Protein phosphorylation identification and new technologies for quantitative












- Slides: 28

Protein phosphorylation – identification and new technologies for quantitative analysis 1. Detection, identification, and mapping of phosphoproteins 2. New methods for quantitative analysis of protein phosphorylation by mass spectrometry 1. SILAC 2. AQUA peptides 3. Monitoring protein phosphorylation by Bio-Plex 1

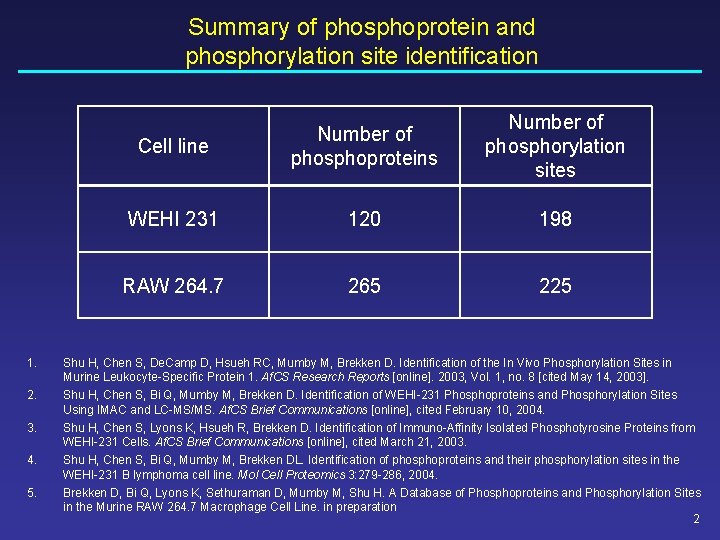
Summary of phosphoprotein and phosphorylation site identification 1. 2. 3. 4. 5. Cell line Number of phosphoproteins Number of phosphorylation sites WEHI 231 120 198 RAW 264. 7 265 225 Shu H, Chen S, De. Camp D, Hsueh RC, Mumby M, Brekken D. Identification of the In Vivo Phosphorylation Sites in Murine Leukocyte-Specific Protein 1. Af. CS Research Reports [online]. 2003, Vol. 1, no. 8 [cited May 14, 2003]. Shu H, Chen S, Bi Q, Mumby M, Brekken D. Identification of WEHI-231 Phosphoproteins and Phosphorylation Sites Using IMAC and LC-MS/MS. Af. CS Brief Communications [online], cited February 10, 2004. Shu H, Chen S, Lyons K, Hsueh R, Brekken D. Identification of Immuno-Affinity Isolated Phosphotyrosine Proteins from WEHI-231 Cells. Af. CS Brief Communications [online], cited March 21, 2003. Shu H, Chen S, Bi Q, Mumby M, Brekken DL. Identification of phosphoproteins and their phosphorylation sites in the WEHI-231 B lymphoma cell line. Mol Cell Proteomics 3: 279 -286, 2004. Brekken D, Bi Q, Lyons K, Sethuraman D, Mumby M, Shu H. A Database of Phosphoproteins and Phosphorylation Sites in the Murine RAW 264. 7 Macrophage Cell Line. in preparation 2

Some phosphotyrosine proteins identified in pervanadate-treated RAW cells Btk Dok 1 Fgr Dok 2 Lck PLC 2 Lyn SHIP FcεRI Shp 1 Fyn Shp 2 Fyb/SLAP 130 SHPS-1 Hematopoietic cell specific Lyn substrate/HS 1 SLP-76 Immunoglobulin superfamily member 4 3

