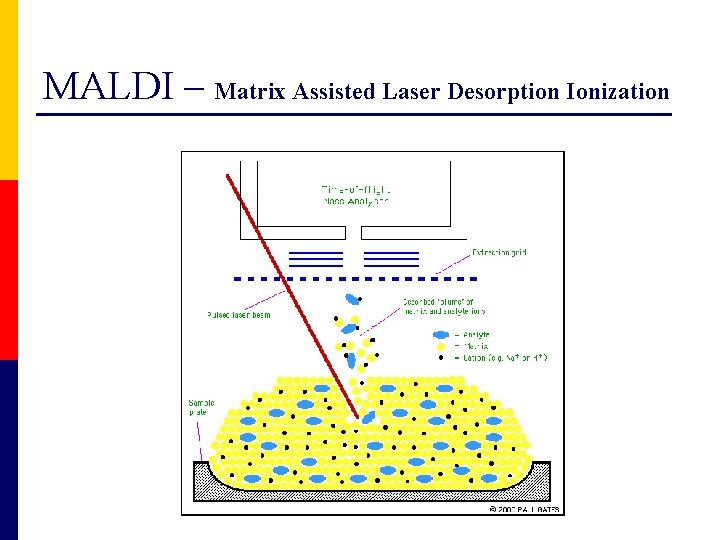
MALDI – Matrix Assisted Laser Desorption Ionization

ESI – Electro Spray Ionization
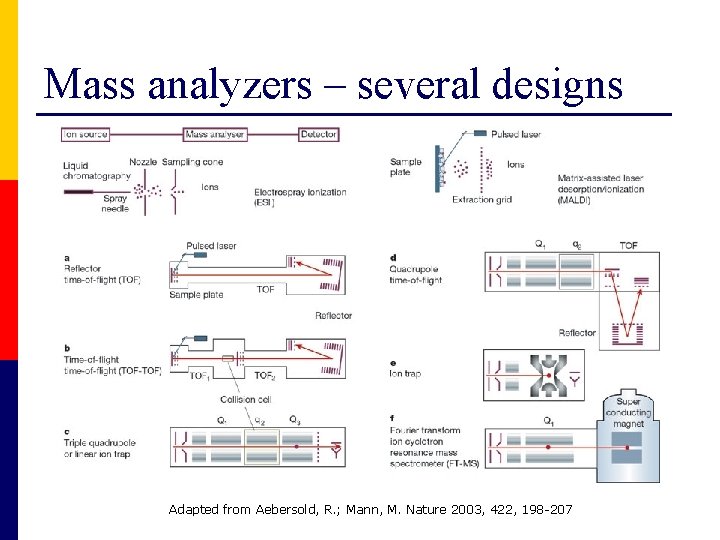
Mass analyzers – several designs Adapted from Aebersold, R. ; Mann, M. Nature 2003, 422, 198 -207

GPCL inventory ABI Voyager DE PRO, walk-up use p ABI 4700 Proteomics Analyzer p Thermoelectron LCQ Deca with Surveyor HPLC p ABI Qstar Elite with Ultimate 3000 HPLC p Bruker micr. OTOF with Ultimate 3000 HPLC p Bruker 12 Tesla FTMS with Ultimate 3000 HPLC p

