Protein Overexpression in E coli and Purification by

Protein Overexpression in E. coli and Purification by Affinity Chromatography Prof. Seung-Taek Lee Teaching Assistant - So Yun Ahn - Min Ju Pyo
Introduction – The feature of E. coli expression system Protein overexpression system - E. coli Prokaryotic cell - Yeast - Insect cell Eukaryotic cell - mammalian - The expression of proteins in Escherichia coli(E. coli) is the easiest, quickest and cheapest method. - There are many commercial and non-commercial expression vectors available with different N- and C-terminal tags and many different strains which are optimized for special applications. - Many foreign proteins are expressed at high level. - However, there is no post-translational modification (e. g. , glycosylation or cleavage at specific site).
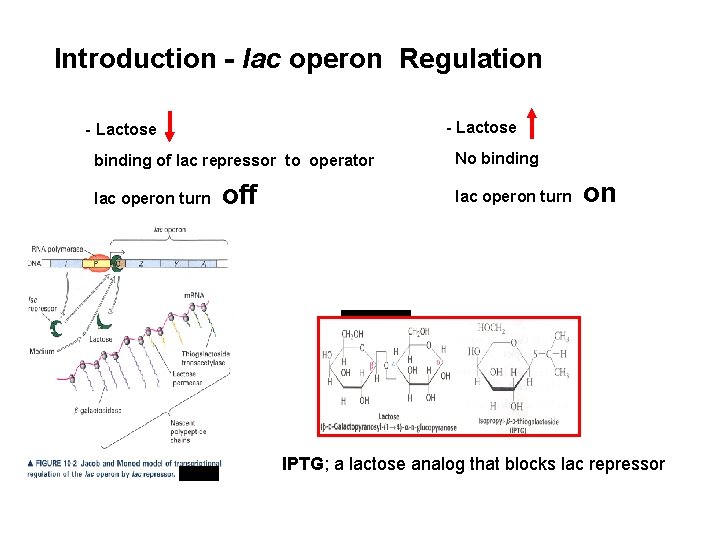
Introduction - lac operon Regulation - Lactose binding of lac repressor to operator No binding lac operon turn off lac operon turn on IPTG; a lactose analog that blocks lac repressor
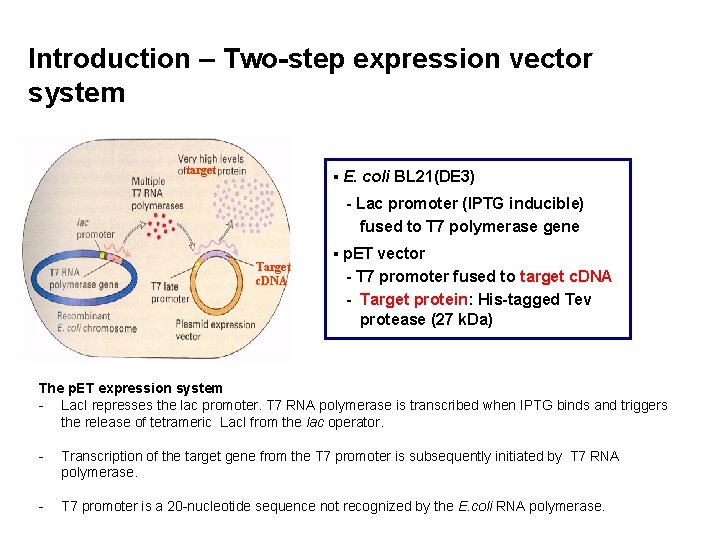
Introduction – Two-step expression vector system target ▪ E. coli BL 21(DE 3) - Lac promoter (IPTG inducible) fused to T 7 polymerase gene Target c. DNA ▪ p. ET vector - T 7 promoter fused to target c. DNA - Target protein: His-tagged Tev protease (27 k. Da) The p. ET expression system - Lac. I represses the lac promoter. T 7 RNA polymerase is transcribed when IPTG binds and triggers the release of tetrameric Lac. I from the lac operator. - Transcription of the target gene from the T 7 promoter is subsequently initiated by T 7 RNA polymerase. - T 7 promoter is a 20 -nucleotide sequence not recognized by the E. coli RNA polymerase.
