Protein Methods Functions Andy Howard Biochemistry Lectures Spring

Protein Methods & Functions Andy Howard Biochemistry Lectures, Spring 2019 Tuesday 5 February 2019

How to study proteins; what they do n n We will complete our discussion of protein methods today Not all proteins are enzymes, but many are; here we’ll discuss the numerous roles that proteins play in biology 02/05/2019 Protein Methods & Functions P. 2 of 56
Topics for today n Methods Purification n Characterization n Structure n n Moonlighting n Protein functions n n n Structure Enzymes Electron-transport Storage & transport Hormones Receptors 02/05/2019 Protein Methods & Functions Page 3 of 56

Electrophoresis (CF&M§ 5. 3) n n n Separating analytes by charge by subjecting a mixture to a strong electric field Gel electrophoresis: field applied to a semisolid matrix Can be used for charge (directly) or size (indirectly) 02/05/2019 Protein Methods & Functions P. 4 of 56
SDS-PAGE n n n Sodium dodecyl sulfate: strong detergent, applied to protein Charged species binds quantitatively Denatures protein n n Good: initial shape irrelevant Bad: it’s no longer folded 02/05/2019 Protein Methods & Functions P. 5 of 56
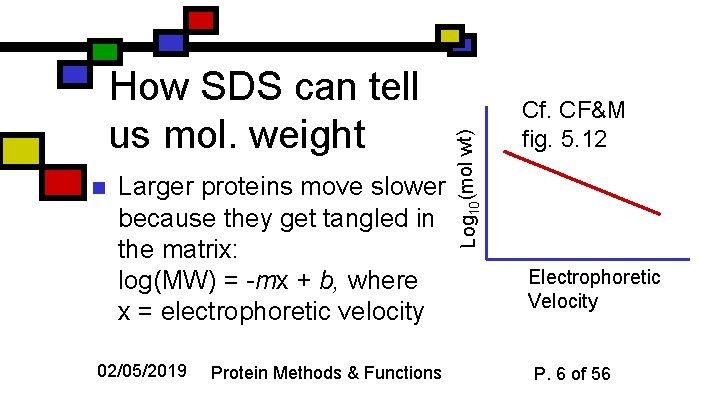
n Larger proteins move slower because they get tangled in the matrix: log(MW) = -mx + b, where x = electrophoretic velocity 02/05/2019 Protein Methods & Functions Log 10(mol wt) How SDS can tell us mol. weight Cf. CF&M fig. 5. 12 Electrophoretic Velocity P. 6 of 56
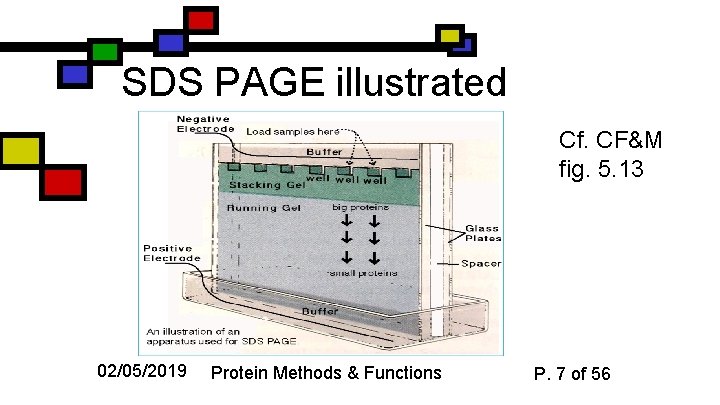
SDS PAGE illustrated Cf. CF&M fig. 5. 13 02/05/2019 Protein Methods & Functions P. 7 of 56
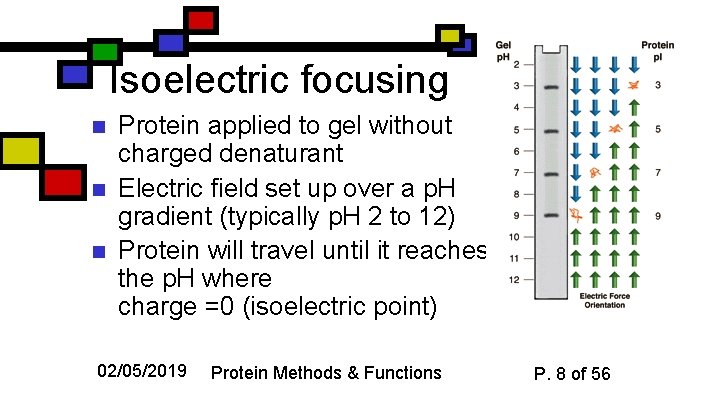
Isoelectric focusing n n n Protein applied to gel without charged denaturant Electric field set up over a p. H gradient (typically p. H 2 to 12) Protein will travel until it reaches the p. H where charge =0 (isoelectric point) 02/05/2019 Protein Methods & Functions P. 8 of 56

