ProteinProtein Interaction Interaction in biological systems Occurs when

Protein-Protein Interaction

Interaction in biological systems • Occurs when two or more proteins bind together • Biochemical interaction (Physical interaction of proteins) • Genetic interaction: the phenomenon where the effects of one gene are modified by one or several other genes. In the pathway, there is no biochemical interaction among gene products, but they are affected by one another. • Important for the majority of biological functions • Signals from the exterior of a cell are mediated to the inside of that cell by protein-protein interactions of the signaling molecules

Methods for Proteins-Protein Interaction Important to understand biological processes Multiple methods exist its own strengths and weaknesses in terms of the sensitivity and specificity of the method 1. Yeast two hybrid 2. Immunoprecipiation (pull-down assay) 3. Phage display 4. Tandem Affinity Purification 5. Bimolecular fluorescence complementation (Bi. FC) 6. Fluorescence resonance energy transfer (FRET) 7. Protein microarray

Yeast two hybrid Yeast Two Hybrid (Y 2 H) • A molecular-biological technique is used to detect protein interaction • A transcription factor is split into two separate fragments called the binding domain (BD) and activation domain (AD) one of a protein fragment complementation method • Usually, library is linked to activation domain and the target protein is fused to DNA binding domain of a transcription factor
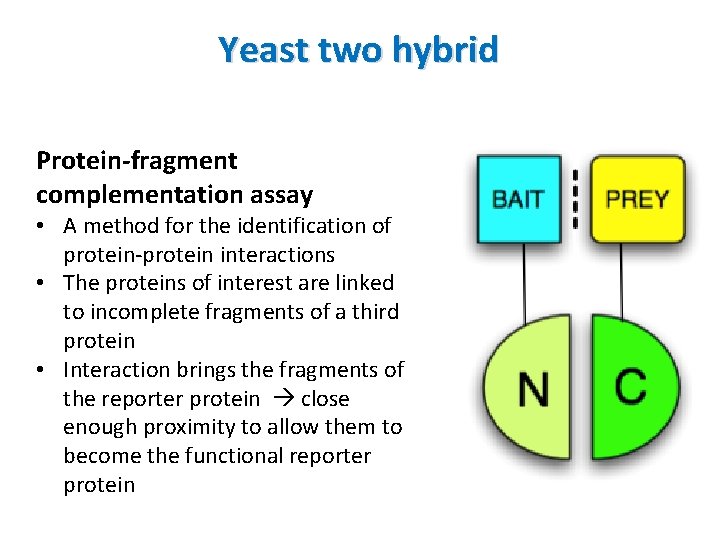
Yeast two hybrid Protein-fragment complementation assay • A method for the identification of protein-protein interactions • The proteins of interest are linked to incomplete fragments of a third protein • Interaction brings the fragments of the reporter protein close enough proximity to allow them to become the functional reporter protein

Yeast two hybrid Host Yeast (S. cerevisiae) • Ability to form tertiary protein structure • Neutral internal p. H • Complete genome sequence is known • Tolerant of diverse culture conditions and harsh chemicals that could not be applied to mammalian tissue culture E. coli • Higher transformation efficiency • Faster rate of growth • The absence of requirement for a nuclear localization signal • Study proteins which would be toxic to yeast

Yeast two hybrid Yeast Two hybrid - term • Transcription factor A protein that binds to specific DNA sequence and controls the flow of genetic information from DNA to m. RNA • Activator: promotes the recruitment of RNA polymerase • Repressor: blocks the recruitment of RNA polymerase Lac operon: CAP (CRP), Repressor I • Binary structure: DNA binding domain and trans-activation domain • Binds to either enhancer or promoter region of DNA up/down regulation • Mechanism: stabilize or block the binding of RNA polymerase to DNA, catalyze acetylation or deacetylation of histone or recruit coactivator or corepressor to the transcription factor DNA complex

