Protein structure validation ECCB 2020 Gent Introduction to

Protein structure validation ECCB 2020 Gent Introduction to protein structure validation (and improvement) Gert Vriend

Protein structure validation The plan for today: Gert Vriend: Robbie Joosten: Jurgen Doreleijers: Bas Vroling (and you): All (and you): Split-up in groups: At the end: Introduction to validation X-ray structure validation and improvement NMR structure validation (and improvement ) YASARA General validation practicals General validation issues X-ray specific issues NMR specific issues Continuation of validation practicals Overview of validation and related facilities And in-between we have coffee, lunch, tea, and whatever else they throw at us at any moment that anybody feels like it.

Structure validation Everything that can go wrong, will go wrong, especially with things as complicated as protein structures.

What is real?
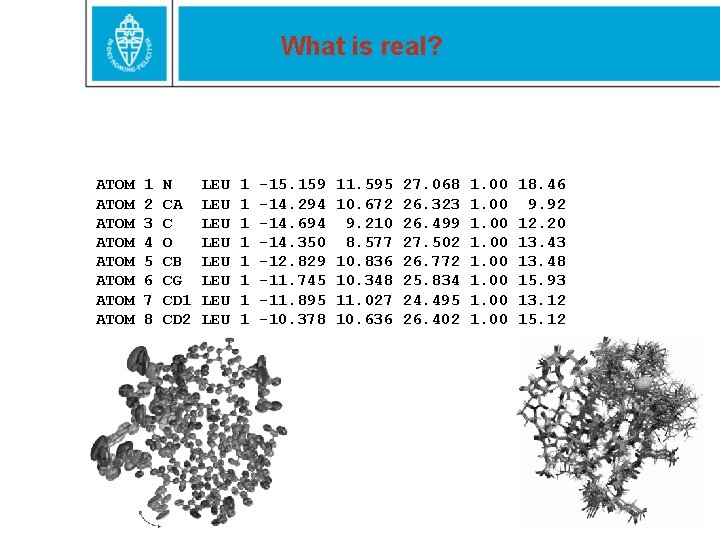
What is real? ATOM ATOM 1 2 3 4 5 6 7 8 N CA C O CB CG CD 1 CD 2 LEU LEU 1 1 1 1 -15. 159 -14. 294 -14. 694 -14. 350 -12. 829 -11. 745 -11. 895 -10. 378 11. 595 10. 672 9. 210 8. 577 10. 836 10. 348 11. 027 10. 636 27. 068 26. 323 26. 499 27. 502 26. 772 25. 834 24. 495 26. 402 1. 00 18. 46 9. 92 12. 20 13. 43 13. 48 15. 93 13. 12 15. 12
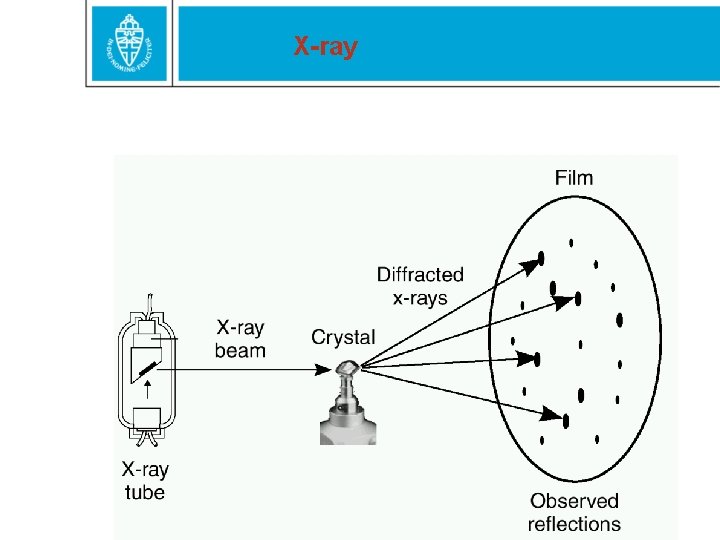
X-ray
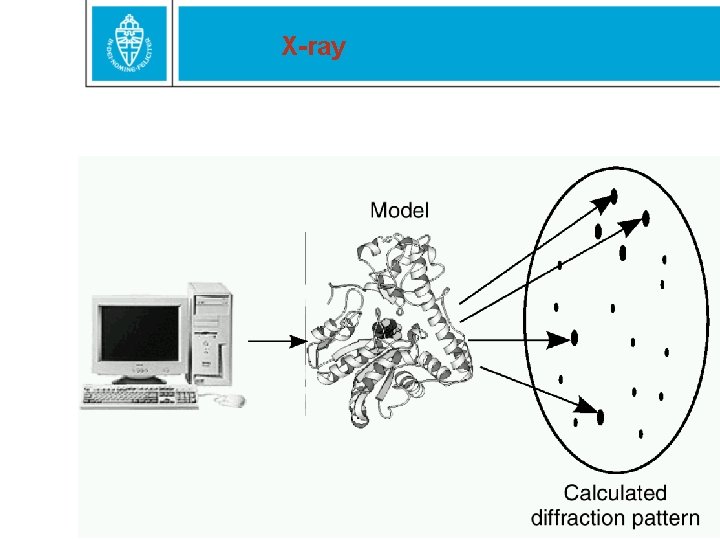
X-ray
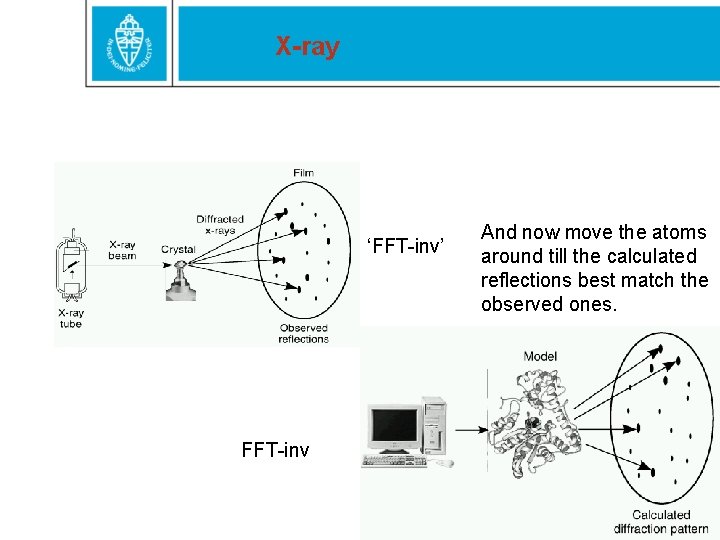
X-ray ‘FFT-inv’ FFT-inv And now move the atoms around till the calculated reflections best match the observed ones.
X-ray refinement / multiple minima Multiple minima