Time-of-flight (TOF) analyzers MALDI TOF Voyager DE PRO ESI TOF Ultimate 3000 with micr. OTOF
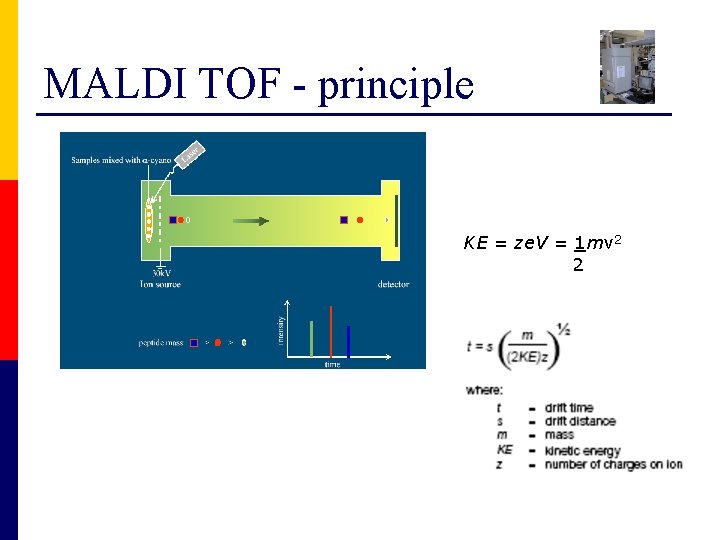
MALDI TOF - principle KE = ze. V = 1 mv 2 2
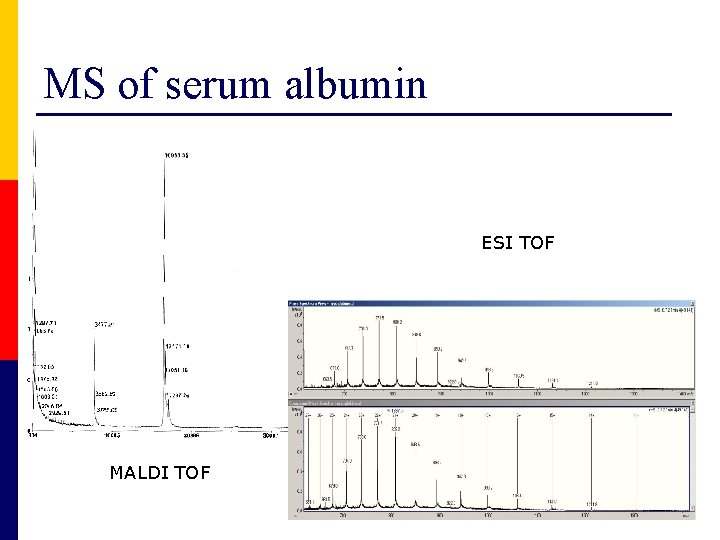
MS of serum albumin ESI TOF MALDI TOF

Tandem mass spectrometer MALDI TOF/TOF MS and MS/MS
Ion Trap MS, MS 2, MS 3, …. MSn

Quadrupole-q-TOF ESI Qq. TOF

…installation phase…. FT MS

…bottom line… . . Resolution and mass accuracy…
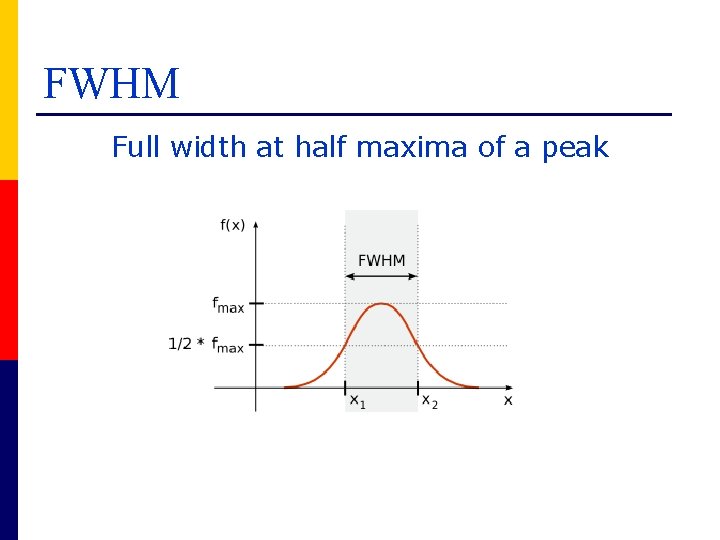
FWHM Full width at half maxima of a peak
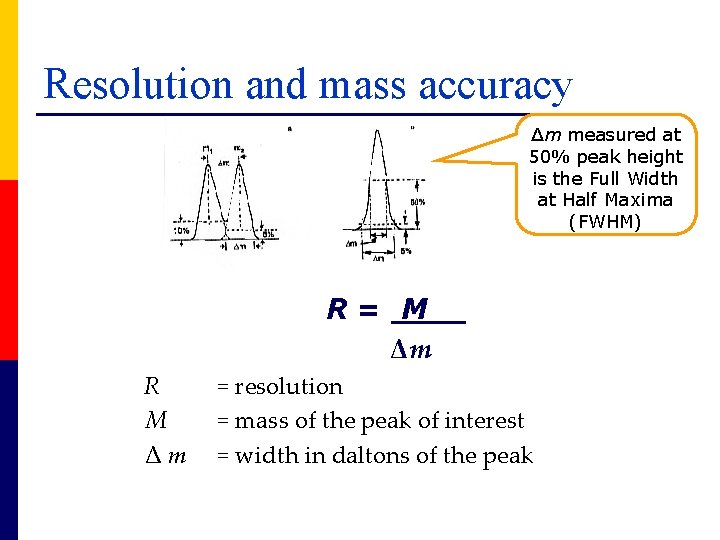
Resolution and mass accuracy Δm measured at 50% peak height is the Full Width at Half Maxima (FWHM) R= M Δm R M Δm = resolution = mass of the peak of interest = width in daltons of the peak
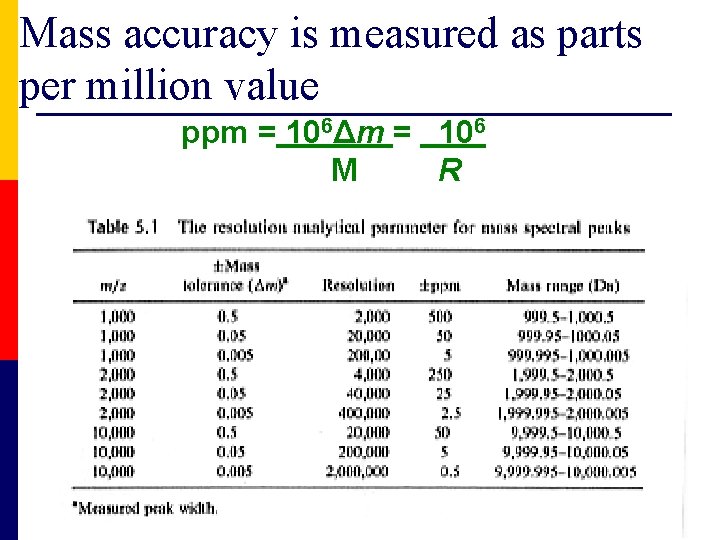
Mass accuracy is measured as parts per million value ppm = 106Δm = 106 M R

