Protein Synthesis Chapter 17 Protein synthesis DNA Responsible

Protein Synthesis Chapter 17

Protein synthesis ¡ ¡ ¡ ¡ DNA Responsible for hereditary information DNA divided into genes Gene: Sequence of nucleotides Determines amino acid sequence in proteins Genes provide information to make proteins
Protein synthesis DNA RNA protein

Central Dogma Mechanism of reading & expressing genes ¡ Information passes from the genes (DNA) to an RNA copy ¡ Directs sequence of amino acids to make proteins ¡
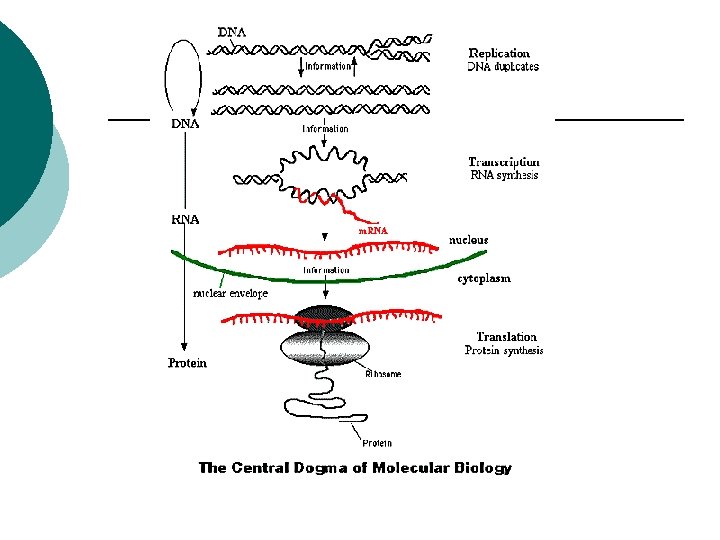

Protein synthesis Transcription: ¡ DNA sequence is copied into an RNA ¡ Translation: ¡ Information from the RNA is turned into an amino acid sequence ¡
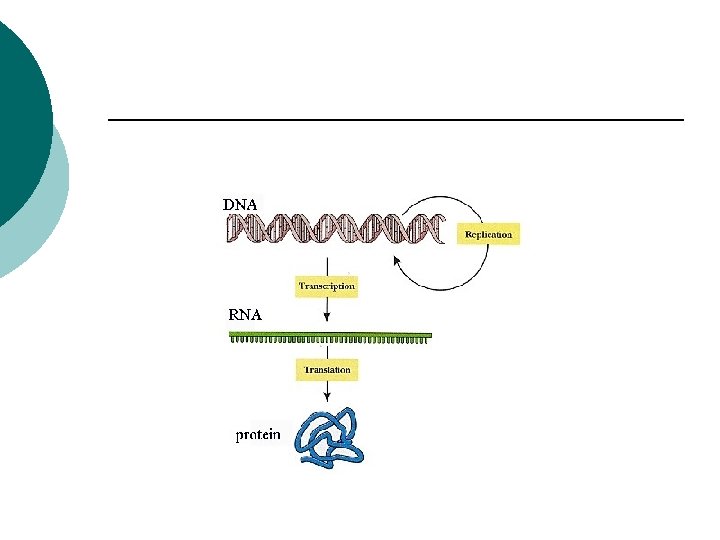
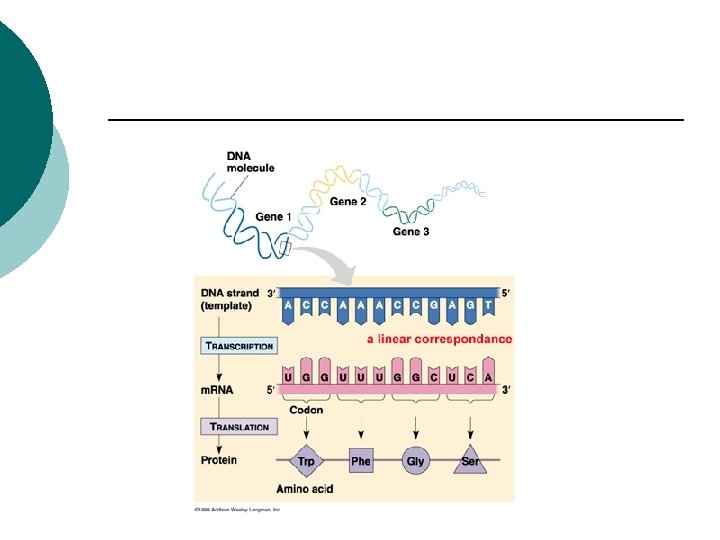
RNA (ribonucleic acid) ¡ Single strand ¡ Sugar –ribose (-OH on 2’ carbon) ¡ Uracil instead of thymine ¡
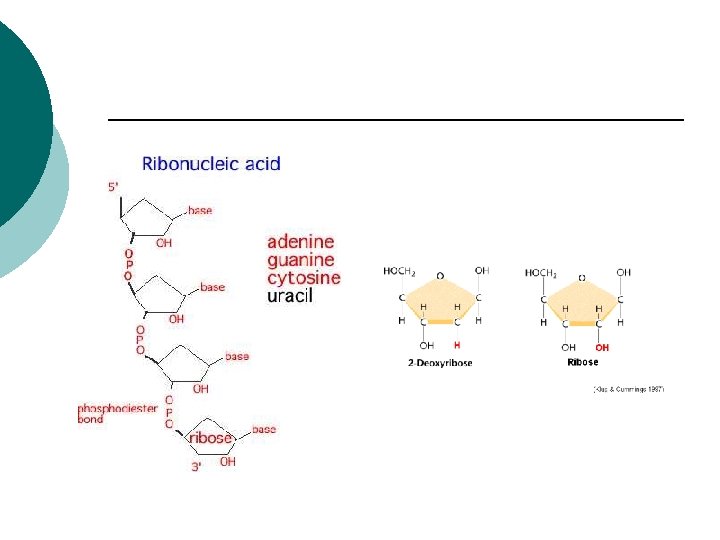
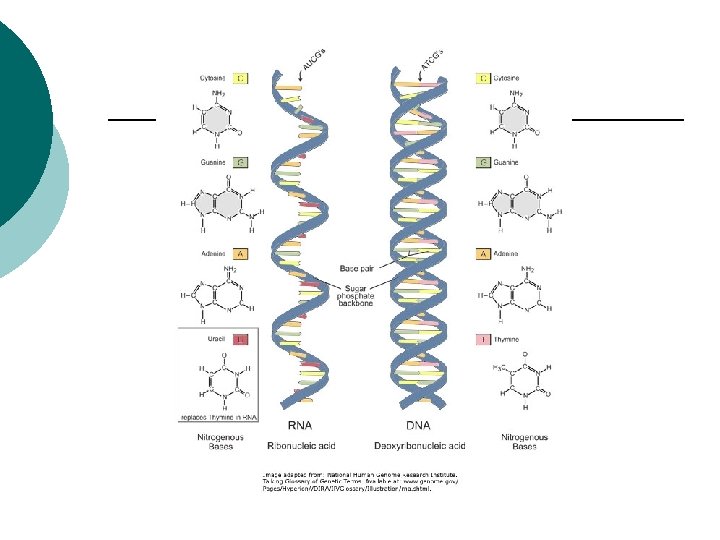