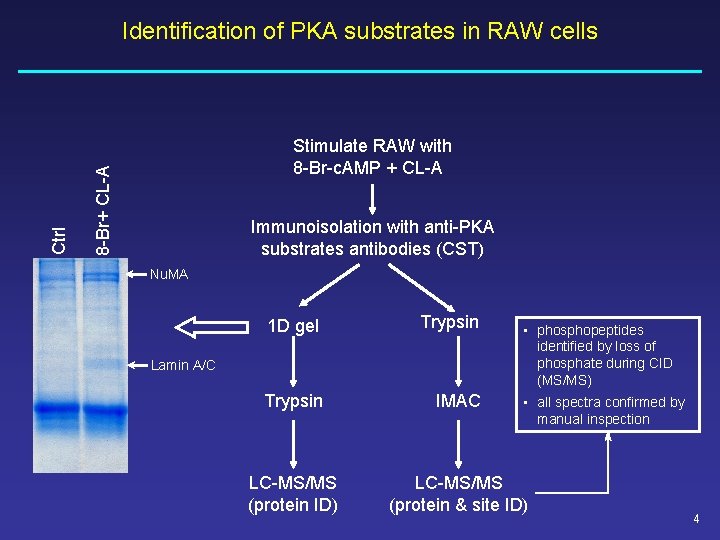
Stimulate RAW with 8 -Br-c. AMP + CL-A 8 -Br+ CL-A Ctrl Identification of PKA substrates in RAW cells Immunoisolation with anti-PKA substrates antibodies (CST) Nu. MA 1 D gel Trypsin IMAC LC-MS/MS (protein ID) LC-MS/MS (protein & site ID) Lamin A/C • phosphopeptides identified by loss of phosphate during CID (MS/MS) • all spectra confirmed by manual inspection 4

Phosphoproteins identified in RAW cells treated with 8 -Br-c. AMP (peptide ID’d is underlined) Contains predicted PKA site Acyl-Coenzyme A binding domain containing 4 PQPLKQRS*PRRTR Yes (T 103) DNA methyltransferase Mmu. I KLESHT*VPVQSR Yes (S 254, T 324) DNA methyltransferase Mmu. I VPALAS*PAGSLPDHVR Yes (S 254, T 324) Heterogeneous nuclear ribonucleoprotein U AAGKSS*GPTSLFAVTVAPPGAR Yes (S 26) Lamin A RSFRS*VGGSGGGSFGDNLVTR Yes (T 10) Lamin A/C LRLS*PSPT*SQR Yes 1 (T 45, S 278) Lamin C SGAQASSTPLS*PTRITRL Yes (T 10) Mitochondrial Solute Carrier protein RDFY*WLR No Nucleolin ALVPT*PGKK Yes (T 57) Raly protein (an hn. RNP) GRLS*PVPVPR Yes (S 119, T 135) Ribosomal protein L 7 VATVPGT*LKKKVP No Vasodilator-stimulated phosphoprotein (VASP) KLRKVS*KQEEASGGPLAPK Yes 1 (S 157, S 239, T 274) Protein name 1 known PKA substrate Peptide sequence 5

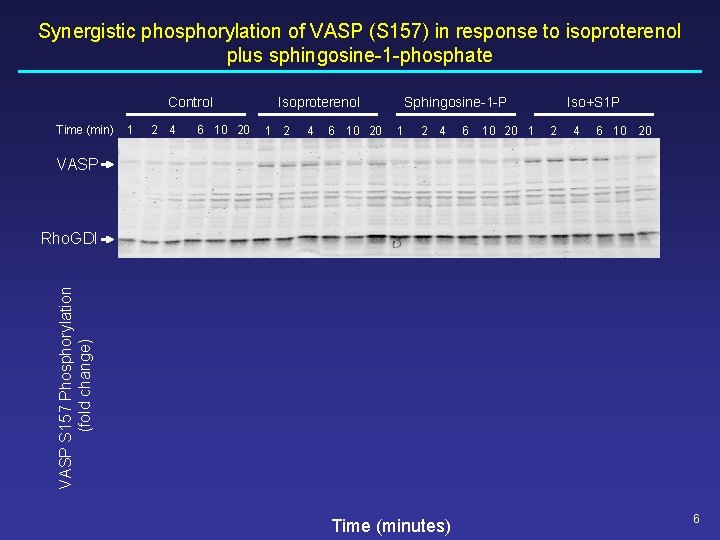
Synergistic phosphorylation of VASP (S 157) in response to isoproterenol plus sphingosine-1 -phosphate Control Time (min) 1 2 4 6 10 20 Isoproterenol 1 2 4 6 10 20 Sphingosine-1 -P 1 2 4 6 10 20 1 Iso+S 1 P 2 4 6 10 20 VASP S 157 Phosphorylation (fold change) Rho. GDI Time (minutes) 6