X-ray R-factor 2 Error = Σ w. (obs-calc) R-factor = Σ w. |obs-calc|
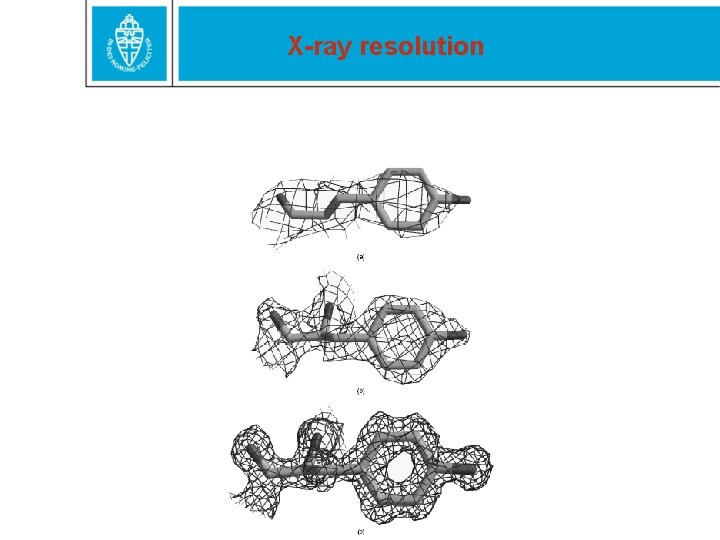
X-ray resolution
NMR data collection

NMR data consists of short inter-atomic distances between atoms. We call these NOEs. Most NOEs are between close neighbours in the sequence. Those hold little information. The ‘good’ NOEs are between atoms far away in the sequence. There are few of those, normally. NOEs are known with low precision. E. g. NOEs are binned 2. 5 -4. 0, 4. 0 -5. 5, and 5. 5 -7. 0. NMR can also measure some angles, and relative orientations. The latter, called RDCs are powerful.

NMR Q-factor Error = Σ NOE/RDC-violations + Energy term 2

NMR versus X-ray With X-ray you measure reflections. Each reflection holds information about each atom. With NMR you measure pair-wise distances, angles, and orientations. These all hold local information. X-ray requires crystals, and crystals cause/are artefacts. NMR is in solution, but provides much less precision.