RNA ¡ ¡ ¡ ¡ ¡ m. RNA: Messenger RNA Transcribes information from DNA Codons (3 nucleotides) CGU m. RNA Codes for amino acids r. RNA: Ribosomal RNA Polypeptides are assembled

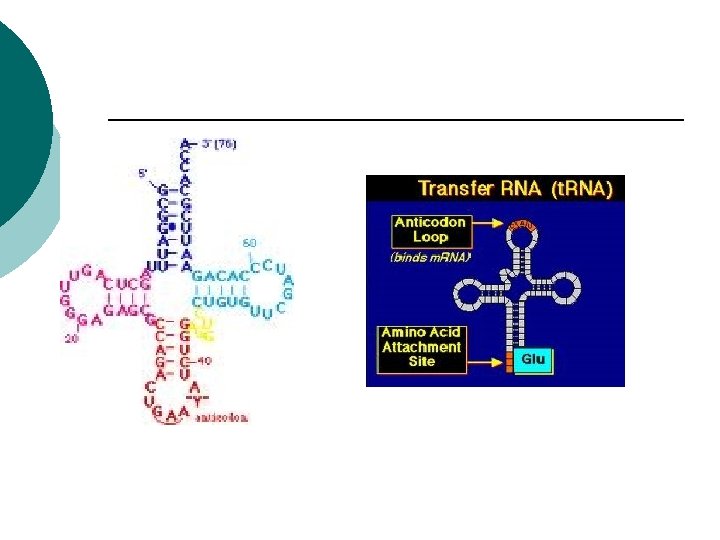
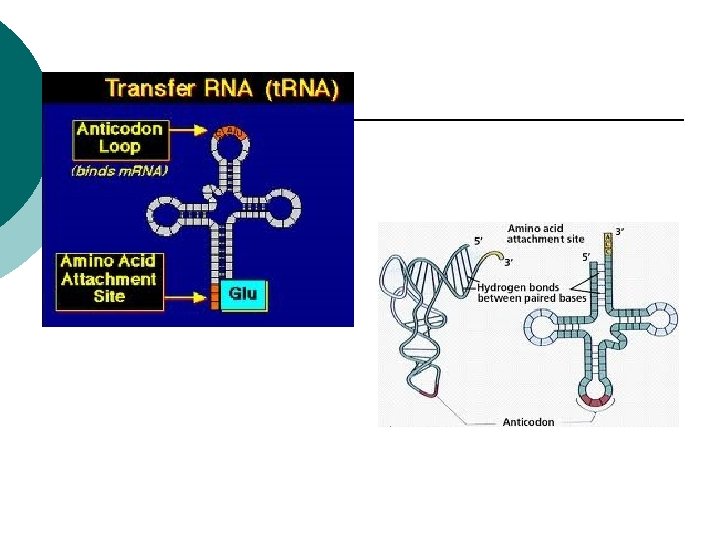
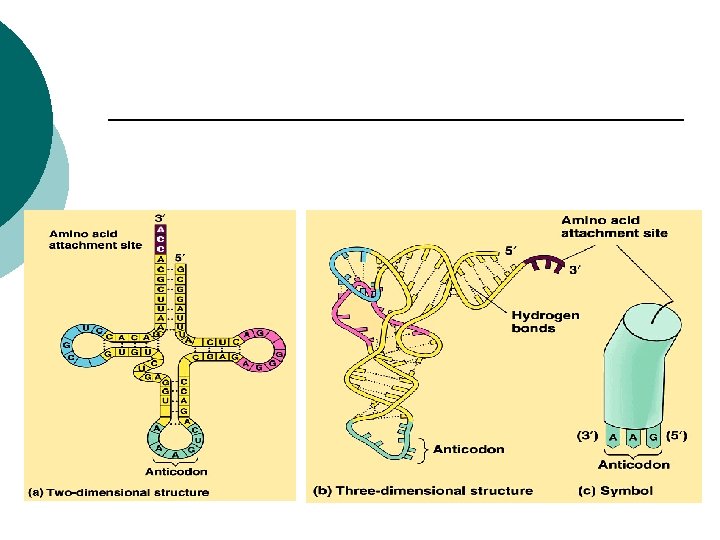
RNA t. RNA: ¡ Transfer RNA ¡ Transports aa to build proteins ¡ Positions aa on r. RNA ¡ Anticodons ¡ (3 complementary nucleotides) GCA ¡

Cracking the code Francis Crick ¡ Codons (Triplet code)-m. RNA ¡ Each codon corresponds to an aa ¡ 20 amino acids ¡ Reading frame ¡ Reading symbols in correct groupings ¡

Cracking the code 1 or 2 deletions or additions ¡ Gene was transcribed incorrectly ¡ 3 deletions ¡ Reading frame would shift ¡ Gene was transcribed correctly ¡