outline Mass spectrometers Protein identification Quantitative proteomics Protein-protein interactions
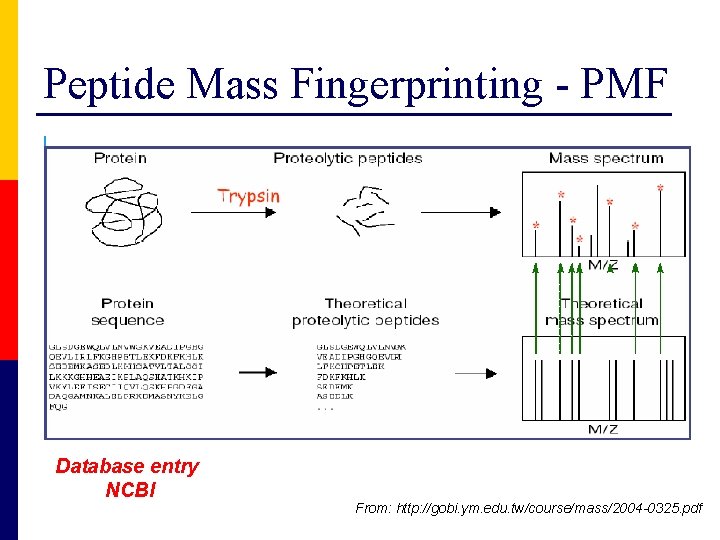
Peptide Mass Fingerprinting - PMF Database entry NCBI From: http: //gobi. ym. edu. tw/course/mass/2004 -0325. pdf

Informatics p Search engines n n p Mascot, Matrix Science Sequest, Thermoelectron Free-ware n n Protein prospector (http: //prospector. ucsf. edu/) TPP tools (http: //tools. proteomecenter. org/TPP. php)