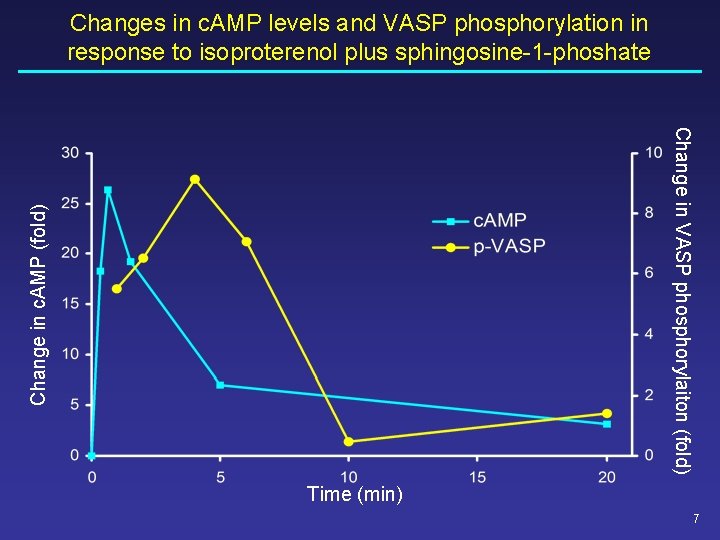
Changes in c. AMP levels and VASP phosphorylation in response to isoproterenol plus sphingosine-1 -phoshate Change in c. AMP (fold) Change in VASP phosphorylaiton (fold) Time (min) 7

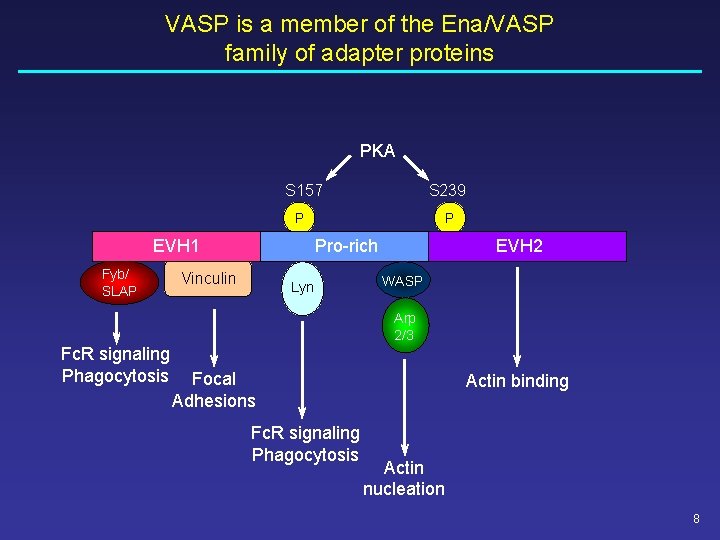
VASP is a member of the Ena/VASP family of adapter proteins PKA S 157 S 239 P EVH 1 Fyb/ SLAP P Pro-rich Vinculin Lyn EVH 2 WASP Arp 2/3 Fc. R signaling Phagocytosis Focal Adhesions Fc. R signaling Phagocytosis Actin binding Actin nucleation 8

New methods for quantitative analysis of phosphorylation of FXM proteins • Phosphopeptides are usually difficult to detect by mass spectrometry • To “hedge our bets”, characterization and validation experiments have utilized RAW cells expressing tagged-FXM proteins • Stable RAW cell populations expressing FLAG-tagged FXM proteins produced via retrovirus transduction and drug selection • Stable cell populations treated with ligands or phosphatase inhibitors • Proteins immunoprecipitated with anti-FLAG antibody • Analyzed by immunoblotting and mass spectrometry Flag-Akt 1 Grb 2 -Flag FTM-Erk 1 Flag-Grk 2 Flag-SHP 2 Flag-Btk Flag-Syk 9

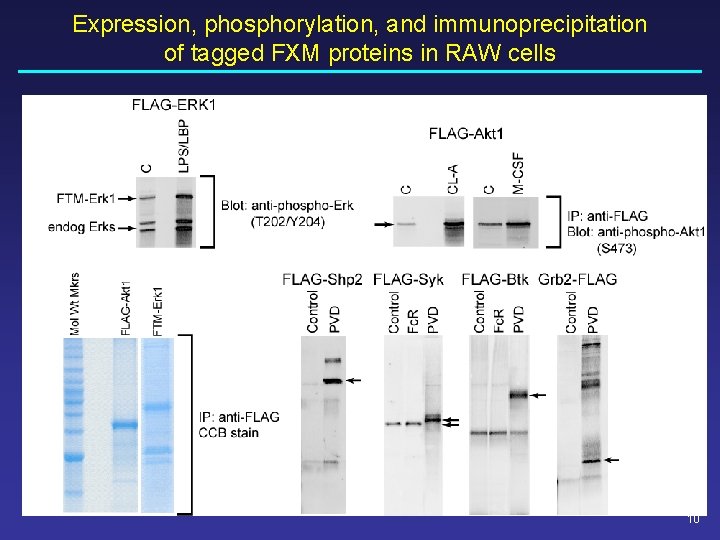
Expression, phosphorylation, and immunoprecipitation of tagged FXM proteins in RAW cells 10