WHYDIDTHEREDCATEATTHEFATRAT WHYDTHEREDCATEATTHEFATRAT WHYTHEREDCATEATTHEFATRAT
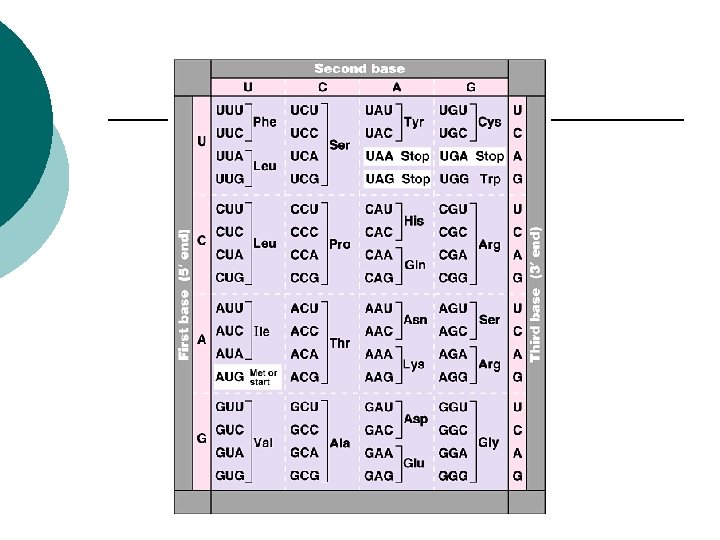

The code Universal code ¡ AGA codes for amino acid Arginine ¡ Humans & bacteria ¡ Genes from humans can be transcribed by m. RNA from bacteria ¡ Produce human proteins ¡ Insulin ¡
Protein synthesis DNA RNA Transcription Protein Translation
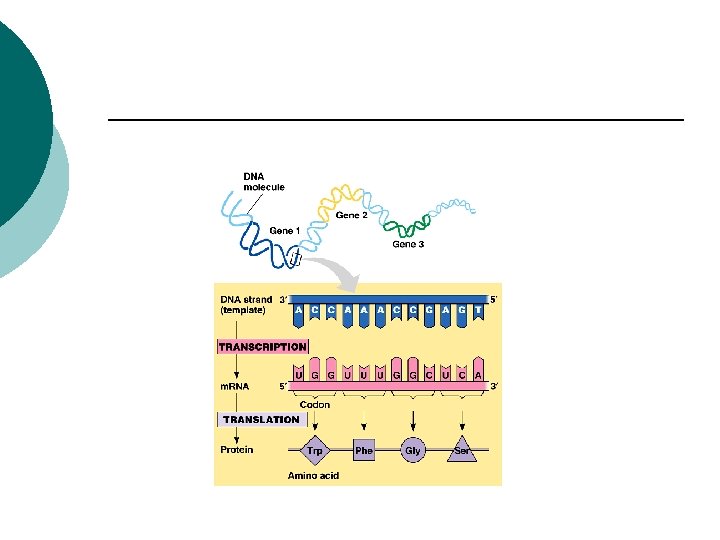
Prokaryotes Transcription ¡ Getting the code from DNA ¡ Template strand ¡ Strand of DNA that is transcribed or read ¡ Transcribed RNA is complementary to the DNA ¡

Prokaryotes Coding strand ¡ DNA strand not coded ¡ Same sequence of nucleotides as the RNA transcript ¡ Only T instead of U. ¡

Prokaryotes RNA polymerase ¡ Enzyme ¡ Adds nucleotides to the 3’end ¡ 5’to 3’ direction ¡ Does not need a primer to start ¡

Prokaryotes Stages of transcription ¡ Initiation ¡ Elongation ¡ Termination ¡

Prokaryotes Initiation ¡ Promoters: ¡ Sequence on DNA where transcription starts ¡ -35 sequence TTGACA ¡ -10 sequence TATAAT ¡ Sequences are not transcribed ¡

Prokaryotes RNA polymerase binds promoter ¡ Unwinds DNA ¡ Uses an ATP or GTP to start ¡ Uses phosphate group ¡ Transcription bubble: ¡ RNA polymerase, DNA & growing RNA strand ¡

Prokaryotes Termination ¡ Stop signal ¡ Sequence on DNA ¡ RNA transcript signals polymerase to detach from DNA ¡ RNA strand separates from the DNA ¡
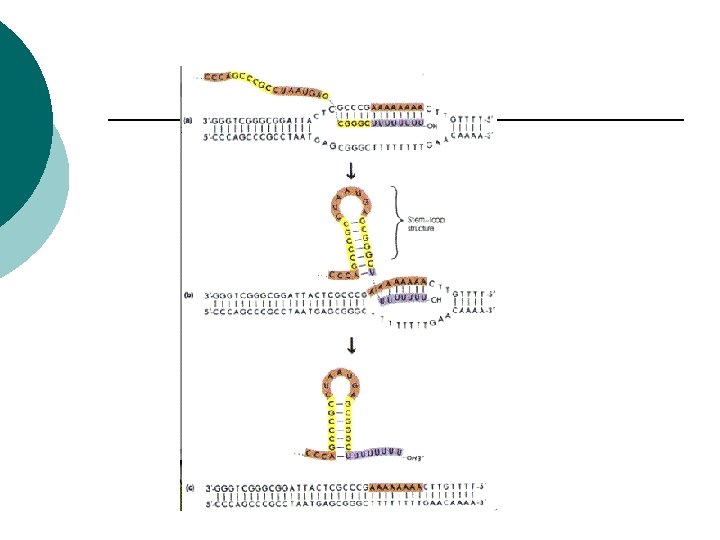
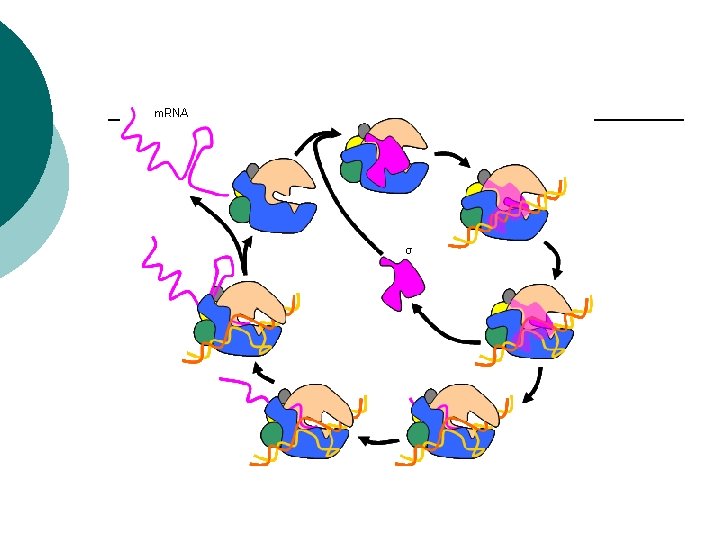

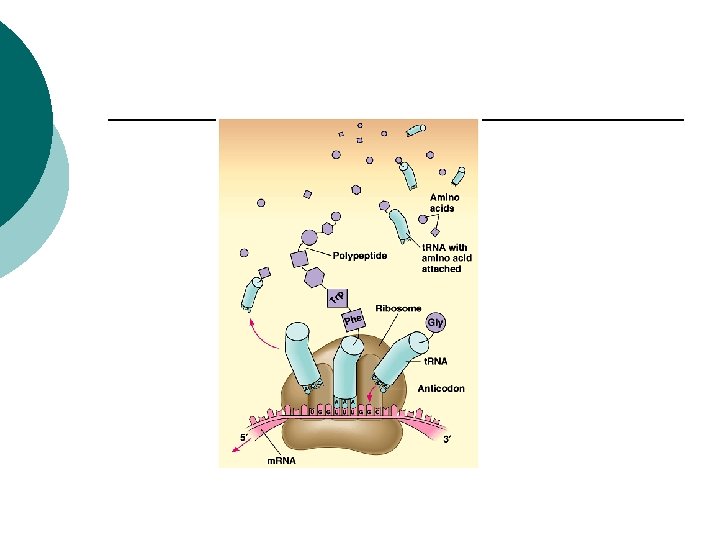
Prokaryotes Translation ¡ Passing the code to make a polypeptide ¡ m. RNA binds to r. RNA on the ribosome ¡ m. RNA attaches so only one codon is exposed at a time ¡