Database searching using MASCOT Overview of the experiment Submission of data to MASCOT webserver
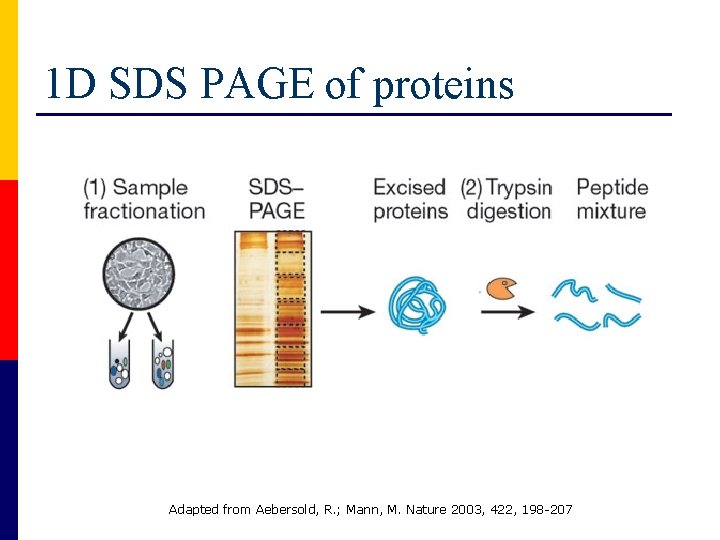
1 D SDS PAGE of proteins Adapted from Aebersold, R. ; Mann, M. Nature 2003, 422, 198 -207
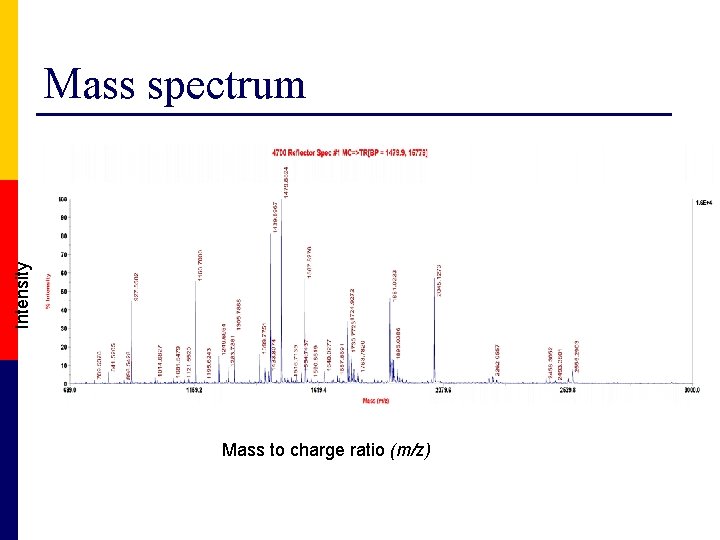
Intensity Mass spectrum Mass to charge ratio (m/z)

Peak list p Compiled from the mass spectra n n p Mass list and intensity Submitted to the search engine
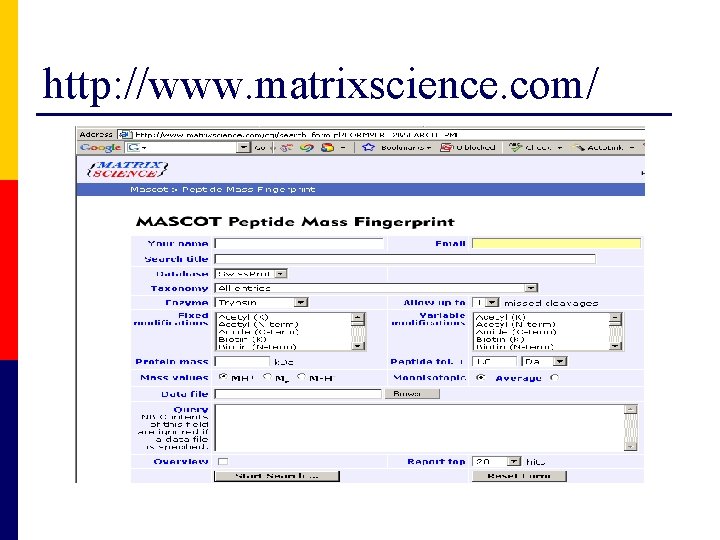
http: //www. matrixscience. com/

Mascot scoring A frequency factor matrix, F, is created, in which each row represents an interval of 100 Da in peptide mass, and each column an interval of 10 k. Da in intact protein mass. As each sequence entry is processed, the appropriate matrix elements fi, j are incremented so as to accumulate statistics on the size distribution of peptide masses as a function of protein mass. The elements of F are then normalised by dividing the elements of each 10 k. Da column by the largest value in that column to give the Mowse factor matrix M: After searching the experimental mass values against a calculated peptide mass database, the score for each entry is calculated according to: Where MProt is the molecular weight of the entry and the product term is calculated from the Mowse factor elements for each match between the experimental data and peptide masses calculated from the entry.