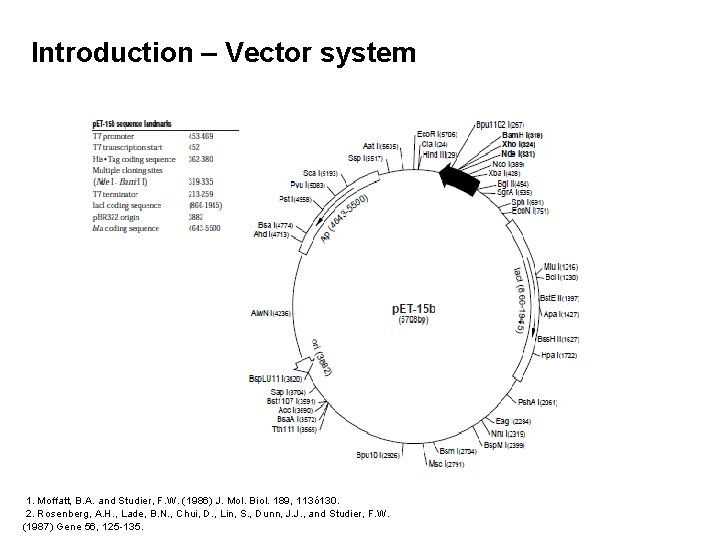
Introduction – Vector system 1. Moffatt, B. A. and Studier, F. W. (1986) J. Mol. Biol. 189, 113ó 130. 2. Rosenberg, A. H. , Lade, B. N. , Chui, D. , Lin, S. , Dunn, J. J. , and Studier, F. W. (1987) Gene 56, 125 -135.

Introduction – p. ET-15 b cloning / expression region
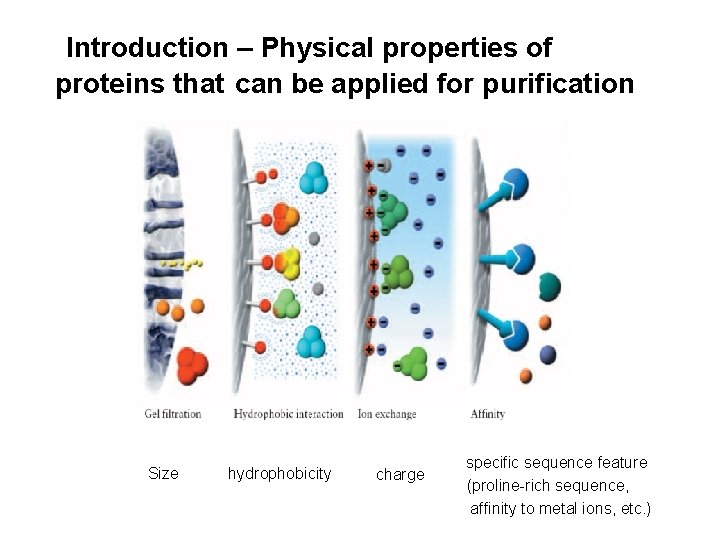
Introduction – Physical properties of proteins that can be applied for purification Size hydrophobicity charge specific sequence feature (proline-rich sequence, affinity to metal ions, etc. )

Introduction – Important matters to consider before starting the purification Sample and target protein properties Strategy Temperature stability p. H stability Use low temp. Selection of buffers Detergent requirement Consider choice of detergents Co-factor for stability or activity Select additives, salt, etc.