Ribosome Located in the cytoplasm ¡ Site of translation ¡ 2 subunits composed of protein & RNA ¡ Small (20 proteins and 1 RNA) ¡ Large (30 proteins and 2 RNA) ¡ 3 sites on ribosome surface involved in protein synthesis ¡ E, P, and A sites ¡
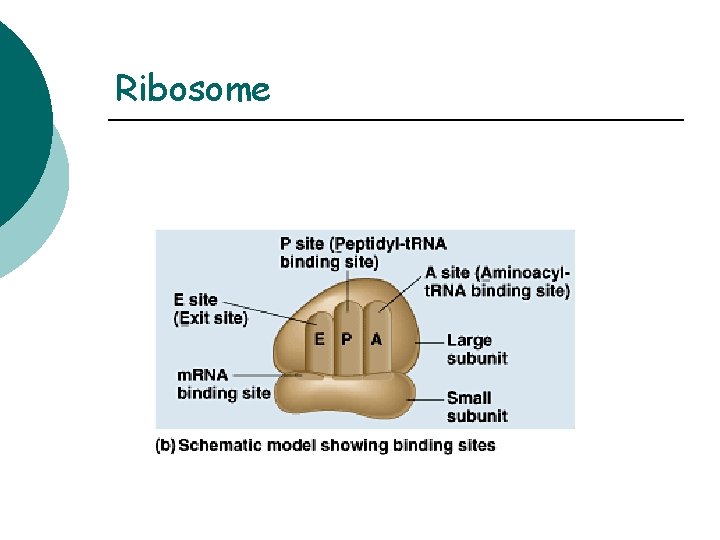
Ribosome
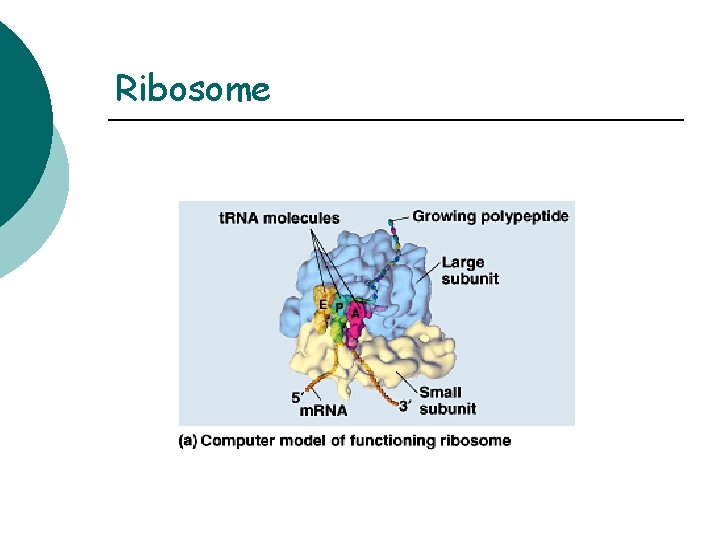
Ribosome

Prokaryotes t. RNA (anti-codon) ¡ Complementary sequence ¡ Binds to m. RNA ¡ t. RNA carries a specific amino acid ¡ Adds to growing polypeptide ¡ 45 t. RNA’s ¡

Prokaryotes Aminoacyl-t-RNA synthetases ¡ Activating enzymes ¡ Link correct t. RNA code to correct aa ¡ One for each 20 amino acids ¡ Some read one code, some read several codes ¡
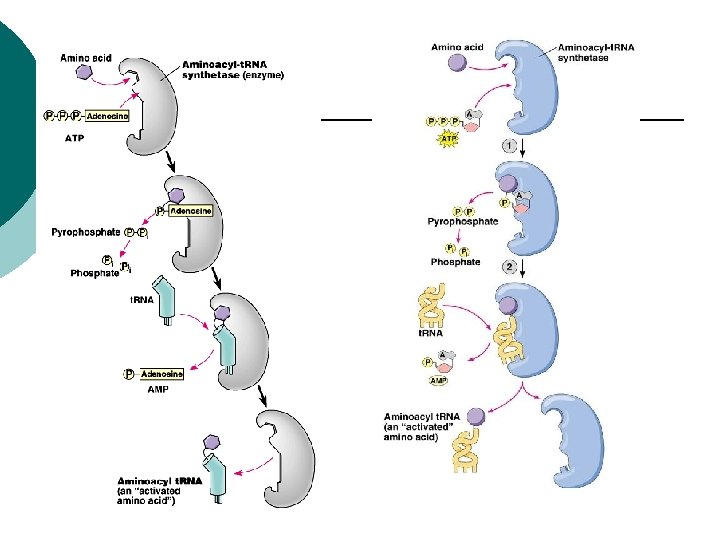

Prokaryotes Nonsense codes ¡ UAA, UAG, UGA code to stop ¡ AUG codes for start as well as methionine ¡ Ribosome starts at the first AUG it comes across in the code ¡

Prokaryotes Translation ¡ 1. Initiation ¡ 2. Elongation ¡ 3. Termination ¡

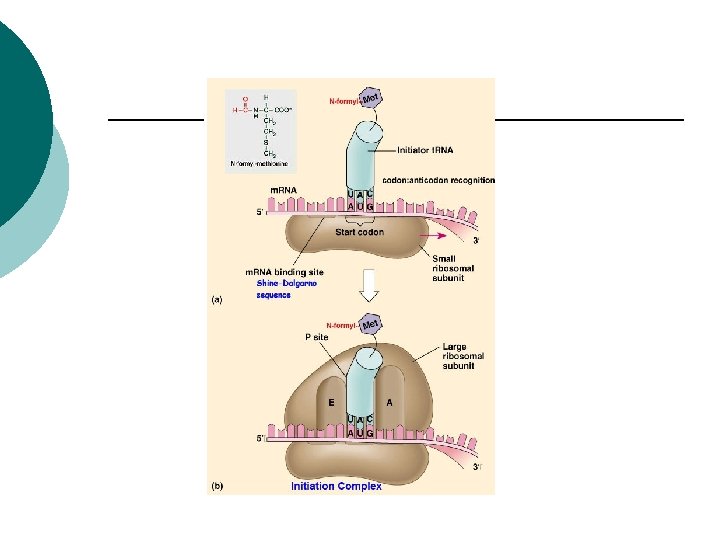
Prokaryotes ¡ ¡ ¡ Initiation complex 1. t. RNA with formylmethionine attached binds to a small ribosome 2. Initiation factors position the t. RNA on the P site 3. A site (aminoacyl) where other t. RNA’s form

Prokaryotes 4. t. RNA is positioned on to the m. RNA at AUG ¡ 5. Attachment of large ribosomal unit ¡