Using Isoelectric Focusing n n Sensitive to single changes in charge (e. g. asp asn) Readily used preparatively with samples that are already semi-pure 02/05/2019 Protein Methods & Functions P. 9 of 56

Applying this method n n Spectroscopy is more relevant for identification of moieties than for structure determination Quenching of fluorescence sometimes provides structural information 02/05/2019 Protein Methods & Functions P. 10 of 56

Mass spectrometry as an analytical tool n n n Mass spectrometry separates molecular species according to their mass/charge value It’s been used in chemistry for a century but couldn’t be applied to proteins until two techniques where developed in the 1980’s that preserved their properties: Electrospray and MALDI; Cf. CF&M section 5 A 02/05/2019 Protein Methods & Functions P. 11 of 56

i. Clicker quiz question 1 1. A protein has p. I = 4. At p. H=7 its charge will be n (a) positive n (b) negative n (c) neutral n (d) insufficient information provided. 02/05/2019 Protein Methods & Functions P. 12 of 56

i. Clicker question 2 2. Which of the following techniques does not separate proteins by size? n (a) SDS-PAGE n (b) Size-exclusion n (c) Isoelectric focusing n (d) Mass spectrometry n (e) All four of these separate by size. 02/05/2019 Protein Methods & Functions P. 13 of 56

Structure Methods!. . . Warning: Specialty Content! n n n I determine protein structures (and develop methods for determining protein structures) as my own research focus So it’s hard for me to avoid putting a lot of emphasis on this material But today I’m allowed to do that, because it’s one of the stated topics of the day. 02/05/2019 Protein Methods & Functions P. 14 of 56

How do we determine structure? (CF&M § 4. 4) n n We can distinguish between methods that require little prior knowledge (crystallography, NMR, Cryo. EM) and methods that answer specific questions (XAFS, fiber, …) This distinction isn’t entirely clear-cut 02/05/2019 Protein Methods & Functions P. 15 of 56

Crystallography: overview n n Crystals are translationally ordered 3 -D arrays of molecules Conventional solids are usually crystals Proteins have to be coerced into crystallizing … but once they’re crystals, they behave like other crystals, mostly 02/05/2019 Protein Methods & Functions P. 16 of 56

How are protein crystals unusual? n n Aqueous interactions required for crystal integrity: they disintegrate if dried Bigger unit cells (~10 nm, not 1 nm) Intermolecular forces are weak ionic forces Small # of unit cells and static disorder means they don’t scatter terribly well Determining 3 D structures is feasible but difficult n 02/05/2019 Protein Methods & Functions P. 17 of 56

Crystal structures: Fourier transforms of diffraction results n n n Position of spots tells you how big the unit cell is Intensity tells you what the contents are We’re using electromagnetic radiation, which behaves like a wave, exp(2 ik • x) = cos 2 k • x + isin 2 k • x 02/05/2019 Protein Methods & Functions P. 18 of 56