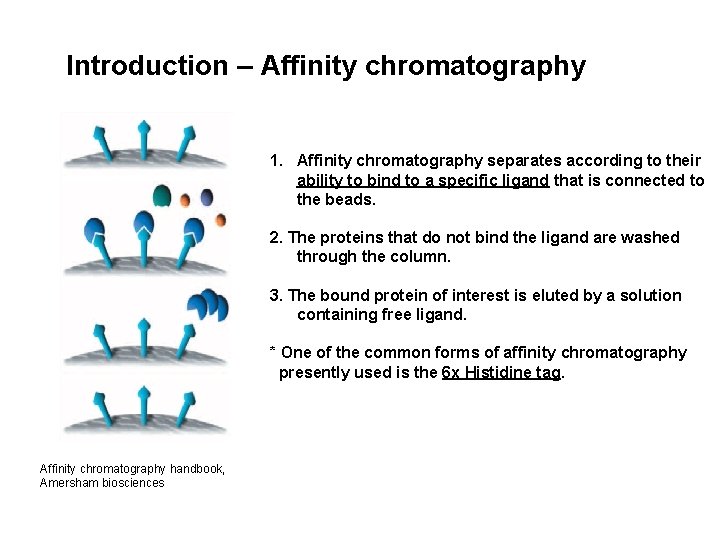
Introduction – Affinity chromatography 1. Affinity chromatography separates according to their ability to bind to a specific ligand that is connected to the beads. 2. The proteins that do not bind the ligand are washed through the column. 3. The bound protein of interest is eluted by a solution containing free ligand. * One of the common forms of affinity chromatography presently used is the 6 x Histidine tag. Affinity chromatography handbook, Amersham biosciences
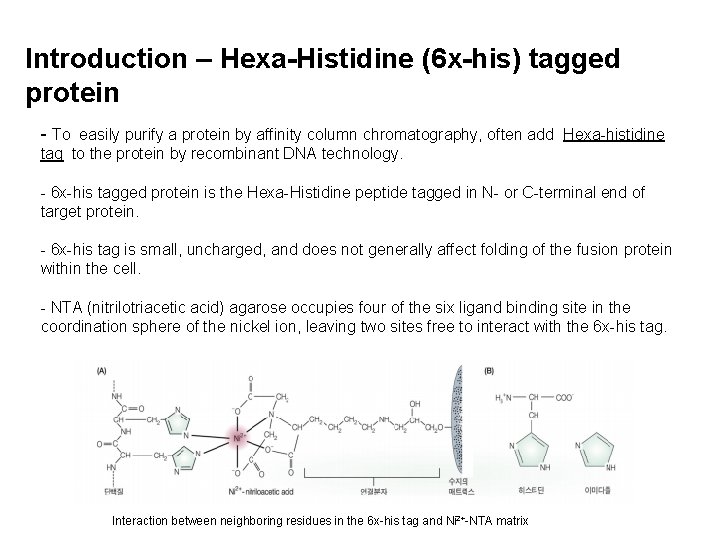
Introduction – Hexa-Histidine (6 x-his) tagged protein - To easily purify a protein by affinity column chromatography, often add Hexa-histidine tag to the protein by recombinant DNA technology. - 6 x-his tagged protein is the Hexa-Histidine peptide tagged in N- or C-terminal end of target protein. - 6 x-his tag is small, uncharged, and does not generally affect folding of the fusion protein within the cell. - NTA (nitrilotriacetic acid) agarose occupies four of the six ligand binding site in the coordination sphere of the nickel ion, leaving two sites free to interact with the 6 x-his tag. Interaction between neighboring residues in the 6 x-his tag and Ni 2+-NTA matrix
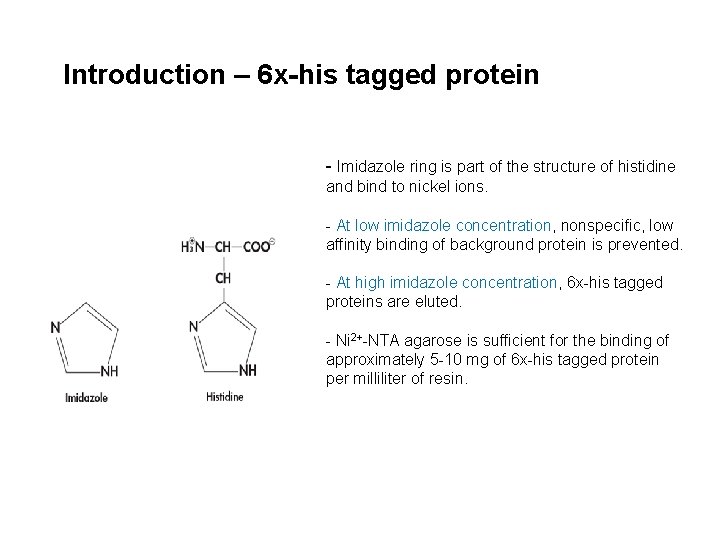
Introduction – 6 x-his tagged protein - Imidazole ring is part of the structure of histidine and bind to nickel ions. - At low imidazole concentration, nonspecific, low affinity binding of background protein is prevented. - At high imidazole concentration, 6 x-his tagged proteins are eluted. - Ni 2+-NTA agarose is sufficient for the binding of approximately 5 -10 mg of 6 x-his tagged protein per milliliter of resin.