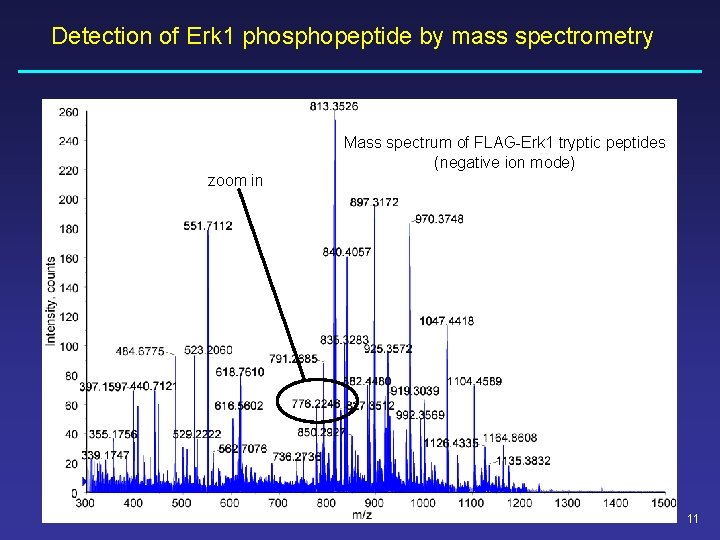
Detection of Erk 1 phosphopeptide by mass spectrometry Mass spectrum of FLAG-Erk 1 tryptic peptides (negative ion mode) zoom in 11

Detection of the Erk 1 phosphopeptides in RAW cells stimulated with LPS doubly phosphorylated peptide 3 IADPEHDHTGFLT*EY*VATR singly phosphorylated peptide 3 - 12

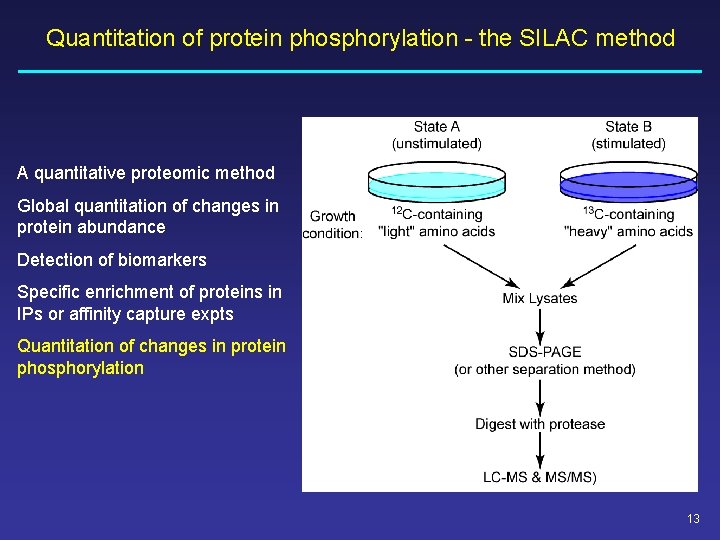
Quantitation of protein phosphorylation - the SILAC method A quantitative proteomic method Global quantitation of changes in protein abundance Detection of biomarkers Specific enrichment of proteins in IPs or affinity capture expts Quantitation of changes in protein phosphorylation 13