Relating ρ(r) to intensities n n n Therefore intensity Ihkl = C*|Fhkl|2 Fhkl is a complex coefficient in the Fourier transform of the electron density in the unit cell: (r) = (1/V) hkl Fhkl exp(-2 ih • r) Inverse of that: Fhkl = �V (r) exp(2 ih • r) 02/05/2019 Protein Methods & Functions P. 19 of 56
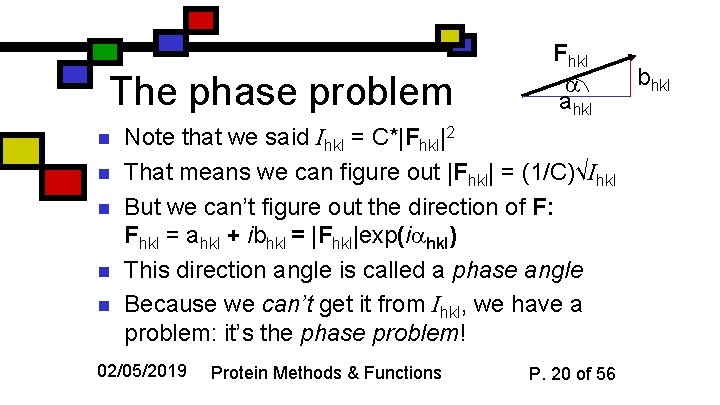
The phase problem n n n Fhkl ahkl Note that we said Ihkl = C*|Fhkl|2 That means we can figure out |Fhkl| = (1/C)√Ihkl But we can’t figure out the direction of F: Fhkl = ahkl + ibhkl = |Fhkl|exp(i hkl) This direction angle is called a phase angle Because we can’t get it from Ihkl, we have a problem: it’s the phase problem! 02/05/2019 Protein Methods & Functions P. 20 of 56 bhkl
What can we learn? n n n Electron density map + sequence we can determine the positions of all the non-H atoms in the protein—maybe! Best resolution possible: Dmin = / 2 Realistic resolution usually poorer than that 02/05/2019 Protein Methods & Functions P. 21 of 56

What else can we learn? n n Hydrogen positions can be inferred, especially if you are able to get highresolution data (see next slide) Atomic mobility can estimated for intermediate to high resolution data 02/05/2019 Protein Methods & Functions P. 22 of 56

Limitations of resolution n n Low values of Dmin mean more detail and better understanding Often the crystal doesn’t diffract ideally, so Dmin is larger than /2— 1. 5Å, 2. 5Å, or worse Dmin ~ 2. 5Å tells us where backbone and most side-chain atoms are Dmin ~ 1. 2Å: all protein non-H atoms, most solvent, some disordered atoms; some H’s 02/05/2019 Protein Methods & Functions P. 23 of 56

What does this look like? n n Takes some experience to interpret Automated fitting programs work pretty well with Dmin < 2. 1Å 02/05/2019 ATP binding to a protein of unknown function: S. H. Kim Protein Methods & Functions P. 24 of 56

Macromolecular NMR n n n NMR is a mature field Depends on resonant interaction between EM fields and unpaired nucleons (1 H, 15 N, 31 S) Raw data yield interatomic distances Conventional spectra of proteins are too muddy to interpret Multi-dimensional (2 -4 D) techniques: initial resonances coupled with others 02/05/2019 Protein Methods & Functions P. 25 of 56
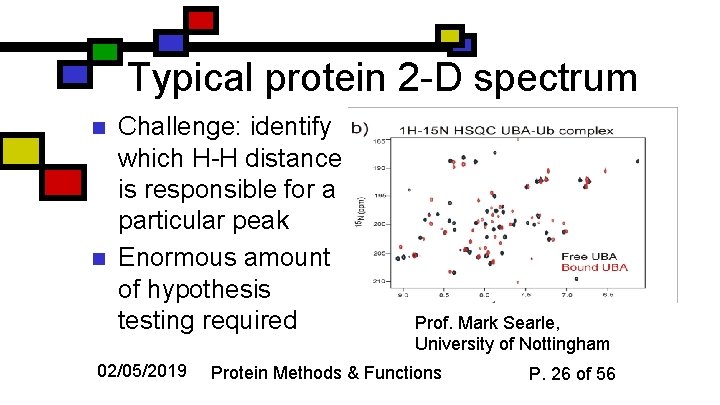
Typical protein 2 -D spectrum n n Challenge: identify which H-H distance is responsible for a particular peak Enormous amount of hypothesis testing required 02/05/2019 Prof. Mark Searle, University of Nottingham Protein Methods & Functions P. 26 of 56