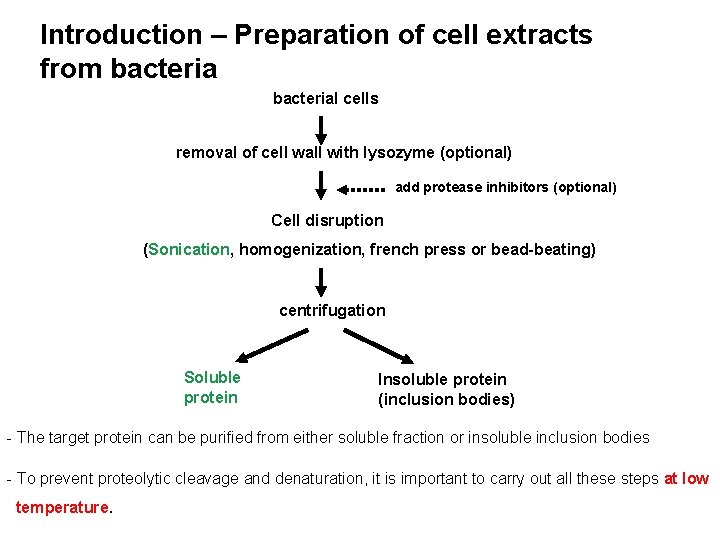
Introduction – Preparation of cell extracts from bacterial cells removal of cell wall with lysozyme (optional) add protease inhibitors (optional) Cell disruption (Sonication, homogenization, french press or bead-beating) centrifugation Soluble protein Insoluble protein (inclusion bodies) - The target protein can be purified from either soluble fraction or insoluble inclusion bodies - To prevent proteolytic cleavage and denaturation, it is important to carry out all these steps at low temperature.
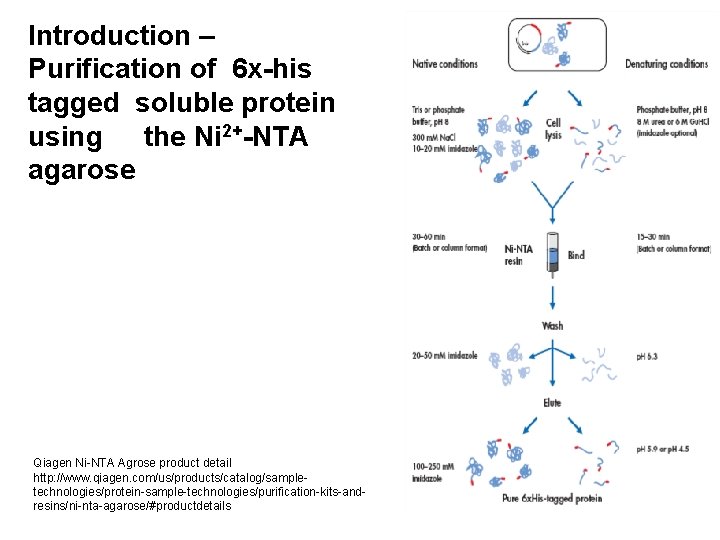
Introduction – Purification of 6 x-his tagged soluble protein using the Ni 2+-NTA agarose Qiagen Ni-NTA Agrose product detail http: //www. qiagen. com/us/products/catalog/sampletechnologies/protein-sample-technologies/purification-kits-andresins/ni-nta-agarose/#productdetails

Procedure < Day 1 > 1. 15 ㎖ tube에 LB media 3 ㎖을 넣고 autoclave하여 보관한다. 2. 15 ㎖ tube에 LB media 3 ㎖, ampicillin(100 ㎎/㎖) 3 ㎕를 넣고 여기에 E. coli colony를 seeding한다. 3. E. coli를 37 ℃, 200 rpm으로 맞추어진 shaking incubator에서 overnight으로 키운다.