Yeast two hybrid Yeast Two hybrid - terms • UAS (upstream activation sequence) Regulatory sequence, distinct from the promoter and increase the expression of a neighboring gene, activator binding site • Reporter gene A gene is attached to a regulatory sequence of another gene of interest (GFP, Luciferase, beta-galactosidase, auxotrophic genes) • Bait/Prey Bait-the gene you have, Prey-the gene you catch with the bait In yeast two hybrid, a bait protein is fused to the DNA binding domain and library (prey) is linked to the activation domain
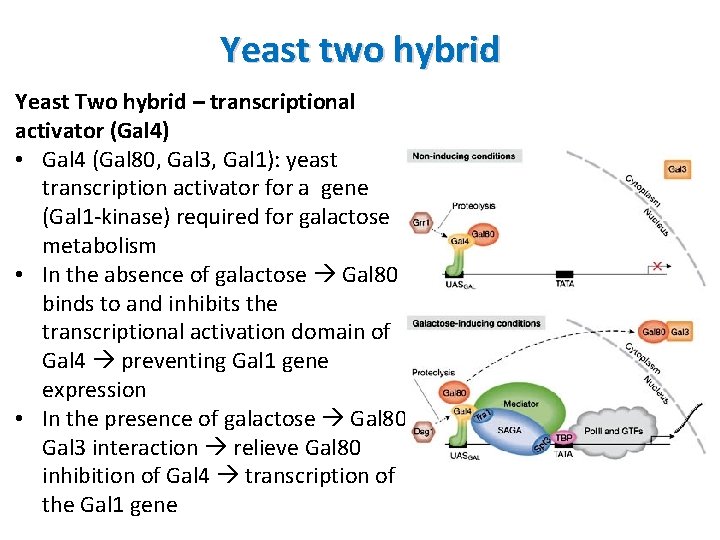
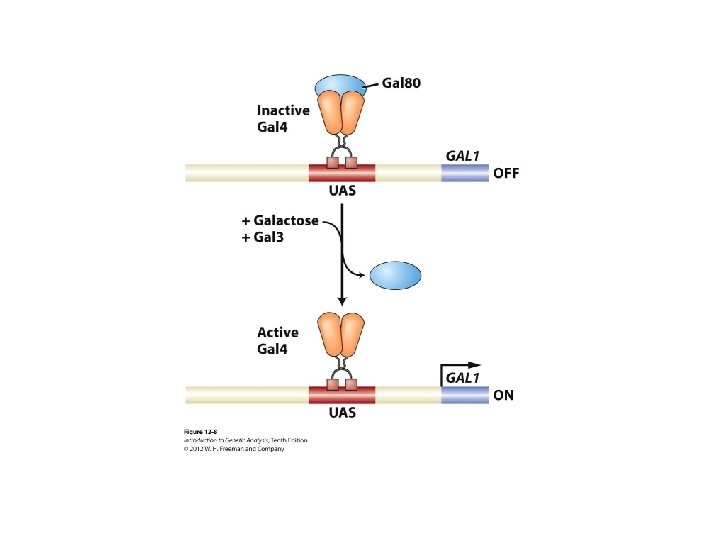
Yeast two hybrid Yeast Two hybrid – transcriptional activator (Gal 4) • Gal 4 (Gal 80, Gal 3, Gal 1): yeast transcription activator for a gene (Gal 1 -kinase) required for galactose metabolism • In the absence of galactose Gal 80 binds to and inhibits the transcriptional activation domain of Gal 4 preventing Gal 1 gene expression • In the presence of galactose Gal 80 Gal 3 interaction relieve Gal 80 inhibition of Gal 4 transcription of the Gal 1 gene
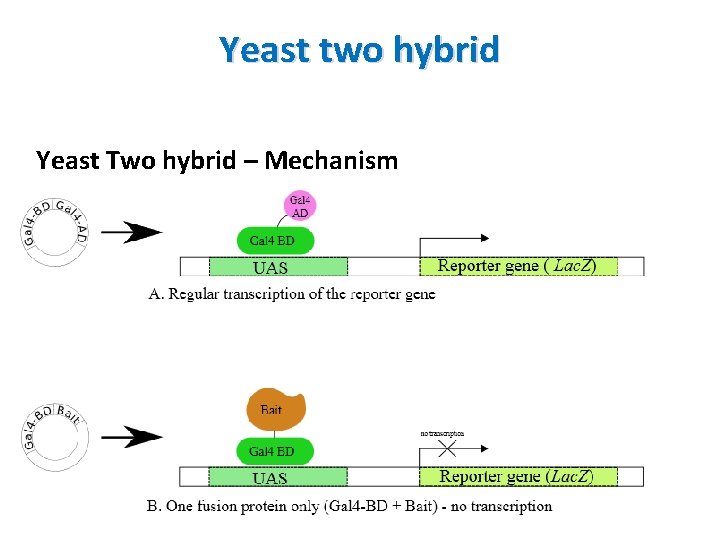
Yeast two hybrid Yeast Two hybrid – Mechanism
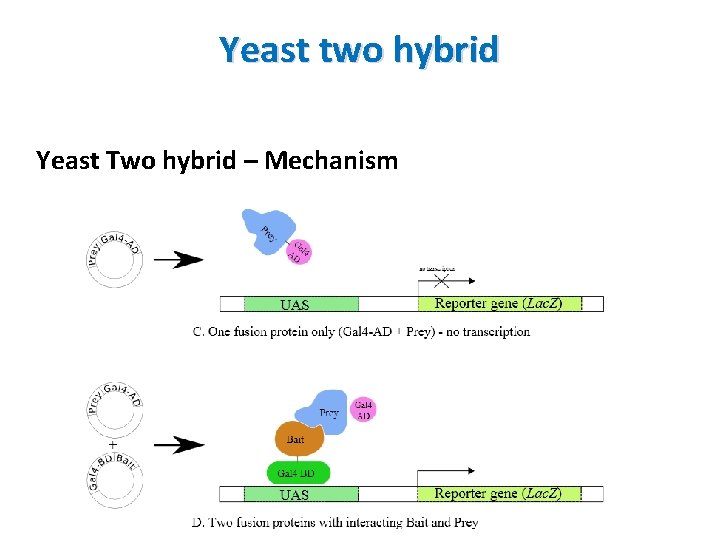
Yeast two hybrid Yeast Two hybrid – Mechanism
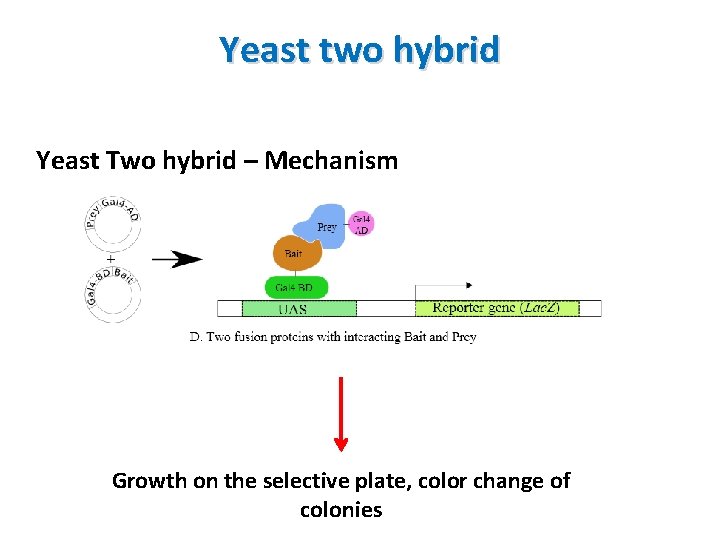
Yeast two hybrid Yeast Two hybrid – Mechanism Growth on the selective plate, color change of colonies