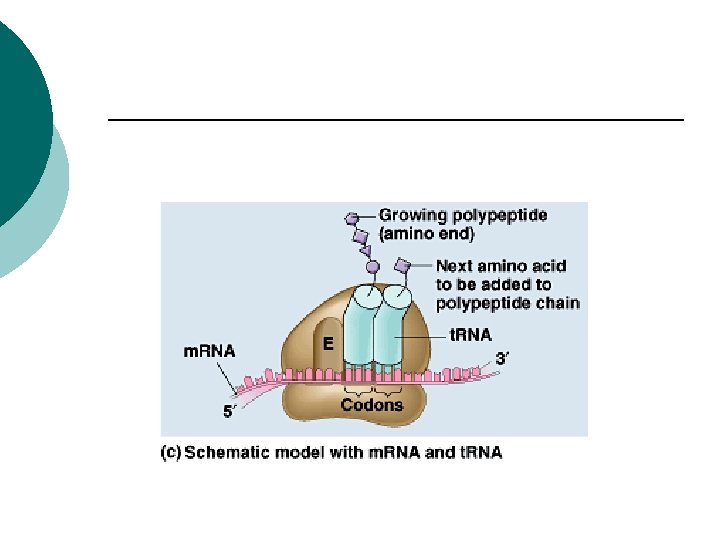
Prokaryotes Elongation factors ¡ Help second t. RNA bind to the A-site ¡ Two amino acids bind (peptide bond) ¡ Translocation: ¡ Ribosome moves 3 more nucleotides along m. RNA in the 5’to 3’ direction ¡

Prokaryotes Initial t. RNA moves to E site ¡ Released ¡ New t. RNA moves into A site ¡ Continues to add more aa to form the polypeptide ¡
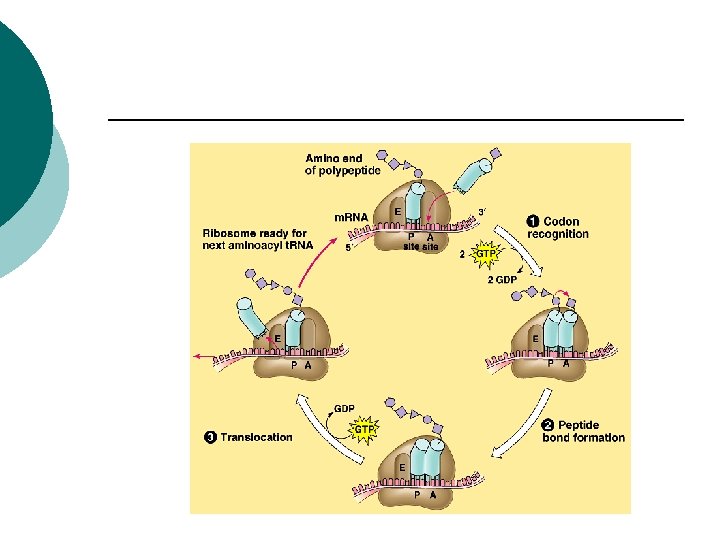

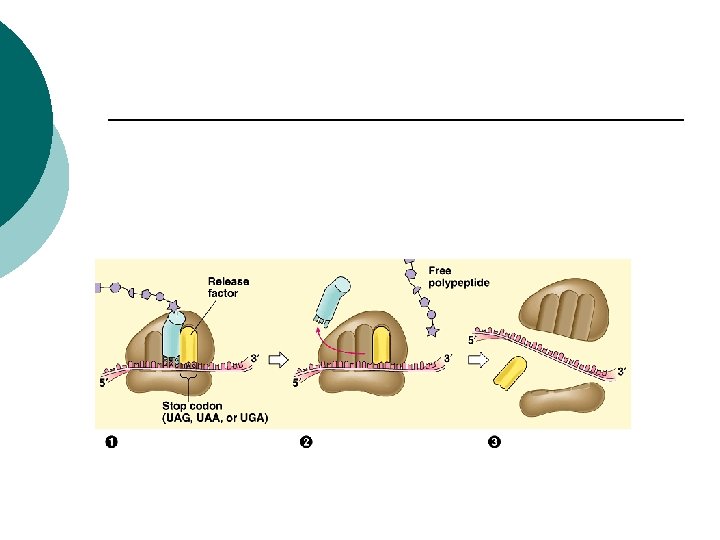
Prokaryotes Release factors: ¡ Proteins that release newly made polypeptides ¡ Codon (UAG, UAA, UGA) ¡ Release factor binds to the codon ¡ Polypeptide chain is released from A site ¡
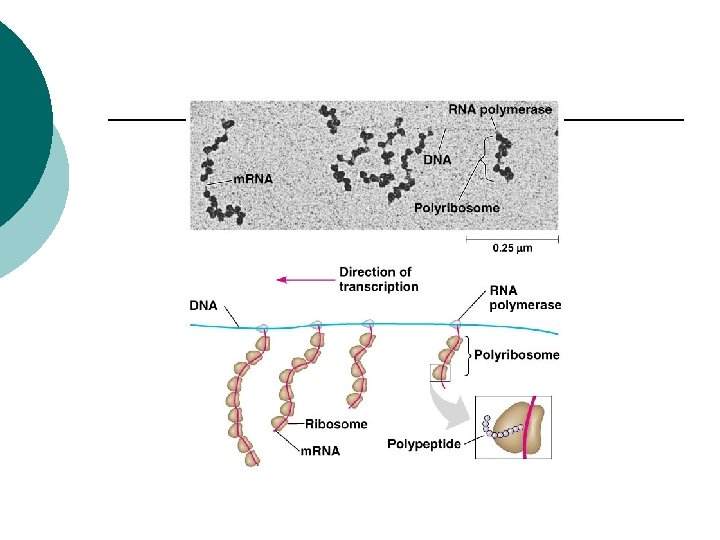