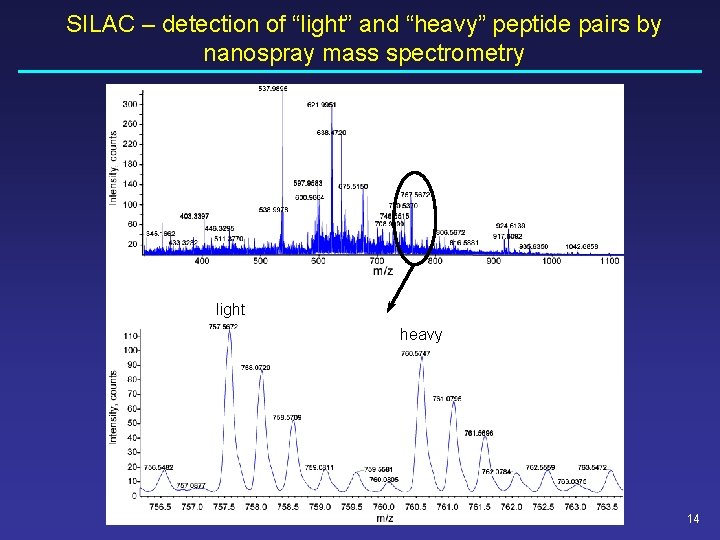
SILAC – detection of “light” and “heavy” peptide pairs by nanospray mass spectrometry light heavy 14

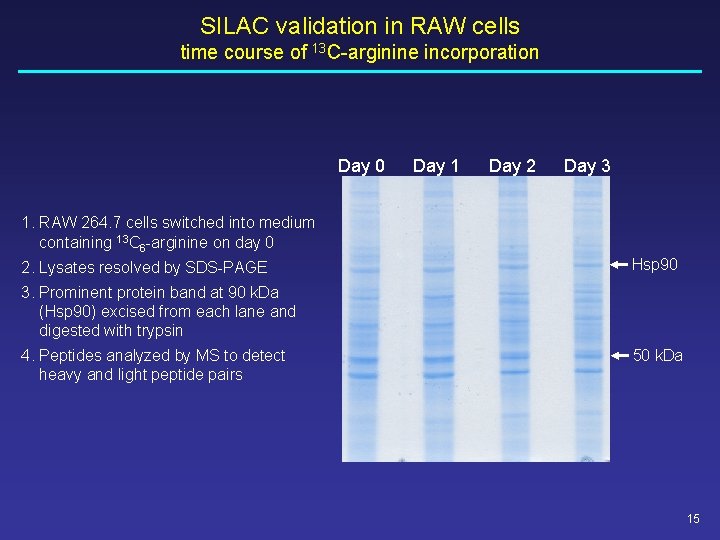
SILAC validation in RAW cells time course of 13 C-arginine incorporation Day 0 Day 1 Day 2 Day 3 1. RAW 264. 7 cells switched into medium containing 13 C 6 -arginine on day 0 2. Lysates resolved by SDS-PAGE Hsp 90 3. Prominent protein band at 90 k. Da (Hsp 90) excised from each lane and digested with trypsin 4. Peptides analyzed by MS to detect heavy and light peptide pairs 50 k. Da 15

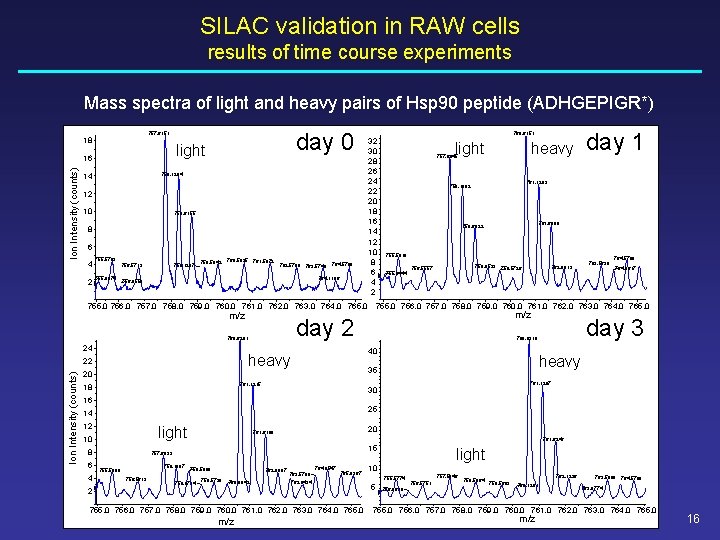
SILAC validation in RAW cells results of time course experiments Mass spectra of light and heavy pairs of Hsp 90 peptide (ADHGEPIGR*) light 16 Ion Intensity (counts) day 0 757. 6151 18 758. 1204 14 12 10 758. 6155 8 6 4 2 755. 5783 756. 5713 755. 0874 759. 1287 759. 5942 760. 5935 761. 5872 762. 5709 763. 5749 764. 5780 764. 1196 756. 0558 32 30 28 26 24 22 20 18 16 14 12 10 8 6 4 2 760. 6151 light heavy 757. 6045 day 1 761. 1203 758. 1092 761. 6098 758. 6022 755. 5810 755. 0444 759. 0533 759. 5726 756. 5557 762. 0913 763. 5638 764. 5780 764. 0617 755. 0 756. 0 757. 0 758. 0 759. 0 760. 0 761. 0 762. 0 763. 0 764. 0 765. 0 m/z day 2 760. 6201 24 Ion Intensity (counts) 761. 1215 18 heavy 35 20 day 3 760. 6319 40 heavy 22 m/z 761. 1297 30 16 25 14 12 light 10 8 6 4 20 761. 6160 761. 6246 15 757. 6033 758. 1087 755. 5809 756. 5613 758. 5869 759. 0714 762. 0987 759. 5730 760. 0842 764. 0547 763. 5799 765. 0367 763. 0434 2 755. 0 756. 0 757. 0 758. 0 759. 0 760. 0 761. 0 762. 0 763. 0 764. 0 765. 0 m/z light 10 755. 5774 5 756. 0610 757. 5946 756. 5751 758. 5864 762. 1236 759. 5683 760. 1281 763. 5809 764. 5798 763. 0774 755. 0 756. 0 757. 0 758. 0 759. 0 760. 0 761. 0 762. 0 763. 0 764. 0 765. 0 m/z 16