Yeast two hybrid Yeast Two hybrid – Strengths • Low-tech: carried out in any lab without sophisticated equipment • Cheap tech: not require expensive reagents • Provide an important first hint for the identification of interaction partners • Scalable: possible to screen for interactions among many proteins • Relatively high quality of interaction data generated (price vs data) • In vivo technique used to screen for interactions

Yeast two hybrid Yeast Two hybrid – weaknesses • Overexpression of fusion proteins cause unnatural protein concentration that lead to unspecific (false) positives • The hybrid proteins are fusion protein inhibit certain interactions, especially an interaction takes place at the N-terminus of a test protein • Prokaryotic protein or mammalian protein improper folding and modification • Test protein should be localized to the nucleus (other localization signals problematic) • High number of false positive and false negative identification
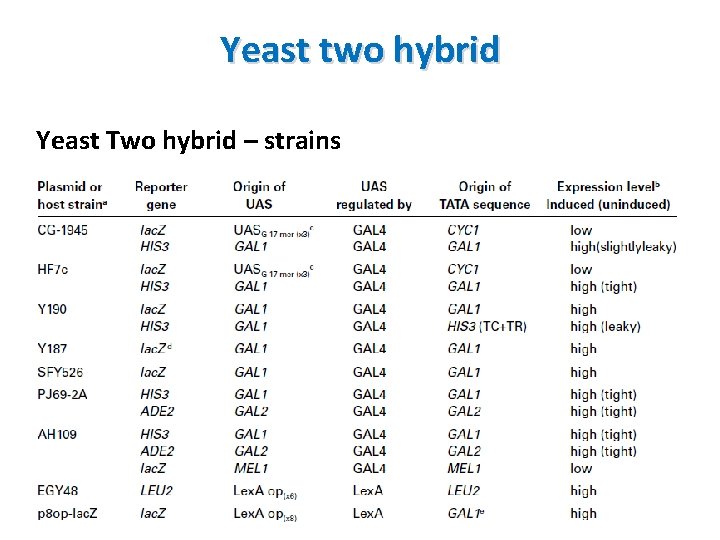
Yeast two hybrid Yeast Two hybrid – strains

Yeast two hybrid Yeast Two hybrid – strains

Yeast two hybrid Yeast Two hybrid-transformation/selection • Yeast transformation is similar to that of bacteria • It has the really thick cells wall hard to transfer of naked DNA into the cells • Li. Ac method loose the cell wall, make yeasts easily uptake DNA • Yeast vectors have autotrophic markers as selection markers in yeast media lacking certain amino acids used to select transformants • Auxotrophy: the inability of an organism to synthesize a particular organic compound required for its growth –amino acids
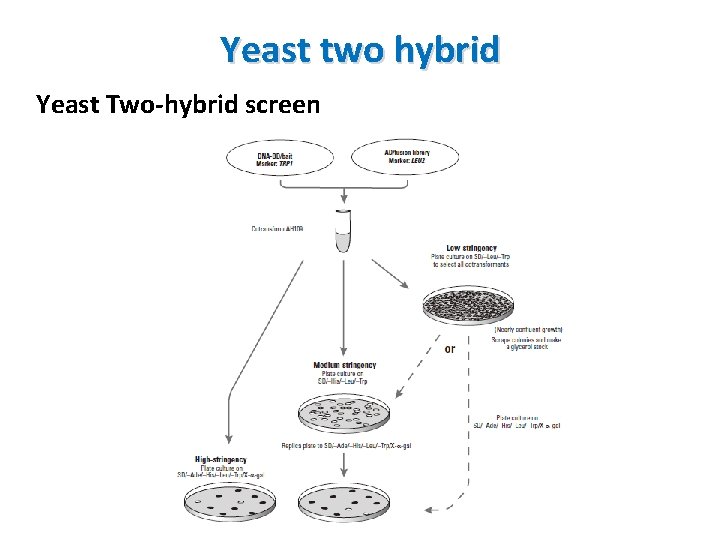
Yeast two hybrid Yeast Two-hybrid screen