NMR versus X-ray ‘Error’ Mobility Crystal artefacts Material needed Cost of hardware Drug design NMR 1 -2 Å yes no 20 mg 4 M Euro no X-ray 0. 1 -0. 5 Å not really yes 1 mg near infinite (share) almost Better combine and use the best of both worlds.
Why validation ? Why does a sane (? ) human being spend twenty years to search for millions of errors in the PDB?

Validation because: Everything we know about proteins comes from PDB files. Errors become less dangerous when you know about them. And, going back to the red thread through this series, if a template is wrong the model will be wrong.

What kind of errors can the software find? Administrative errors. Crystal-specific errors. NMR-specific errors. Really wrong things. Improbable things. Things worth looking at. Ad hoc things.
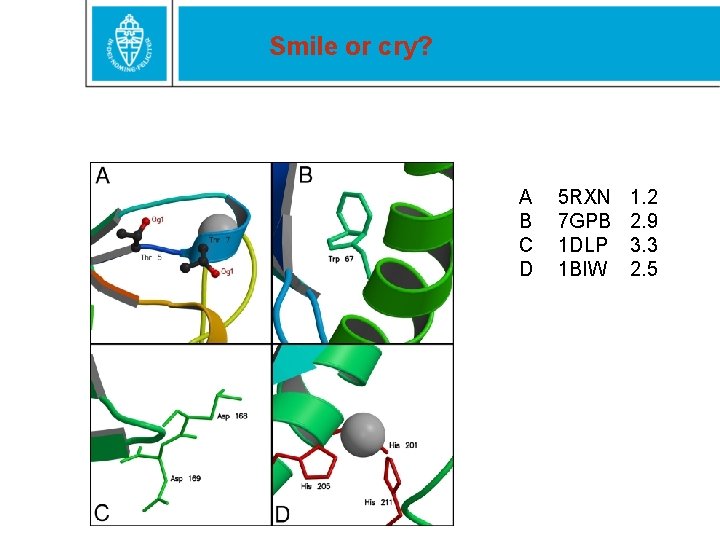
Smile or cry? A B C D 5 RXN 7 GPB 1 DLP 1 BIW 1. 2 2. 9 3. 3 2. 5

Why? Simple, proteins are very complex.

X-ray specific
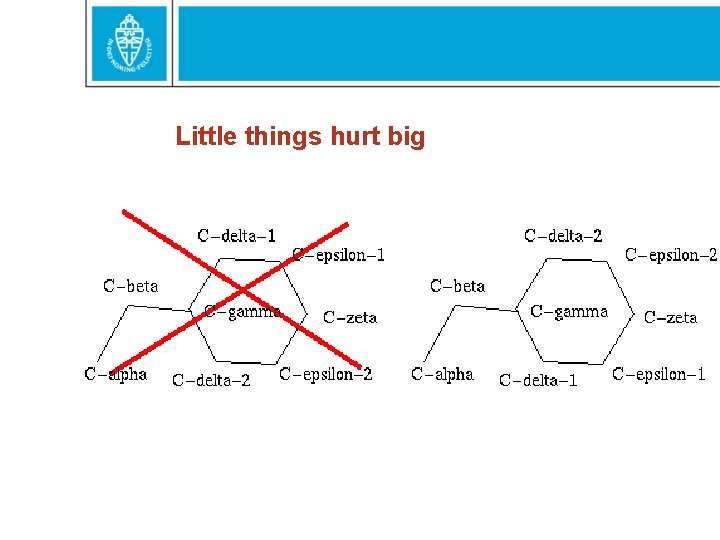
Little things hurt big
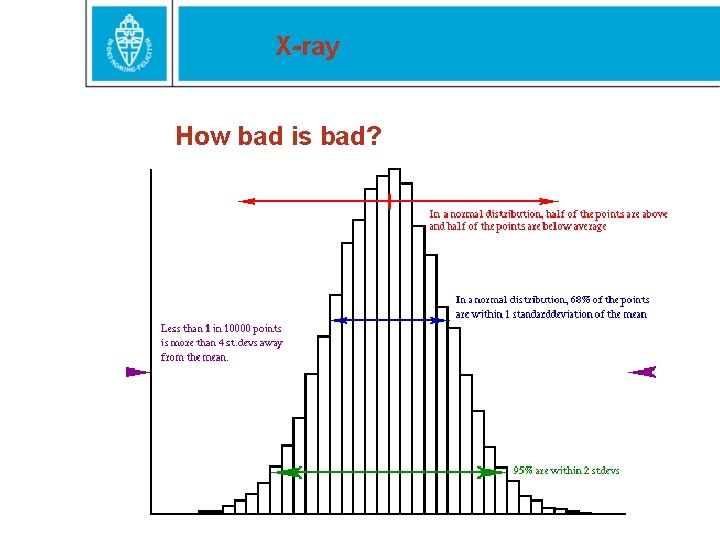
X-ray How bad is bad?