List of common contaminants Trypsin autolysis peptides p Matrix peaks p Keratin from skin, hair p Other contaminants p
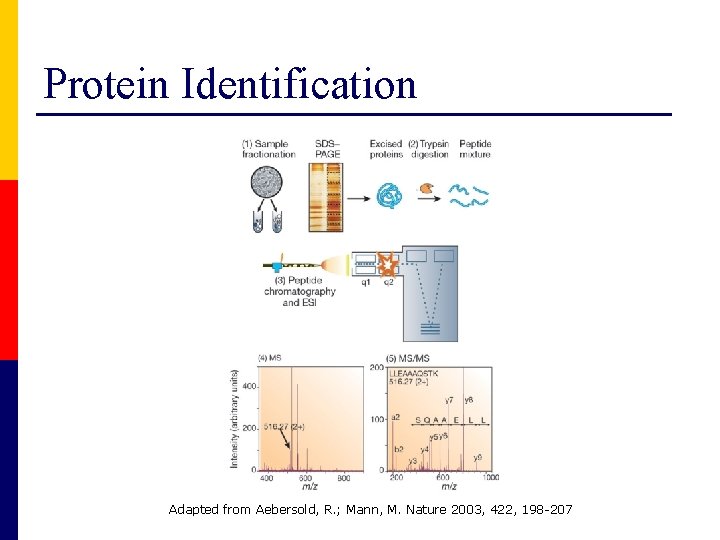
Protein Identification Adapted from Aebersold, R. ; Mann, M. Nature 2003, 422, 198 -207
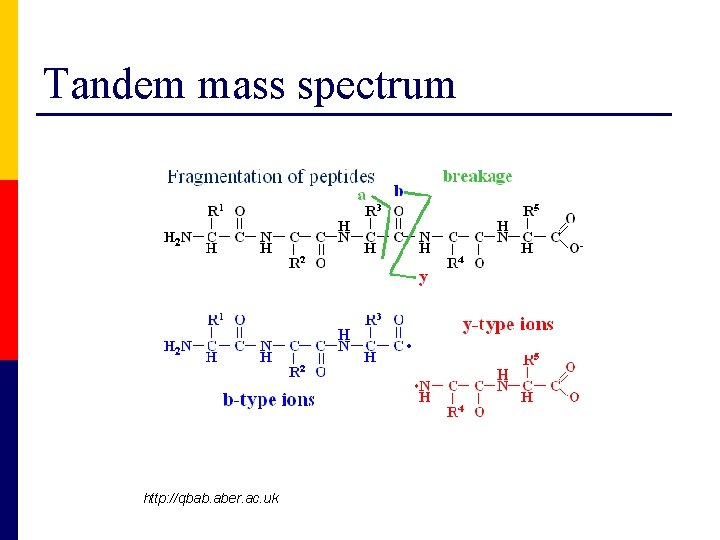
Tandem mass spectrum http: //qbab. aber. ac. uk
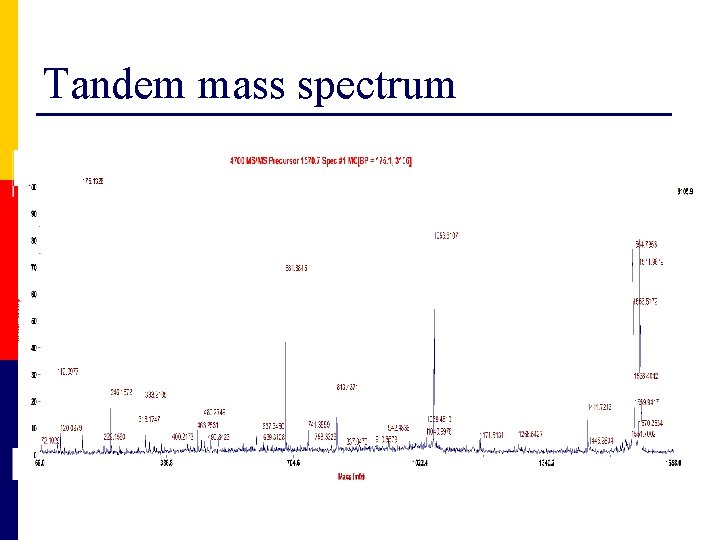
Tandem mass spectrum
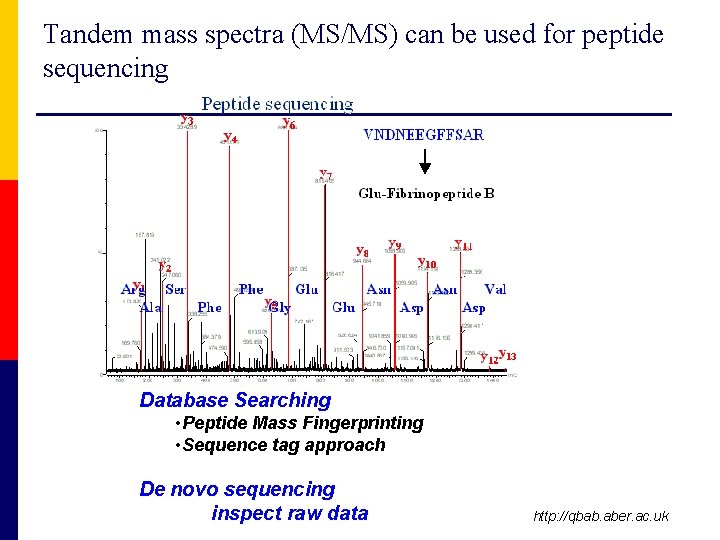
Tandem mass spectra (MS/MS) can be used for peptide sequencing Database Searching • Peptide Mass Fingerprinting • Sequence tag approach De novo sequencing inspect raw data http: //qbab. aber. ac. uk
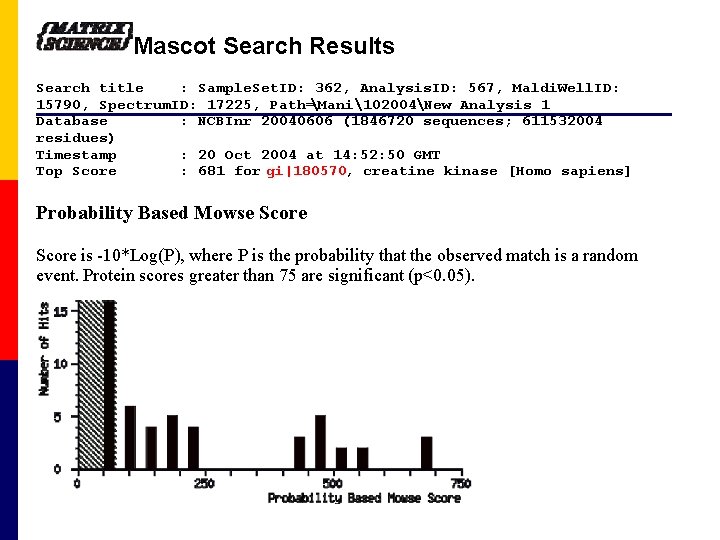
Mascot Search Results Search title : Sample. Set. ID: 362, Analysis. ID: 567, Maldi. Well. ID: 15790, Spectrum. ID: 17225, Path=Mani102004New Analysis 1 Database : NCBInr 20040606 (1846720 sequences; 611532004 residues) Timestamp : 20 Oct 2004 at 14: 52: 50 GMT Top Score : 681 for gi|180570, creatine kinase [Homo sapiens] Probability Based Mowse Score is -10*Log(P), where P is the probability that the observed match is a random event. Protein scores greater than 75 are significant (p<0. 05).