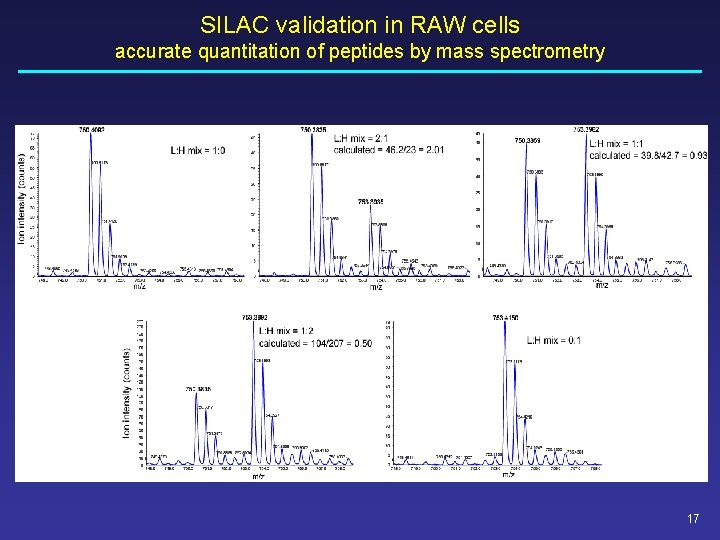
SILAC validation in RAW cells accurate quantitation of peptides by mass spectrometry 17

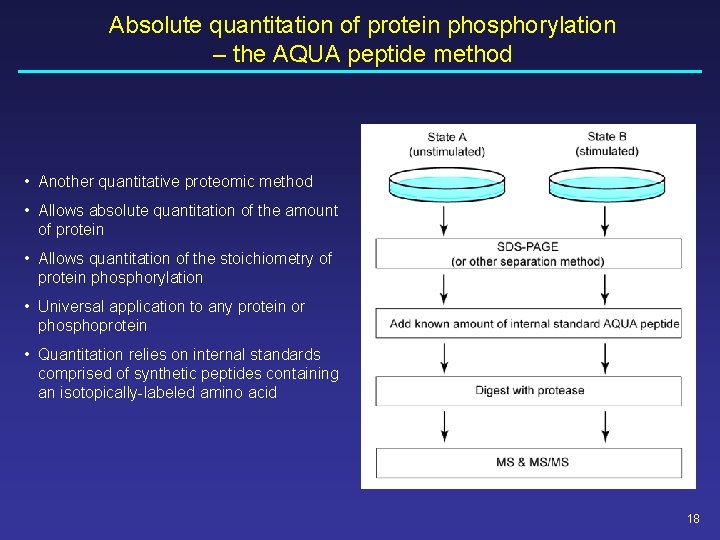
Absolute quantitation of protein phosphorylation – the AQUA peptide method • Another quantitative proteomic method • Allows absolute quantitation of the amount of protein • Allows quantitation of the stoichiometry of protein phosphorylation • Universal application to any protein or phosphoprotein • Quantitation relies on internal standards comprised of synthetic peptides containing an isotopically-labeled amino acid 18