Eukaryotes Transcription (nucleus) ¡ Initiation ¡ Elongation ¡ Termination ¡

Eukaryotes Initiation ¡ Transcription Initiation Complex is formed ¡ Transcription factors bind first to the promoter ¡ RNA pol II binds DNA ¡ Starts to transcribe ¡
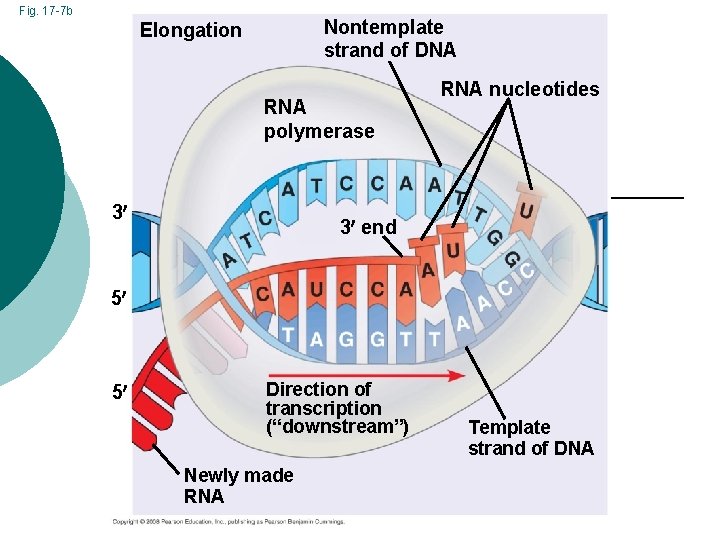
Fig. 17 -7 b Nontemplate strand of DNA Elongation RNA polymerase 3 RNA nucleotides 3 end 5 5 Direction of transcription (“downstream”) Newly made RNA Template strand of DNA
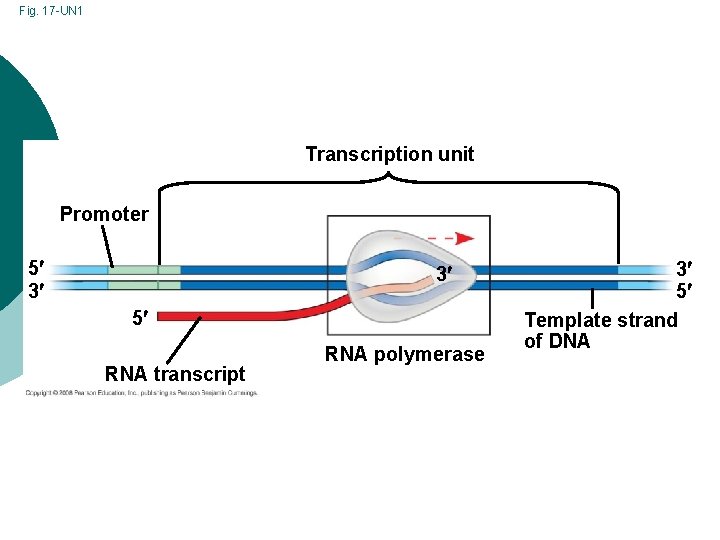
Fig. 17 -UN 1 Transcription unit Promoter 5 3 3 5 RNA transcript RNA polymerase 3 5 Template strand of DNA

Eukaryotes Termination ¡ Polyadenylation signal sequence ¡ Recognized by RNA polymerase II ¡ m. RNA is released ¡

Transcription D: Chapter_17A_Power. Point_Lectures17_Lectu re_Presentation1707 Transcription. Intro. A. html
Eukaryotes m. RNA is modified ¡ Nucleus ¡ RNA processing ¡

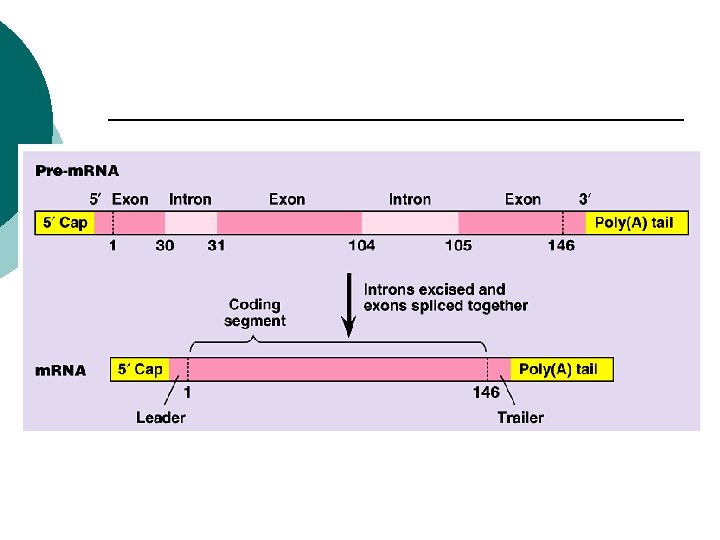
Eukaryotes 5’ cap ¡ Addition of a GTP ¡ 5’ phosphate of the first base of m. RNA ¡ Methyl group is added to the GTP ¡ 3’poly-A-tail ¡ Several A’s on the end of the m. RNA ¡