Top hits from Mascot Search – there are multiple accession numbers for the same protein
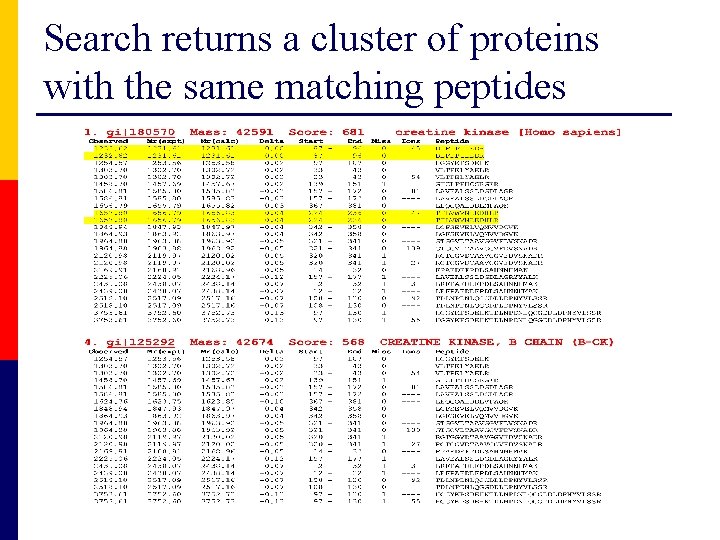
Search returns a cluster of proteins with the same matching peptides