AQUA peptide design for FLAG-Akt 1 (Ser 473) FLAG + att. B 1 sequence MDYKDDDDKGAGAGSSSGHQTSLYKKAGSTMNDVAIVKEGWLHKRGE YIKTWRPRYFLLKNDGTFIGYKERPQDVDQRESPLNNFSVAQCQLMK TERPRPNTFIIRCLQWTTVIERTFHVETPEEREEWATAIQTVADGLK RQEEETMDFRSGSPSDNSGAEEMEVSLAKPKHRVTMNEFEYLKLLGK GTFGKVILVKEKATGRYYAMKILKKEVIVAKDEVAHTLTENRVLQNS RHPFLTALKYSFQTHDRLCFVMEYANGGELFFHLSRERVFSEDRARF YGAEIVSALDYLHSEKNVVYRDLKLENLMLDKDGHIKITDFGLCKEG IKDGATMKTFCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGR LPFYNQDHEKLFELILMEEIRFPRTLGPEAKSLLSGLLKKDPTQRLG GGSEDAKEIMQHRFFANIVWQDVYEKKLSPPFKPQVTSETDTRYFDE EFTAQMITITPPDQDDSMECVDSERRPHFPQFSYSASGTA Ser 473 tryptic peptide endogenous phosphopeptide RPHFPQFS*YSASGTA + PO 3 m/z of M 2 - = 865 synthetic “AQUA” phosphopeptide RPHFPQF(13 C 915 N 1)S*YSASGTA + PO 3 m/z of M 2 - = 870 19

AQUA method – mass spectrum of FLAG-Akt 1 from calyculin-A treated RAW cells Zoom in on this area 20

AQUA peptide – FLAG-Akt 1 (Ser 473) zoomed scan Endogenous phosphopeptide 13 C, 15 N-labeled AQUA phosphopeptide 21

Planned uses of SILAC and AQUA methods • • • SILAC – in a targeted way to quantitate phosphorylation of specific proteins (e. g. , FXM) – in a global way to identify known and novel proteins whose phosphorylation is altered by ligands/perturbations – coupled with antibody and IMAC enrichment methods AQUA – in a targeted manner to quantitate phosphorylation of FXM proteins where a suitable phospho-specific antibody is not available – quantitate the absolute amounts of phosphoproteins and the stoichiometry of phosphorylation in response to stimuli/perturbations Current efforts – finish implementing both methods – increase sensitivity – selected reaction monitoring (20 X); new instrumentation (50 X) 22

The Af. CS Protein Chemistry Lab Deirdre Brekken Kathy Lyons Lead Scientist Farah El Mazouni Deepa Sethuraman Hongjun Laura Draper Robert Cox Qun Bi. Shu Lead Scientist 23

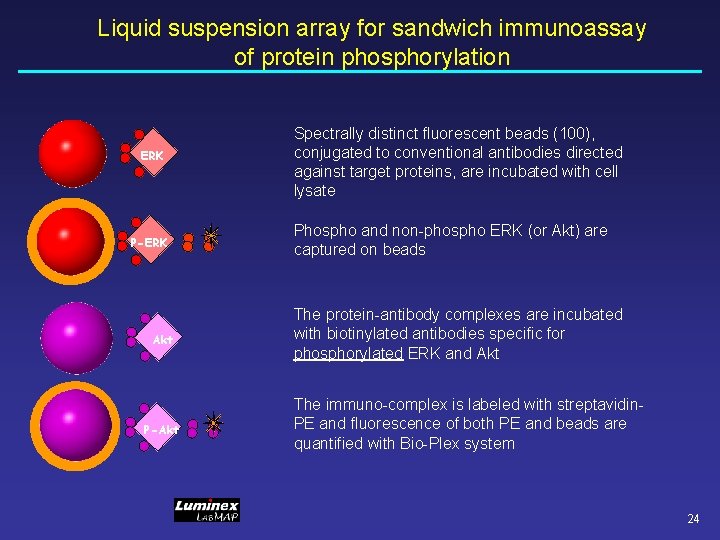
Liquid suspension array for sandwich immunoassay of protein phosphorylation ERK P-ERK Akt P-Akt Spectrally distinct fluorescent beads (100), conjugated to conventional antibodies directed against target proteins, are incubated with cell lysate Phospho and non-phospho ERK (or Akt) are captured on beads The protein-antibody complexes are incubated with biotinylated antibodies specific for phosphorylated ERK and Akt The immuno-complex is labeled with streptavidin. PE and fluorescence of both PE and beads are quantified with Bio-Plex system 24