Yeast two hybrid One hybrid • Designed to investigate Protein-DNA interactions • Use a single fusion protein: AD is linked directly to library which you want to test • Library can be selected against the desired target sequence which is inserted in the promoter region of the reporter gene
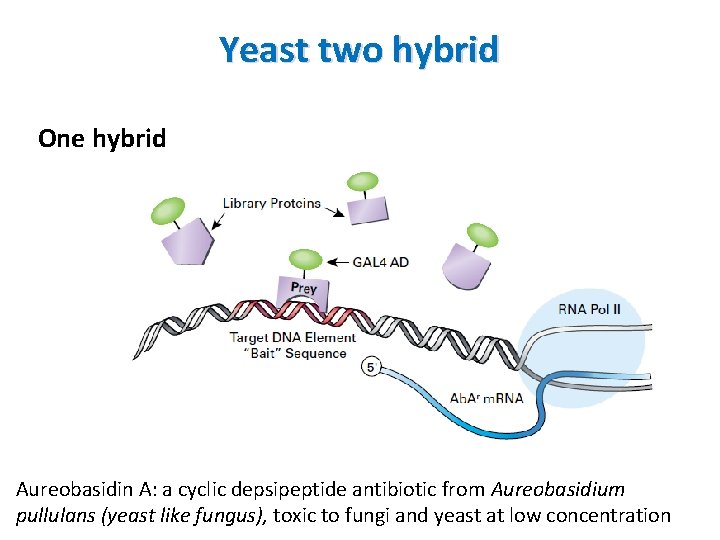
Yeast two hybrid One hybrid Aureobasidin A: a cyclic depsipeptide antibiotic from Aureobasidium pullulans (yeast like fungus), toxic to fungi and yeast at low concentration

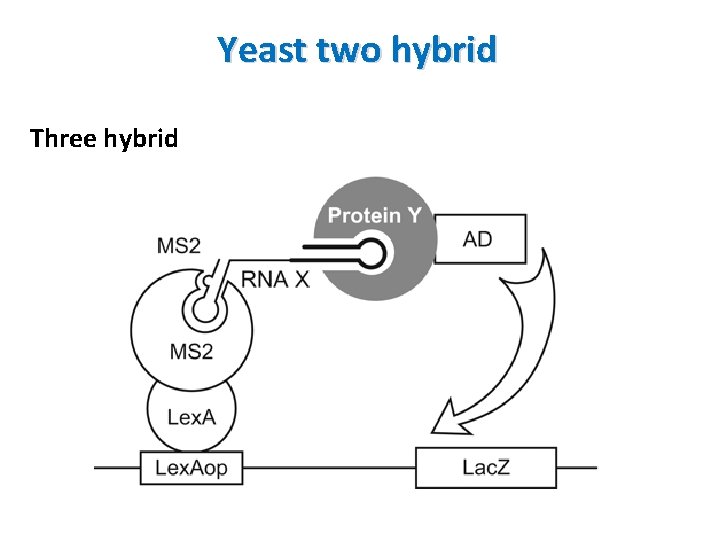
Yeast two hybrid Three hybrid • Used to detect RNA-protein interactions • Two fusion proteins interact each other through intermediary RNA molecule • BD is fused to a known RNA binding protein (MS 2 -the bacteriophage coat protein) and MS 2 binding RNA is linked to a target RNA of interest • Library is fused to activation domain • Hybrid RNA binds to a protein fused to activation domain (complex formation) reporter gene expression
Yeast two hybrid Three hybrid

Immunoprecipitation (IP) • The technique of precipitating a protein antigen out of solution using an antibody specifically binding to the protein • Used to isolate and concentrate a particular protein from a sample containing many thousands of different proteins • Requires that the antibody should be coupled to a solid substrate at some point in the procedure • Types of immunoprecipitation • Individual protein immunoprecipitation (IP) • Protein complex immunoprecipitation (Co-IP) • Chromatin immunoprecipitation (Ch. IP) • RNA immunoprecipitation (RIP) • Tagged proteins
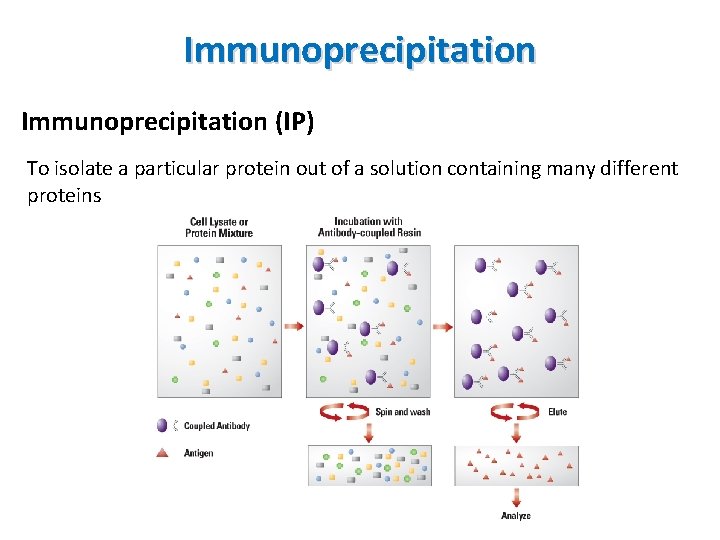
Immunoprecipitation (IP) To isolate a particular protein out of a solution containing many different proteins
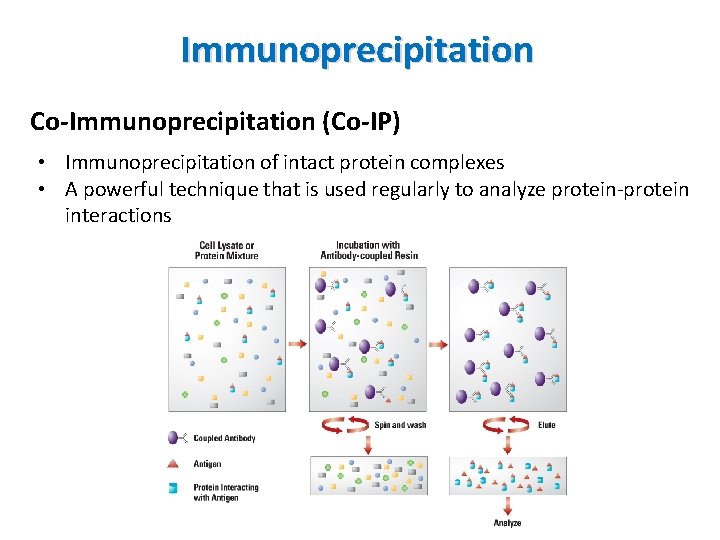
Immunoprecipitation Co-Immunoprecipitation (Co-IP) • Immunoprecipitation of intact protein complexes • A powerful technique that is used regularly to analyze protein-protein interactions

