Protein Structure Prediction by Comparative Modelling Dr Abdelkrim

Protein Structure Prediction by Comparative Modelling Dr. Abdelkrim Rachedi, Dept. of Biology, Saida University

Comparative Modelling methods - Homology modelling. Targets that have homologous proteins with known structures in the PDB. - Threading or fold recognition. Targets that are estimated to have fold-level homology. The assumption is that the number of folds are limited in the known structures (PDB) Both are template-based methods.

Comparative Modelling basis - Proteins sharing a good level sequence similarity & share common ancestor "homology" would adopt similar structural fold. - In parctice: Comparative modelling approximates the 3 D structure of a target protein for which only the sequence is available. + Homology modelling: requires empirical 3 D "template" structure available with >25% sequence identity. + Threading: requires fold similarity and can be useful in cases of < 25% sequence identity.

Modelling Process Needs - The sequence of the protein with unknown 3 D structure, the "target sequence". - A 3 D template is chosen by virtue of having the highest sequence identity with the target sequence. - The 3 D structure of the template must be determined by reliable empirical methods such as crystallography or NMR, flawless, high resolution and is typically a published atomic coordinate "PDB" file from the Protein Data Bank.

Modelling Process Needs … Templates discovery: - Templates can be found using: - Sequence alignment methods such as FASTA or BLAST. (templates with low E-valued alignments are used) or - Protein threading / fold recognition methods. By placing amino acids in target sequence to positions in the template structure … using some scoring matrices. (better in finding distantly related templates. )

Modelling Major Step - Selection of a reliable template structure. (preliminary sequence alignment) - Sequence alignment of target sequence and template sequence. - Main chain atoms building. - Building loop regions. (database of turns & loops) - Building side chains - Energy minimization.

Template & seq. alignment
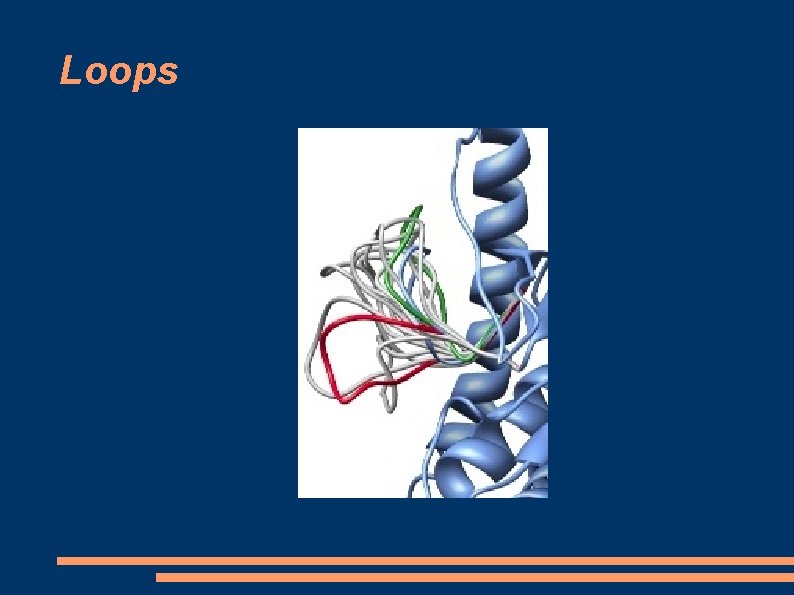
Loops
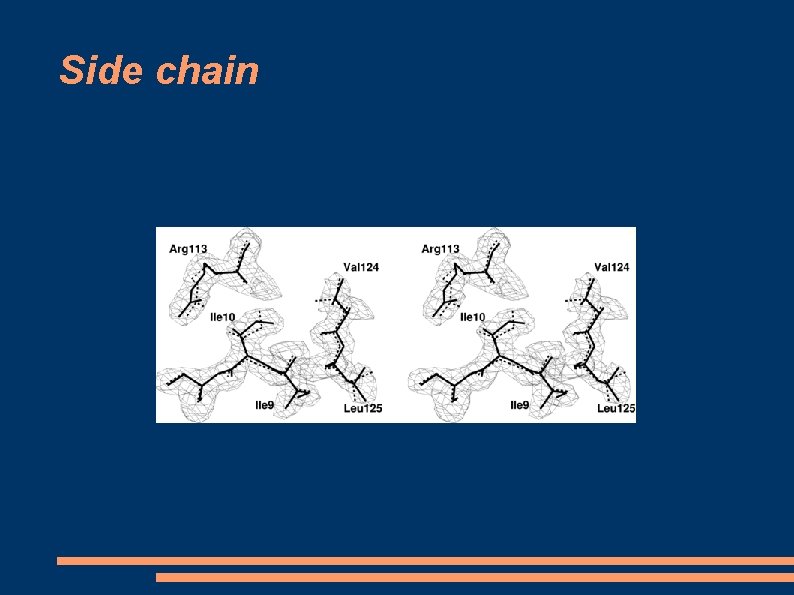
Side chain

Models Uses: - Conserved regions in the model can provide clues about putative active sites and ligand pokets. - Models can guide mutagenesis experiments. - Models hypotheses about structure-function relationships.

Models drawbacks - Models are unreliable in predicting the conformations of loop regions (insertions or deletions) This simply because loops are parts of the target sequence that don't align with the sequence of the template. - Models do not provide reliable side-chain positions. - Models are unlikely to be useful in modelling ligand docking (drug design) unless the sequence identity with the template is >70%, and even then, less reliable than an empirical crystallographic or NMR structure.

Online Modelling tools - Swiss-Model at http: //www. expasy. ch/swissmod/SWISSMODEL. html - The Protein Model Portal (PMP) at http: //www. proteinmodelportal. org/.

Models Databases Note: Because of the general unreliability of models, This category of “structures" has been removed from the PDB. - Mod. Base (Andrej Sali et al. , Rockefeller U, NY) at http: //pipe. rockefeller. edu/modbase - 3 DCrunch (Manuel Peitsch et al. , Glaxo. Wellcome) at http: //www. expasy. ch/swissmod/SM_3 DCrunch. html

References # D. Baker, A. Sali. Protein structure prediction and structural genomics. Science 294(5540): 93 -6 (2001). # C. Chothia, A. M. Lesk. The relation between the divergence of sequence and structure in proteins. EMBO J. 5: 823 -826 (1986). # D. Vitkup, E. Melamud, J. Moult, and C. Sander. Completeness in structural genomics. Nature Structal Biology 8: 482 -4 (2001).
- Slides: 14