Procedure < Day 2 > 1. E. coli 가 자란 것을 확인한다. 2. autoclave해둔 10 ㎖ LB media에 ampicillin (100 ㎎/㎖) 10 ㎕를 가하고, overnight으로 배양한 E. coli 100 ㎕ (1%)을 inoculation 한다. 3. spectrophotometer로 600 ㎚에서 흡광도를 측정하여 O. D. 값이 0. 5가 될 때까지 키운다. 4. O. D. 가 0. 5가 되면 1 M IPTG 10 ㎕를 tube에 넣어주어 25℃, 100 rpm으로 8시간 동안 더 키워서 target 단백질을 induction한다. 5. 50 ㎖ tube를 4, 000 rpm, 10 min centrifuge하여 E. coli를 모은다.

Procedure 1. Resuspending of cells cell pellet에 10 m. M imidazole을 포함하는 lysis buffer를 cell culture 부피의 1 / 10 을 가해 resuspend 한다. 10 ㎖ culture한 cell pellet이므로 1 ㎖로 resuspend함. (lysis buffer 조성: 25 m. M Sodium Phosphate (p. H 7. 5), 100 m. M Na. Cl , 2 m. M βmercaptoethanol) 2. Lysis of cells resuspending 한 sample을 microtube에 옮긴다. 이 시료를 sonication으로 cell을 파쇄한다. (30 sec burst, 90 sec cooling, 5 -8 times) - 주의! sonication 시, 거품이 나지 않도록 rod가 sample의 중간에 오도록 한다. 3. Separation of soluble proteins microtube centrifuge로 14, 000 rpm, 4℃, 10 min centrifuge 하여 supernatant를 얻는다. (Input 시료 : Supernatant 중 20 ㎕를 취하여 5 X sample buffer 5 ㎕ 가 들어있는 tube 에 넣는다. )

Procedure 4. Ni 2+- NTA agarose affinity column chromatography <각 단계 buffer 조성> binding buffer : 10 m. M imidazole이 포함된 lysis buffer (p. H 7. 5) washing buffer : 30 m. M imidazole이 포함된 lysis buffer (p. H 7. 5) elution buffer : 500 m. M imidazole이 포함된 lysis buffer (p. H 7. 5) 1) 20% ethanol에 equilibrium 되어있는 resin (200 ㎕)-> slurry로 400 ㎕ 흔들어준 후 column에 붓는다. 2) resin이 가라앉으면 D. W를 500 ㎕ 씩 2번 흘려주어 column bed 바로 위까지 비운다. 이때 resin이 마르지 않도록 주의한다. 3) bed volume(200 ㎕)의 3배의 binding buffer를 600 ㎕ 흘려준다. 4) Soluble protein이 들어있는 Sup을 loading하고, flow through를 받는다. F. T. 를 다시 loading하고 F. T. 를 받는 과 정을 4회 반복한다. (최종적으로 받은 F. T. 는 총 5회의 F. T. 