Immunoprecipitation Chromatin immunoprecipitation (Ch. IP) • A method to determine the location of DNA binding sites on the genome for a particular protein of interest • Based on DNA-protein interaction • Using an antibody specific to a putative DNA binding proteins immunoprecipitate the protein-DNA complex out of cellular lysates • Determine identity and quantity of the DNA fragment by PCR or cloning the DNA into a plasmid vector
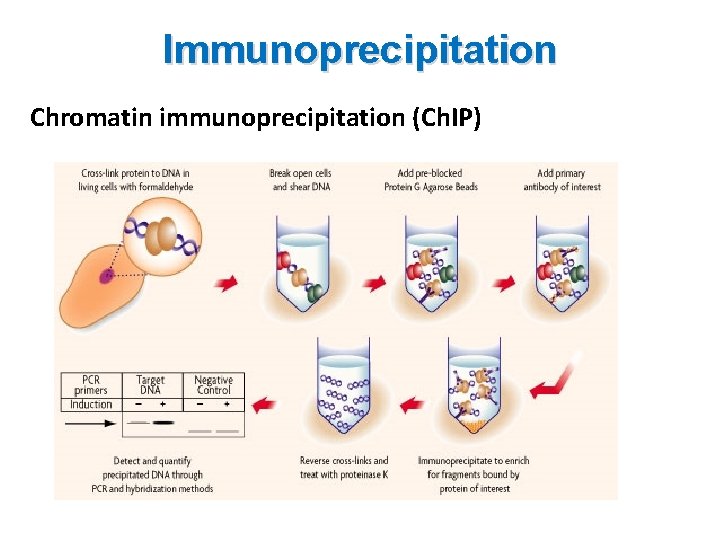
Immunoprecipitation Chromatin immunoprecipitation (Ch. IP)

Immunoprecipitation RNA immunoprecipitation (RIP) • Similar to chromatin immunoprecipitation • Targets RNA binding proteins rather than DNA binding proteins • Isolate RNA-protein complex using an antibody specific for a protein of interest • Determination of identity of RNA by RT-PCR and c. DNA sequencing
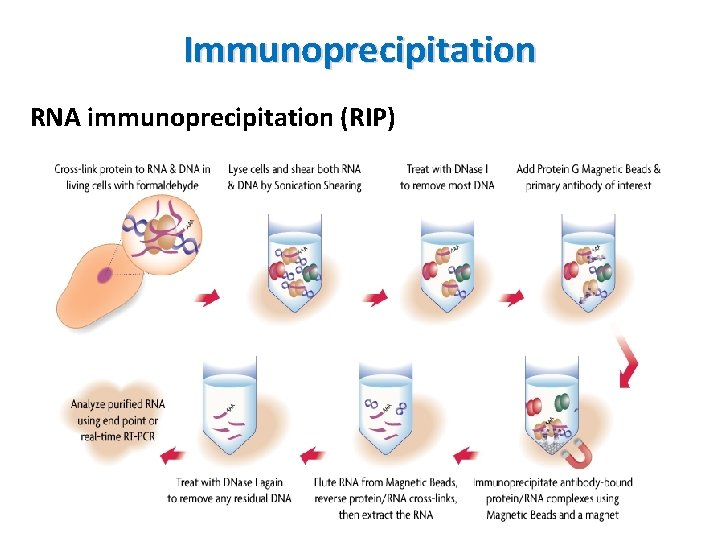
Immunoprecipitation RNA immunoprecipitation (RIP)