Check with force fields A force field is a set of parameters together with a set of rules to use those parameters to see how normal something is. Most force fields are designed to score events, or to predict the future. In structure validation we often look at structures and count ‘things’ For example, we count that the number of buried hydrogen bond donors that do not make a hydrogen bond is 4. 6+/-1. 2 per 100 amino acids in well-solved proteins. So we call that normal, and now, using ΔG=-RTln(K), we can calculate the energy penalty for proteins with more than 4. 6 unsatisfied buried unsatisfied hydrogen bonds.
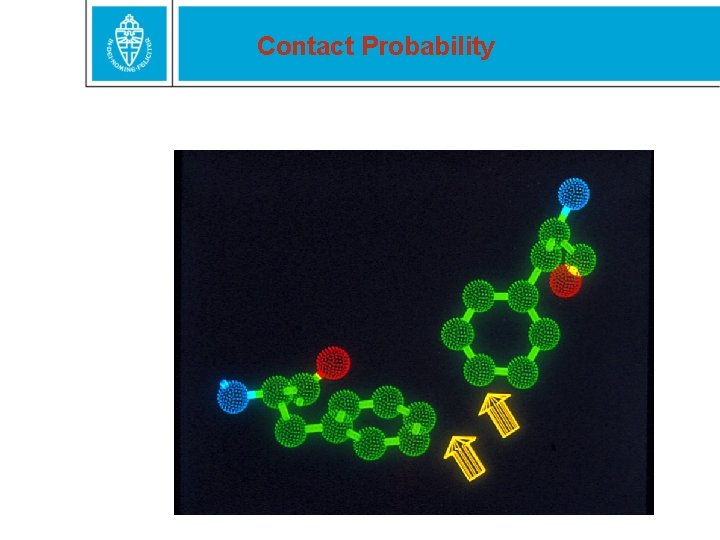
Contact Probability
Contact Probability