를 진행한 것이어야 함. ) ( FT 시료 : 최종 flow-through 중 20 ㎕을 취하여 5 X sample buffer 5 ㎕가 들어있는 tube에 넣는다. ) 5) bed volume(200 ㎕) 9배의 washing buffer를 600 ㎕씩 3번 흘려준다. (3배의 Bed volume X 3회 : W 1, W 2, W 3) ( W 2, W 3 시료 : W 2, W 3 시료 중 12 ㎕를 취하여 5 X sample buffer 3㎕ 들어있는 tube 에 넣는다. ) 6) bed volume(200 ㎕) 10배의 elution buffer를 400 ㎕씩 5번에 나누어 흘려주어 column을 지난 elution fraction 을 받는다. (E 1, E 2 , E 3, E 4, E 5 시료 : elution의 각 fraction 별 sample 8 ㎕ 를 취하여 5 X sample buffer 2 ㎕가 들어있는 tube 에 넣는다. )

Procedure 5. SDS-PAGE 확인 1) 12. 5% SDS-PAGE 을 만든다. 2) sample에 sample buffer를 가하여 준비한 microtube를 3 min동안 끓인다. 3) sample을 loading한다. 4) 80 V에서 10 min동안 초기 running 한다. 5) 130 V 에서 dye가 끝에 올 때까지 running한다. 5) Coomassie brilliant blue (staining solution)로 30 min동안 staining한다. 6) destaining solution에 넣고 dye를 제거한다. - destaining된 gel을 scan하여 생화학과 게시판에 올릴 예정
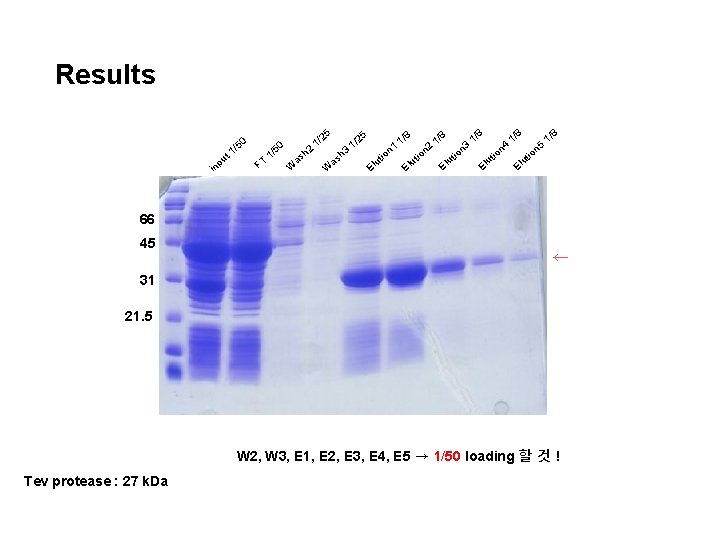
Results t u np i 50 1/ / 1 T F 5 /2 50 1 2 h as W 5 h W as 8 1/ 1/ /2 1 3 El ut io n 1 El ut io n 2 8 El 1/ ut io n 3 8 El /8 1 5 n tio 8 1/ ut io n 4 u El 66 45 ← 31 21. 5 W 2, W 3, E 1, E 2, E 3, E 4, E 5 → 1/50 loading 할 것 ! Tev protease : 27 k. Da
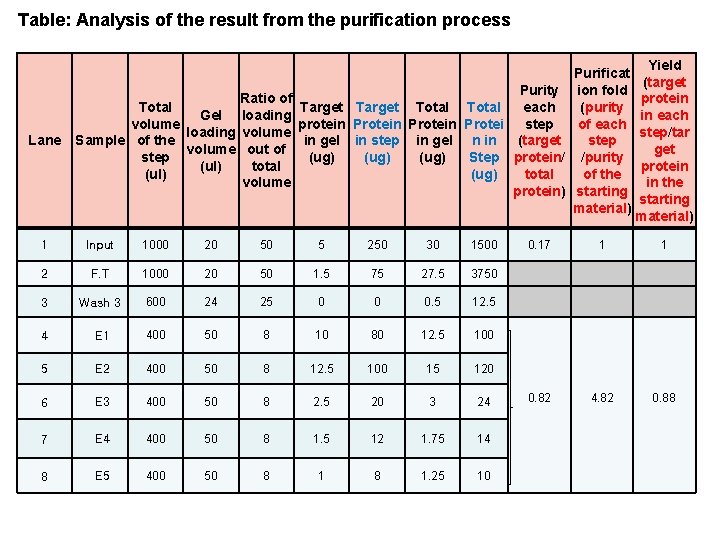
Table: Analysis of the result from the purification process Lane Purificat Purity ion fold Ratio of Target Total each Total (purity Gel loading volume protein Protein Protei step of each loading volume in gel in step in gel n in (target Sample of the step volume out of step (ug) Step protein/ /purity (ug) (ul) total (ul) (ug) total of the volume protein) starting material) 1 Input 1000 20 50 5 250 30 1500 2 F. T 1000 20 50 1. 5 75 27. 5 3750 3 Wash 3 600 24 25 0 0 0. 5 12. 5 4 E 1 400 50 8 10 80 12. 5 100 5 E 2 400 50 8 12. 5 100 15 120 6 E 3 400 50 8 2. 5 20 3 24 7 E 4 400 50 8 1. 5 12 1. 75 14 8 E 5 400 50 8 1. 25 10 0. 17 1 0. 82 4. 82 Yield (target protein in each step/tar get protein in the starting material) 1 0. 88

- Slides: 21