Immunoprecipitation Tagged proteins • Technical difficulty of immunoprecipiation: generation of an antibody specifically targeting a single known protein is the great difficulty • To get around this obstacle, tags onto either C- or Nterminal end of the protein of interest (GFP, Flag, HA tags, GST) • Presence of a good antibody against these tags and substances specifically binding to tags (immunoprecipitation or pull-down)

Pull down assay Common fusion tags and their affinity binding ligands Fusion tag Affinity ligand Glutathione S-transferase (GST) Glutathione Poly-histidine (polyhis or 6 xhis) Nickel or cobalt chelate complexes Biotin (vitamin B 7), Streptavidin (bacterial proteins), avidin
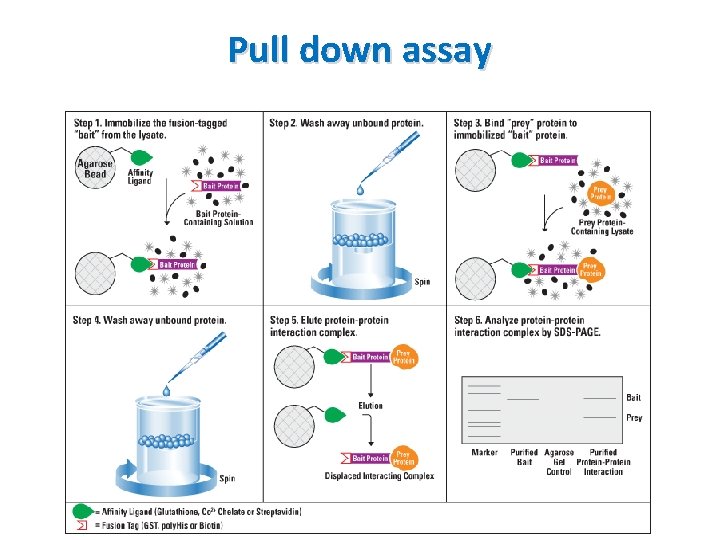
Pull down assay

Phage display • A laboratory technique for the study of protein-protein, proteinpeptide and protein-DNA interaction that uses bacteriphage to connect proteins with the gene encoding proteins/peptide • Bacteriophage: any one of a number of viruses that infect bacteria • Common phages in phage display: M 13, T 4, T 7 and l phage

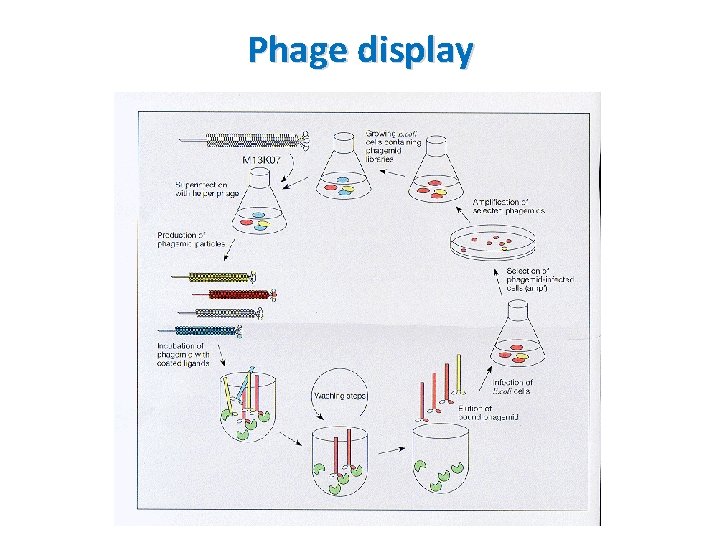
Phage display Principle • DNA encoding the protein or peptide of interest (or library, prey) is ligated into the p. III or p. VIII gene encoding either the minor or major coat protein of the virus • Exogenous proteins are expressed at the virus surface ( on the coat protein of virus) • A target molecules (bait-protein, peptide or DNA) is immobilized on the surface of a plate • Phage displaying a protein/peptide specifically binding to the target molecules (protein, peptide or DNA) will remain after washing unbound phages off
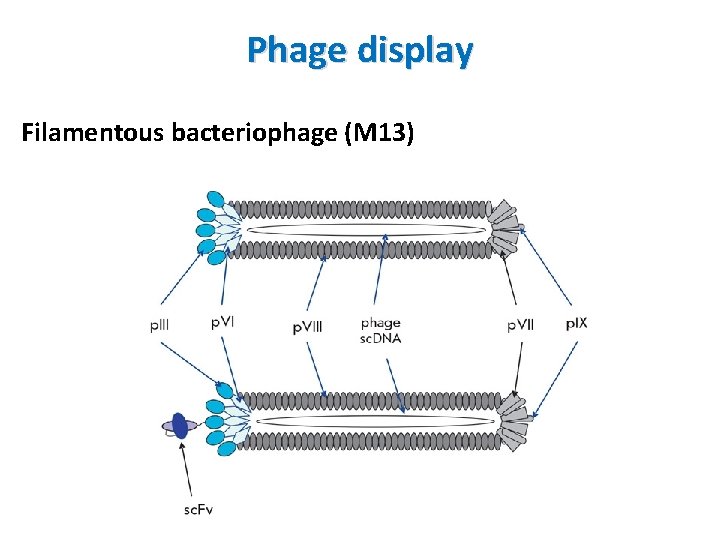
Phage display Filamentous bacteriophage (M 13)

Phage display General protocol • Target (bait) proteins or DNA sequences are immobilized to the wells of a microtiter plate. • Many genetic sequences are expressed in a bacteriophage library in the form of fusions with the bacteriophage coat protein, so that they are displayed on the surface of the viral particle. The protein displayed corresponds to the genetic sequence within the phage. • This phage-display library is added to the dish and after allowing the phage time to bind, the dish is washed.