Creatine kinase B is the highest scoring protein Match to: gi|21536286 ; Score: 681 Creatine kinase - B [Homo sapiens] Nominal mass (Mr): 42591; Calculated p. I value: 5. 34 Observed Mass & p. I: 43 kd, 6. 2 -6. 27 Sequence Coverage: 46% 1 MPFSNSHNAL KLRFPAEDEF PDLSAHNNHM AKVLTPELYA ELRAKSTPSG 51 FTLDDVIQTG VDNPGHPYIM TVGCVAGDEE SYEVFKDLFD PIIEDRHGGY 101 KPSDEHKTDL NPDNLQGGDD LDPNYVLSSR VRTGRSIRGF CLPPHCSRGE 151 RRAIEKLAVE ALSSLDGDLA GRYYALKSMT EAEQQQLIDD HFLFDKPVSP 201 LLSASGMARD WPDARGIWHN DNKTFLVWVN EEDHLRVISM QKGGNMKEVF 251 TRFCTGLTQI ETLFKSKDYE FMWNPHLGYI LTCPSNLGTG LRAGVHIKLP 301 NLGKHEKFSE VLKRLRLQKR GTGGVDTAAV GGVFDVSNAD RLGFSEVELV 351 QMVVDGVKLL IEMEQRLEQG QAIDDLMPAQ K

outline Mass spectrometers Protein identification Quantitative proteomics Protein-protein interactions
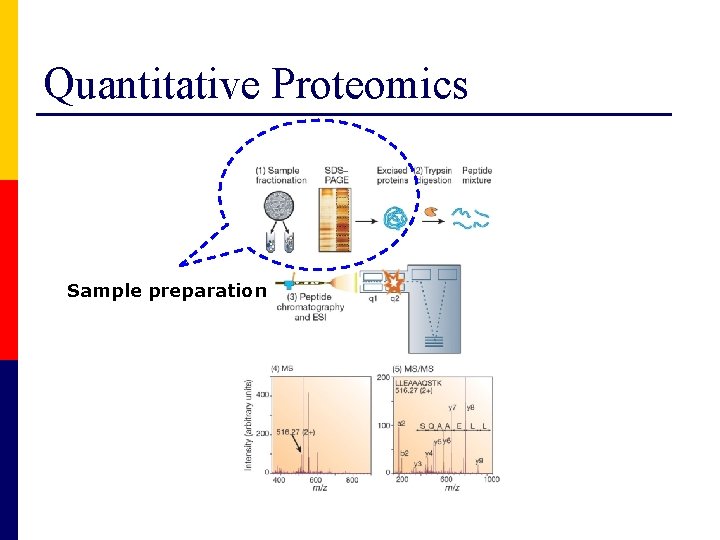
Quantitative Proteomics Sample preparation

From 2 D gels …. to MALDI or ESI MS Control Test Pool Cy 3 Cy 5 Image analysis with Delta 2 D, Decodon Quantitate Export spot list to robotic picker