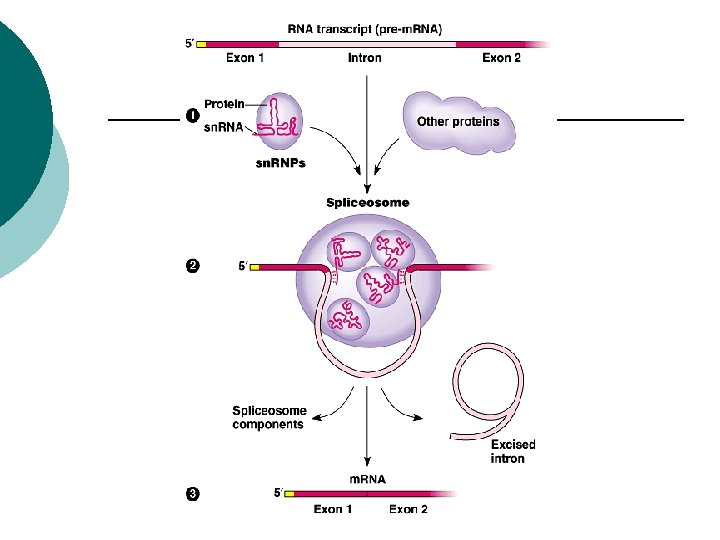
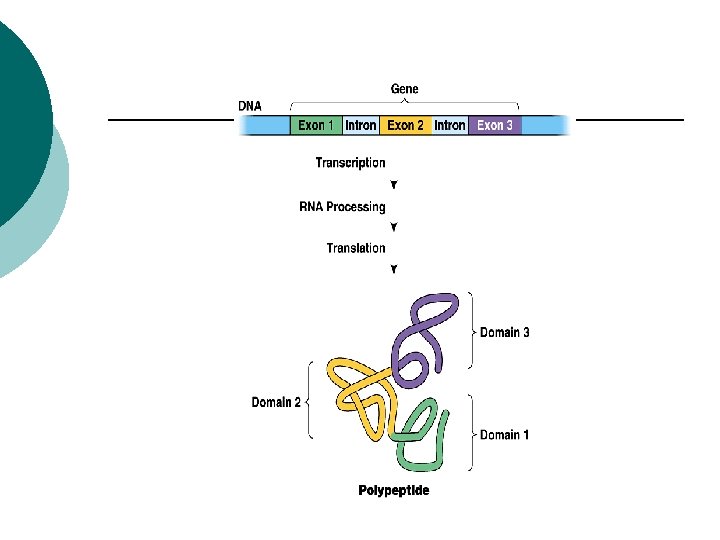
Eukaryotes Introns: ¡ non-coding sequences of nucleic acids ¡ Exons: ¡ coding sequences of nucleic acids ¡
Euraryotes RNA splicing ¡ Cut out introns ¡ Reconnect exons ¡ sn. RNP’s (small nuclear RNA’s) ¡ Spliceosome: ¡ Many sn. RNP’s come together & remove introns ¡
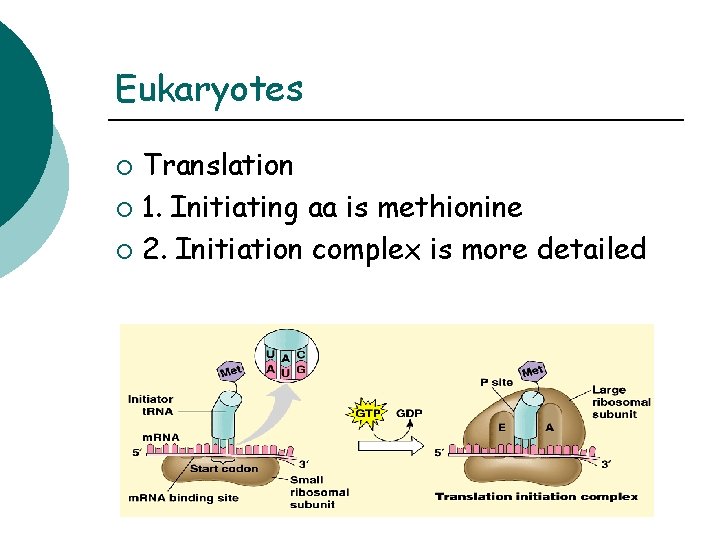
Eukaryotes Translation ¡ 1. Initiating aa is methionine ¡ 2. Initiation complex is more detailed ¡
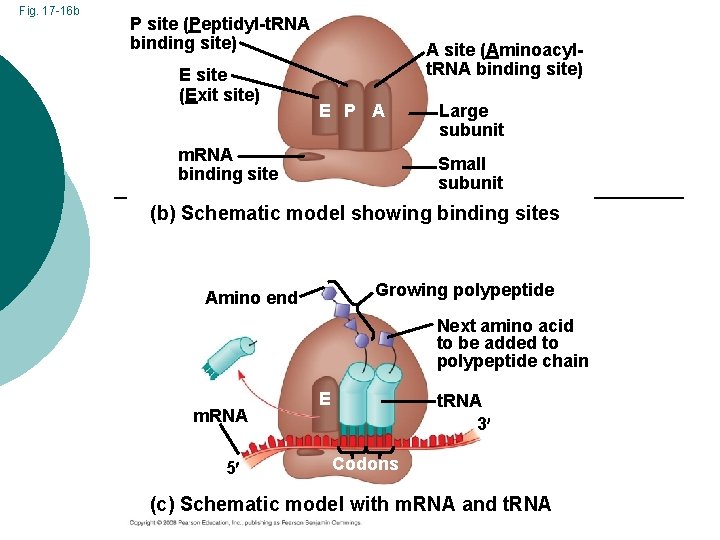
Fig. 17 -16 b P site (Peptidyl-t. RNA binding site) E site (Exit site) A site (Aminoacylt. RNA binding site) E P A m. RNA binding site Large subunit Small subunit (b) Schematic model showing binding sites Growing polypeptide Amino end Next amino acid to be added to polypeptide chain m. RNA 5 E t. RNA 3 Codons (c) Schematic model with m. RNA and t. RNA
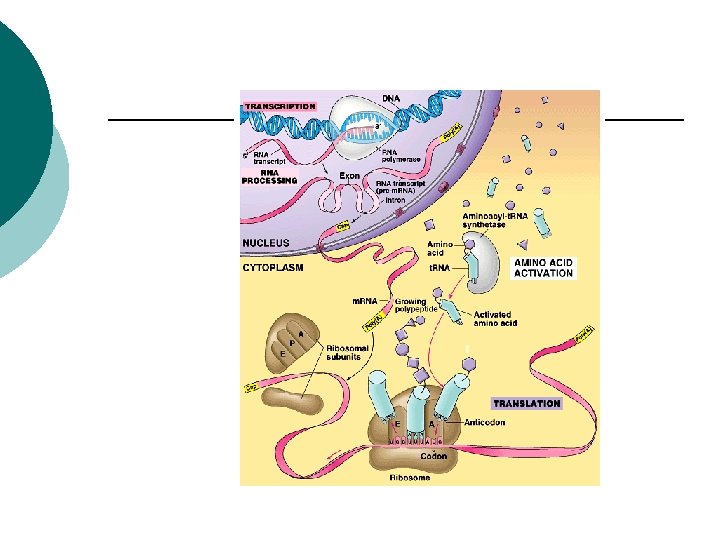
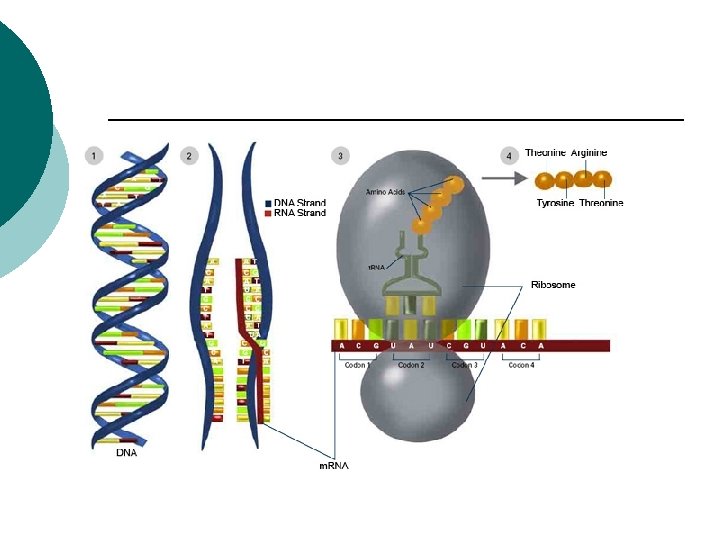
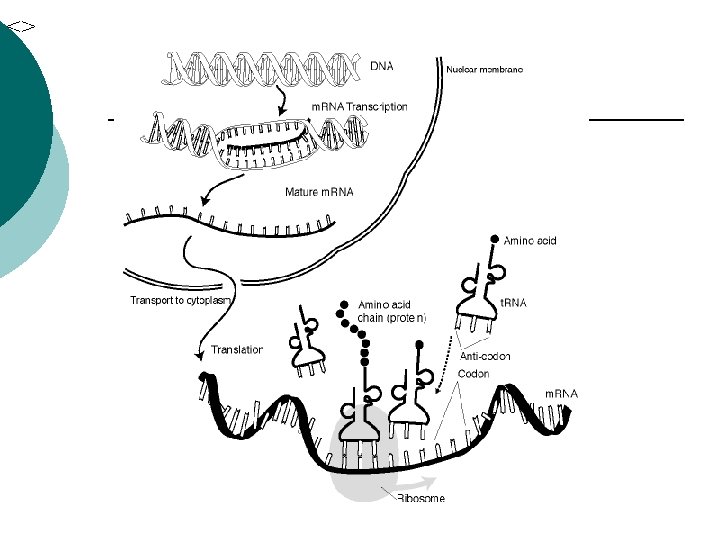
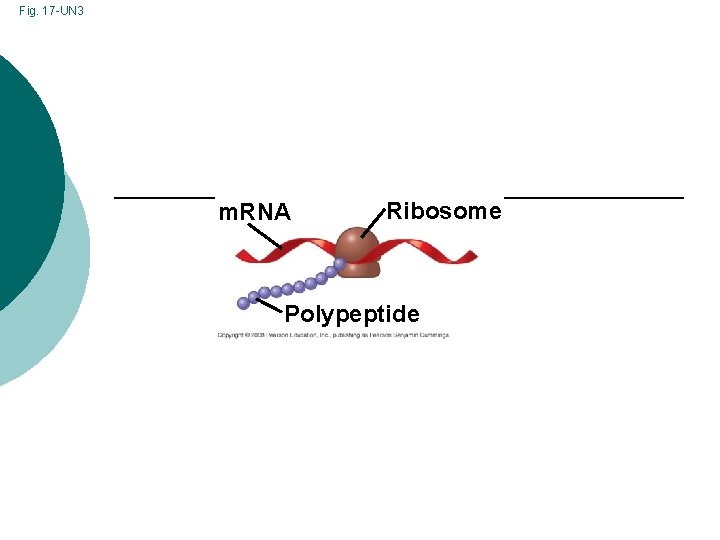
Fig. 17 -UN 3 m. RNA Ribosome Polypeptide