Phage display General protocol • Phage-displaying proteins that interact with the target molecules remain attached to the dish, while all others are washed away. • Attached phage may be eluted and used to create more phage by infection of suitable bacterial hosts. The new phage constitutes an enriched mixture, containing considerably less irrelevant phage (i. e. non-binding) than were present in the initial mixture. • Above steps are optionally repeated one or more times, further enriching the phage library in binding proteins. • Following further bacterial-based amplification, the DNA within in the interacting phage is sequenced to identify the interacting proteins or protein fragments.
Phage display

Phage display Advantages • A means of rapidly screening large number of proteins against potential binding partners billions of clones can be screened within a week Disadvantages • The possibility of sterically hindering access to the displayed protein • Difficulty to express a protein larger than a coat protein

Tandem Affinity Purification (TAP) Principle • A technique for studying protein-protein interactions • A fusion protein with the TAP tag on the end • TAP tag consists of calmodulin binding peptide (CBP), tabacco etch virus protease (TEV protease) cleavage sit and protein A • TEV protease: a highly specific cysteine protease found in tabacco etch virus, recognize Glu-Asn-Leu-Tyr-Phe-Gln. Gly/Ser • Protein A – cell wall protein of bacteria, binds to immunoglobulins (Ig. G, Fc region of Ab) disrupts opsonization and phagocytosis
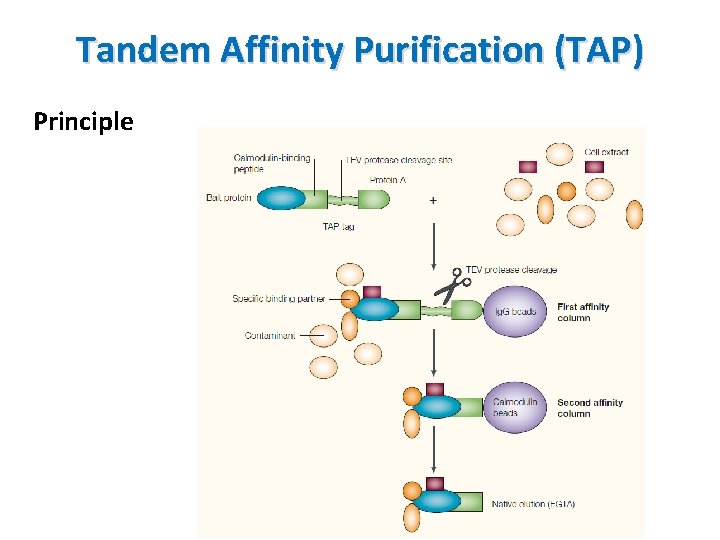
Tandem Affinity Purification (TAP) Principle

Tandem Affinity Purification (TAP) Advantages • Simple to execute and often provide high yield • Decrease the contamination of the target protein (two successive affinity purification) • It can also provide effective and highly specific means to purify a protein Disadvantages • Tag can obscure binding of interacting proteins to the target protein • Tag also can affect protein expression levels • Tag may not be sufficiently exposed to the affinity beads • The target protein may have TEV protease cleavage sites • Hard to detect transient interactions (due to successive steps of purification)

Bimolecular fluorescence complementation (Bi. FC) Principle • A technology used to validate protein interactions • Proteins to be tested are fused to unfolded complementary fragments of a fluorescent reporter protein • Interaction of proteins brings the fluorescent fragments within proximity and allows the reporter protein to reform its 3 -D structure and emit its fluorescent signal • The intensity of fluorescent signal is proportional to the intensity of interaction • Biochemical complementation proteins: ribonuclease, bglactosidase, GFP , YFP
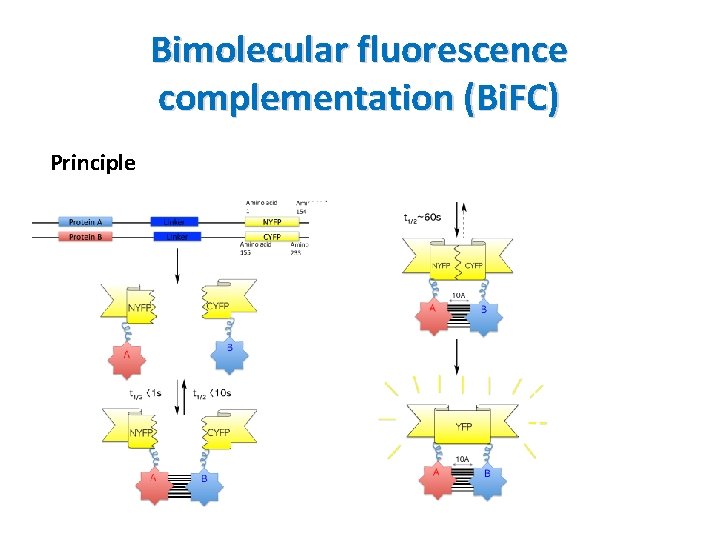
Bimolecular fluorescence complementation (Bi. FC) Principle