Contact probability box
One slide about homology modelling
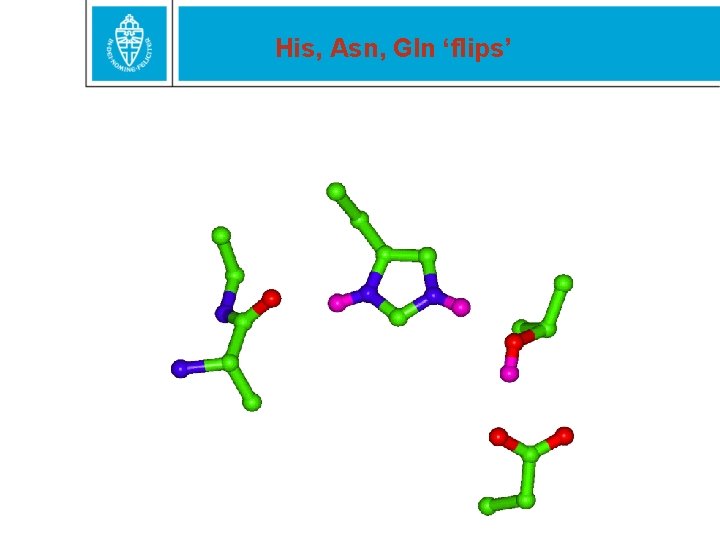
His, Asn, Gln ‘flips’
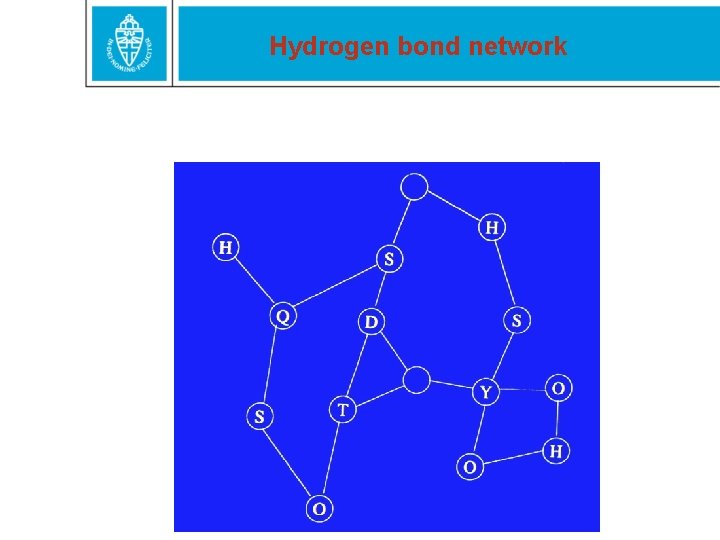
Hydrogen bond network
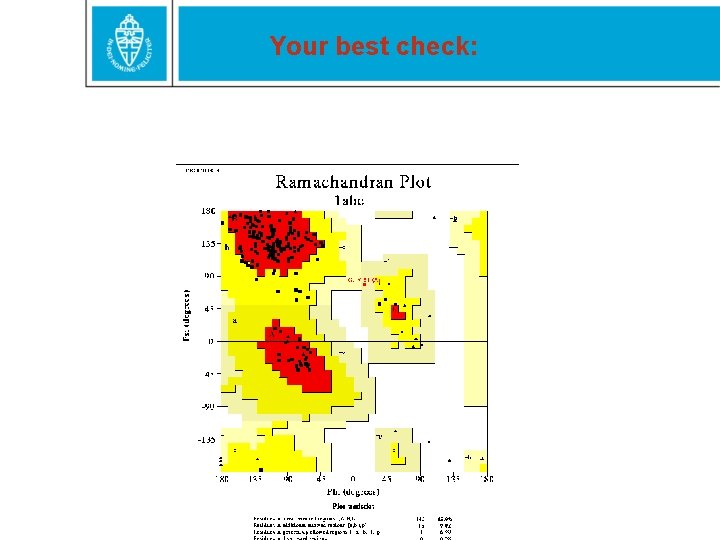
Your best check:
How difficult can it be? 1 CBQ 2. 2 A
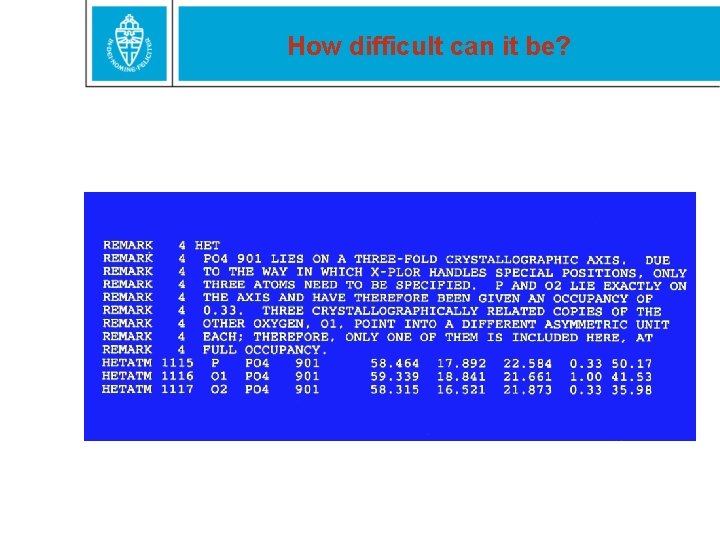
How difficult can it be?
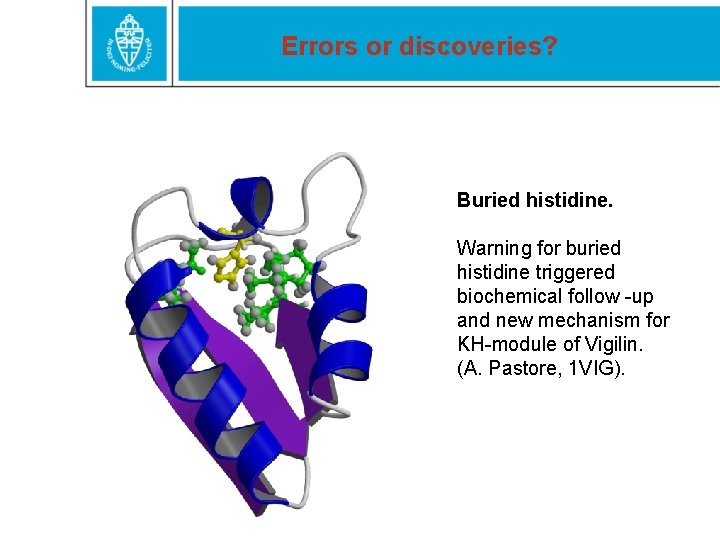
Errors or discoveries? Buried histidine. Warning for buried histidine triggered biochemical follow -up and new mechanism for KH-module of Vigilin. (A. Pastore, 1 VIG).

Acknowledgements: Rob Hooft Robbie Joosten Elmar Krieger Sander Nabuurs Chris Spronk Maarten Hekkelman
- Slides: 36