. . its high-throughput… 1 st Dimension - Isoelectric focussing 2 nd Dimension – SDS PAGE Spot picking Trypsin gel digest
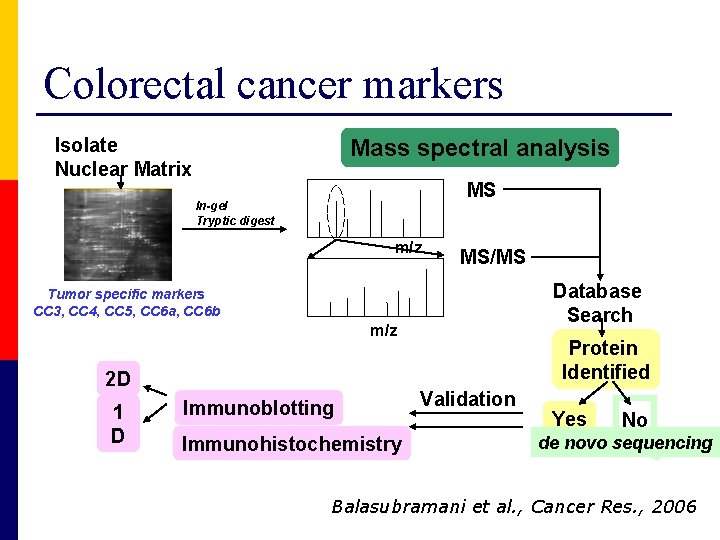
Colorectal cancer markers Isolate Nuclear Matrix Mass spectral analysis MS In-gel Tryptic digest m/z MS/MS Database Search Tumor specific markers CC 3, CC 4, CC 5, CC 6 a, CC 6 b m/z 2 D 1 D Immunoblotting Immunohistochemistry Protein Identified Validation Yes No de novo sequencing Balasubramani et al. , Cancer Res. , 2006
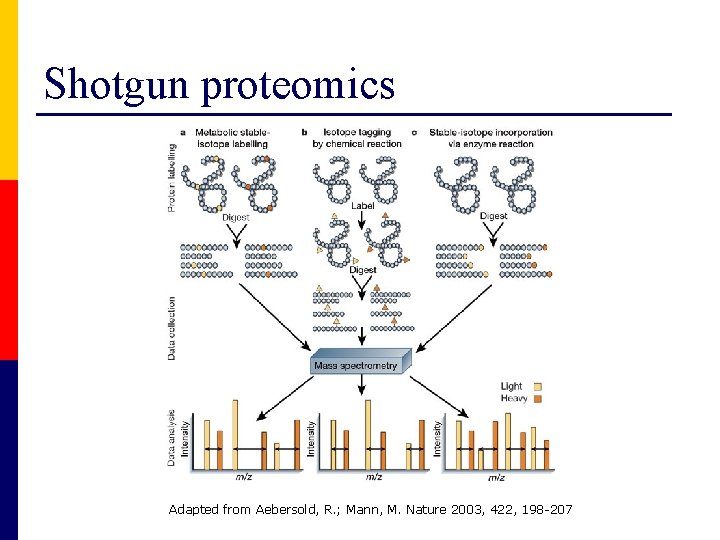
Shotgun proteomics Adapted from Aebersold, R. ; Mann, M. Nature 2003, 422, 198 -207
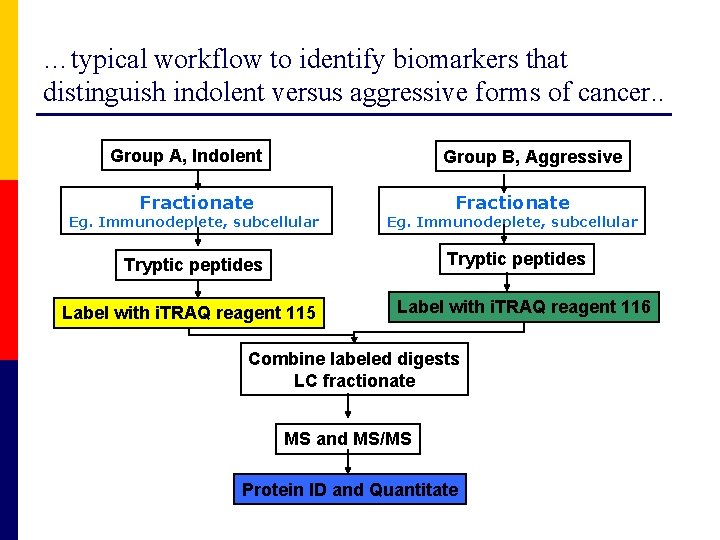
…typical workflow to identify biomarkers that distinguish indolent versus aggressive forms of cancer. . Group A, Indolent Group B, Aggressive Fractionate Eg. Immunodeplete, subcellular Tryptic peptides Label with i. TRAQ reagent 115 Label with i. TRAQ reagent 116 Combine labeled digests LC fractionate MS and MS/MS Protein ID and Quantitate

Sample handling In-solution Isoelectric focussing HPLC 1 D or 2 D LC MALDI

Protein-protein interaction studies Immunoaffinity pull-downs p Tandem affinity purification p

GPCL Billy W Day Paul Wood Mirunalni Thangavelu Tamanna Sultana Emanuel M Schreiber Chris Bolcato Chris Myers Patrick Miller Robert Wolfe

definitions The amu is defined as 1/12 th the mass of one neutral 6 C 12 atom p Amu is also called the dalton p 1 amu =1/12 ( 12 g 12 C/mol 12 C 6. 0221 x 1023 atoms 12 C/mol 12 C 1. 6605 x 10 -24 g/atom 12 C p

Isotopic species of M p (M + H)+ (M + 1 H)/1 H+ p (M + 2 H)2+ (M + 2 H)/2 H+ p (M + 3 H)3+ (M + 3 H)/3 H+
- Slides: 51