Bimolecular fluorescence complementation (Bi. FC) Strengths • Direct visualization: direct visualization rather than secondary effects or staining by exogenous molecules • Sensitivity: can detect weak interactions and low expression of proteins due to the stable complementation of the reporter proteins, strength of interaction measured by fluorescent signal strength • Spatial resolution: measure spatial protein-protein interaction (for example: drug responses) • No specialized equipment: a inverted fluorescent microscope is good enough • Live cell image

Bimolecular fluorescence complementation (Bi. FC) Limitations • Unable to detect real time detection of protein interactions (the signal is not produced right after interaction, takes hours) • Irreversible Bi. FC formation • Altering protein structure and steric hindrance • Fluorescence complementation may indicate either direct or indirect interaction • Obligate anaerobes: due to the requirement of oxygen for fluorophore formation, Bi. FC cannot be used in obligate anaerobes (used in aerobic organisms)

Fluorescence Resonance Energy Transfer (FRET) Principle • Useful tool to quantify molecular dynamics in biophysics and biochemistry such as protein-protein interactions, protein-DNA interactions and protein conformational changes • Proteins are labeled with a donor and acceptor • Two proteins are separated the donor emission is detected upon the donor excitation • The donor and acceptor are in proximity due to interaction the acceptor emission is predominantly observed because of the intermolecular fret from the donor to the acceptor • In order to detect conformational changes of a protein similar method, a target protein is labeled with both the donor and acceptor molecules
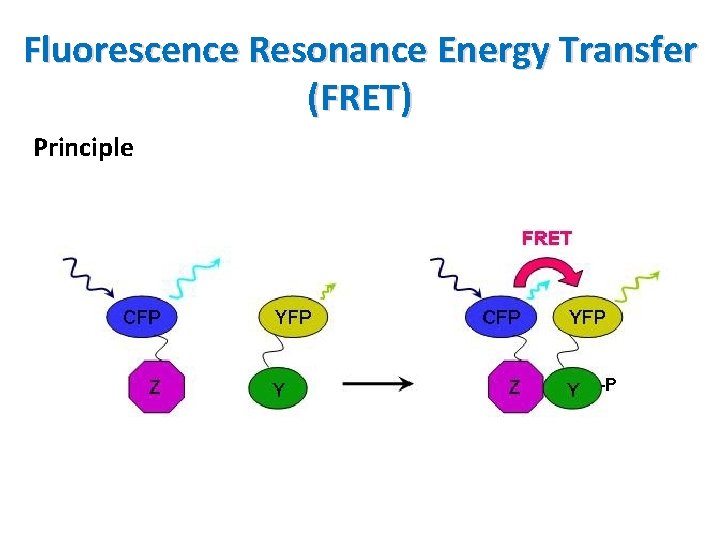
Fluorescence Resonance Energy Transfer (FRET) Principle

Bi. FC VS FRET Similarity • Ability to detect and locate protein interaction sites within live cells Differences • FRET need close spatial proximity (60 -100 A, Bi. FC: 7 nm) • FRET has decreased sensitivity (background difficult to detect weak interactions) • FRET has irreversible photo-bleaching (the fluorescence will be destroyed over time by the light needed to excite the fluorophores • FRET can detect instantaneous real-time protein interactions and dynamics of interaction due to reversible fluorophore interaction (advantage)

Protein Microarray • Also called a protein binding microarray • Provide a multiplex approach to identify protein-protein interactions, to identify the substrates of protein kinases, to identify transcription factor or to identify the targets of biologically active small molecules Principle § A piece of glass on which different molecules of protein are affixed at separate locations in an ordered manner labeled samples are added interacted proteins remained after washing
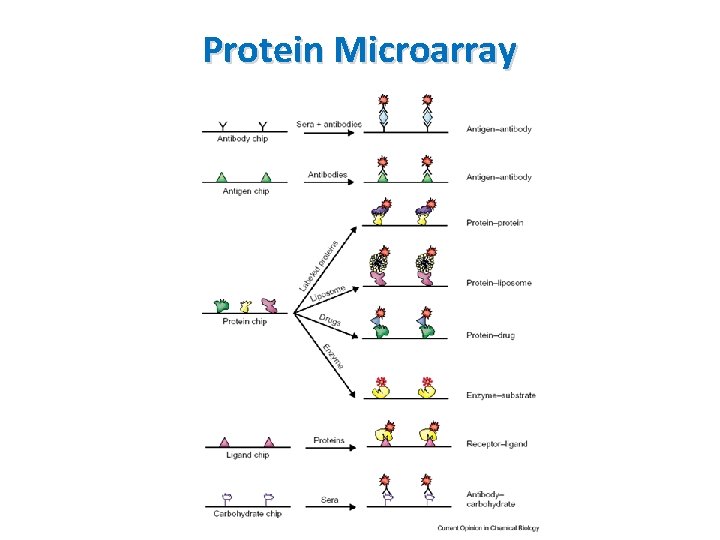
Protein Microarray
- Slides: 52