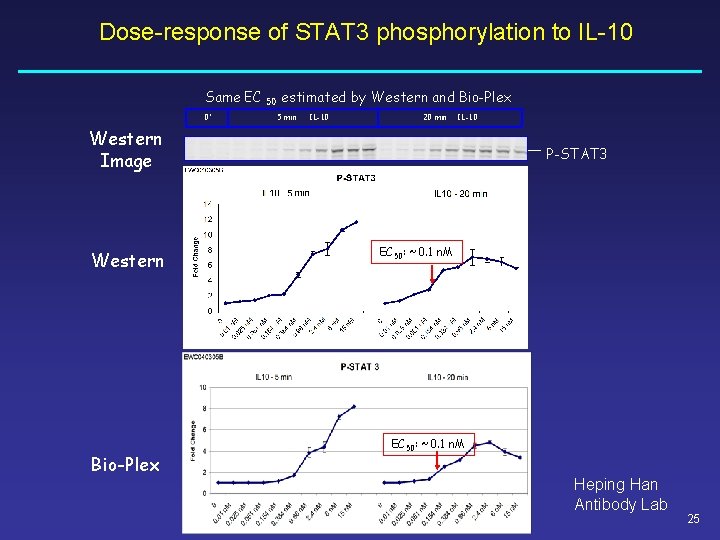
Dose-response of STAT 3 phosphorylation to IL-10 Same EC 0’ 50 estimated by Western and Bio-Plex 5 min IL-10 20 min IL-10 Western Image Western Bio-Plex P-STAT 3 EC 50: ~0. 1 n. M Heping Han Antibody Lab 25

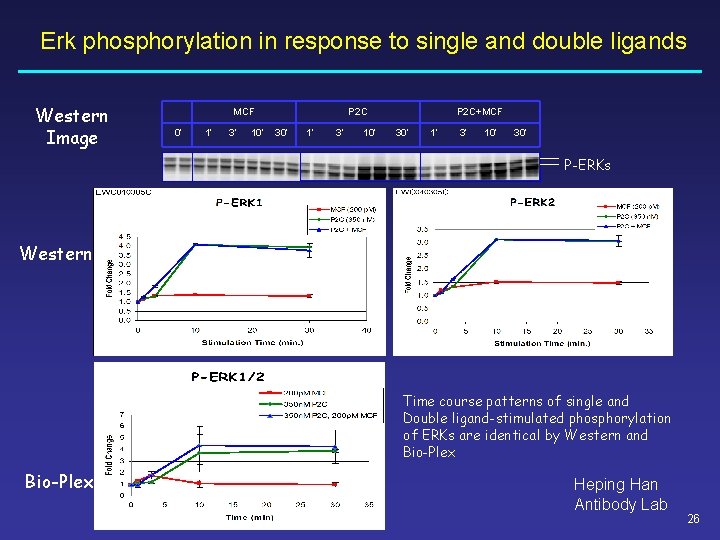
Erk phosphorylation in response to single and double ligands Western Image MCF 0’ 1’ 3’ 10’ P 2 C 30’ 1’ 3’ 10’ P 2 C+MCF 30’ 1’ 3’ 10’ 30’ P-ERKs Western Time course patterns of single and Double ligand-stimulated phosphorylation of ERKs are identical by Western and Bio-Plex Heping Han Antibody Lab 26

Summary comparison of Bio-Plex and Western Blotting for quantifying ligand-induced phosphorylation of ATF-2, ERKs, and STAT 3 Conclusions: • Bio-Plex results are VERY similar to blotting results • Bio-Plex has the potential to triple or quadruple the number of phosphoproteins currently monitored • Bio-Plex has the potential to massively increase throughput in screening phosphoproteins • More sets of antibodies for phosphoproteins are needed to help Af. CS (currently 11) 27

Bio-Plex Validation of the Bio-Plex assay for protein phosphorylation is an ongoing collaboration between The Af. CS Antibody Lab and Bio-Rad Laboratories and Cell Signaling Technology 28