<> D: Chapter_17A_Power. Point_Lectures17_Le cture_Presentation1718 Translation. Intro. A. html
Similarities DNA RNA Transcription Protein Translation

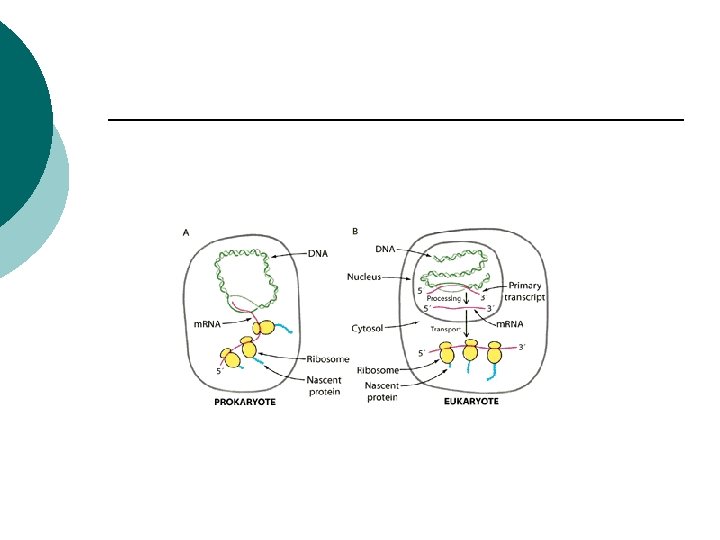
Differences in gene expression ¡ ¡ ¡ Transcription 1. Prokaryotes one RNA polymerase Eukaryotes 3 RNA polymerases (poli-II m. RNA synthesis) 2. Prokaryotes m. RNA contain transcripts of several genes Eukaryotes only one gene 3. Prokaryotes no nucleus so start translation before transcription is done

Differences in gene expression ¡ ¡ 3. Eukaryotes complete transcription before leaving the nucleus 4. Eukaryotes modify RNA l ¡ ¡ ¡ Introns/exons 5. Prokaryotes Polymerase binds promoters Eukaryotes transcription factors bind first then enzyme 6. Termination

Differences in gene expression Translation ¡ 1. Prokaryotes start translation with AUG ¡ Eukaryotes 5’cap initiates translation ¡ 2. Prokaryotes smaller ribosomes ¡

Mutations Changes in genetic information ¡ Point mutations: ¡ Change in a single base pair ¡ Sickle cell mutation ¡

Mutations Two types ¡ 1. Base-pair substitution ¡ Exchange one nucleotide and base pair with another ¡ Silent mutations ¡ No effect on proteins ¡

Mutations Missense mutations: ¡ Substitutions that change one aa for another ¡ Little effect ¡

Mutations Nonsense mutations ¡ Point mutation codes for stop codon ¡ Stops translation too soon ¡ Shortens protein ¡ Non-functional proteins ¡

Mutations 2. Insertions or deletions ¡ Additions or losses of nucleotides ¡ Frameshift mutations ¡ Improperly grouped codons ¡ Nonfuctional proteins ¡
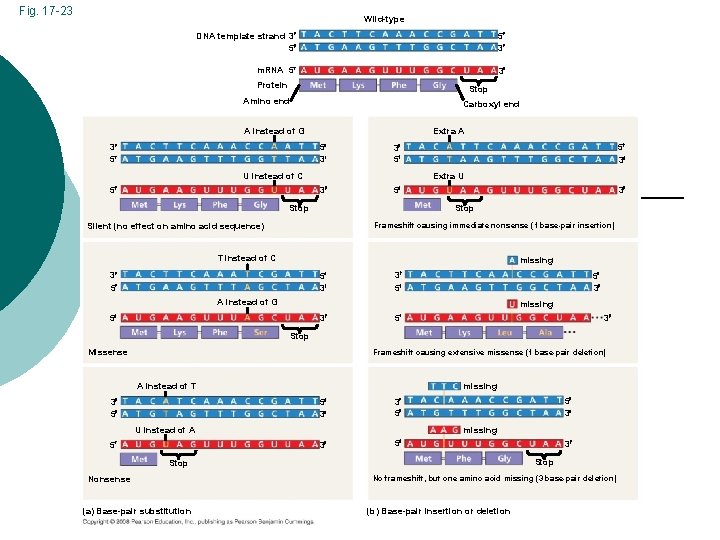
Fig. 17 -23 Wild-type DNA template strand 3 5 5 3 m. RNA 5 3 Protein Stop Amino end Carboxyl end A instead of G 3 5 Extra A 5 3 U instead of C 5 5 3 3 5 Extra U 3 5 3 Stop Silent (no effect on amino acid sequence) Frameshift causing immediate nonsense (1 base-pair insertion) T instead of C missing 3 5 5 3 A instead of G missing 3 5 5 3 Stop Missense Frameshift causing extensive missense (1 base-pair deletion) missing A instead of T 3 5 5 3 3 5 3 5 U instead of A 5 5 3 missing Stop Nonsense (a) Base-pair substitution 3 No frameshift, but one amino acid missing (3 base-pair deletion) (b) Base-pair insertion or deletion

Mutagens Chemical or physical agents ¡ Mutations in DNA ¡
- Slides: 86