Results n n Often there’s a family of structures that satisfy the NMR data equally well Can be portrayed as a series of threads tied down at unambiguous assignments They portray the protein’s structure in solution Ambiguities partly represent real molecular diversity; but they also represent atoms that area in truth well-defined, but the NMR data don’t provide the unambiguous assignment 02/05/2019 Protein Methods & Functions P. 27 of 56

Comparing NMR to X-ray n n n NMR family of structures often reflects real conformational heterogeneity Nonetheless, it’s hard to visualize what’s happening at the active site at any instant Hydrogens sometimes well-located in NMR; they’re often the least defined atoms in an Xray structure 02/05/2019 Protein Methods & Functions P. 28 of 56

NMR vs. X-ray, continued n n The NMR structure is obtained in solution! Hard to make NMR work if MW > 55 k. Da, and even when you can, it takes a lot of computer time 02/05/2019 Protein Methods & Functions P. 29 of 56

What does it mean when NMR and X-ray structures differ? n n Lattice forces may have tied down or moved surface amino acids in X-ray structure NMR may have errors in it X-ray may have errors in it (measurable) X-ray structure often closer to true atomic resolution X-ray structure has built-in reliability checks n 02/05/2019 Protein Methods & Functions P. 30 of 56

n n n Cryoelectron microscopy Like X-ray crystallography, EM damages the samples Samples analyzed < 100 K survive better 2 -D arrays of molecules n Spatial averaging to improve resolution n Discerning details ≥ 4Å resolution Can be used with crystallography n 02/05/2019 Protein Methods & Functions P. 31 of 56

Solution scattering n n n Proteins in solution scatter X-rays in characteristic ways Low-resolution structural information available Does not require crystals Until ~ 2000 you needed high [protein] Thanks to Bio. CAT, SAXS on dilute proteins is becoming more feasible Hypothesis-based analysis n 02/05/2019 Protein Methods & Functions P. 32 of 56

Fiber Diffraction Some proteins, like many DNA molecules, possess approximate fibrous order (2 -D ordering) n Produce characteristic fiber diffraction patterns n Collagen, muscle proteins, filamentous viruses 02/05/2019 n Protein Methods & Functions P. 33 of 56
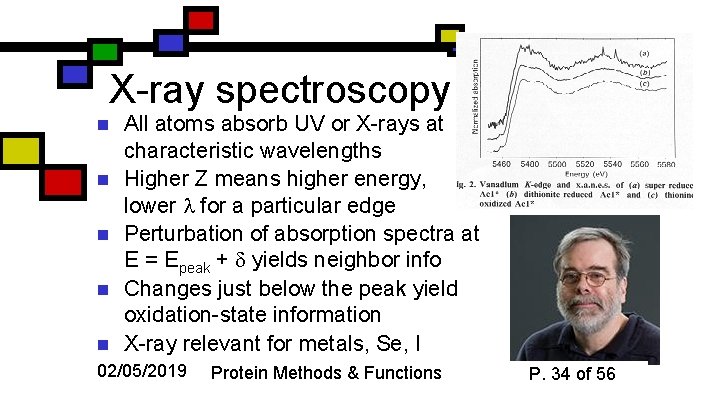
X-ray spectroscopy n n n All atoms absorb UV or X-rays at characteristic wavelengths Higher Z means higher energy, lower for a particular edge Perturbation of absorption spectra at E = Epeak + yields neighbor info Changes just below the peak yield oxidation-state information X-ray relevant for metals, Se, I 02/05/2019 Protein Methods & Functions P. 34 of 56

n n n Mass spectrometry as a structural tool MS tells you molecular weights Can give high precision in m/m Not inherently a way of determining structure Can distinguish oligomeric state Coupled with proteolytic digestion, it can be used to find fragmentation patterns 02/05/2019 Protein Methods & Functions P. 35 of 56

Circular dichroism n n n Proteins in solution can rotate polarized light Amount of rotation varies with Effect depends on interaction with secondary structure elements, esp. 02/05/2019 Protein Methods & Functions P. 36 of 56
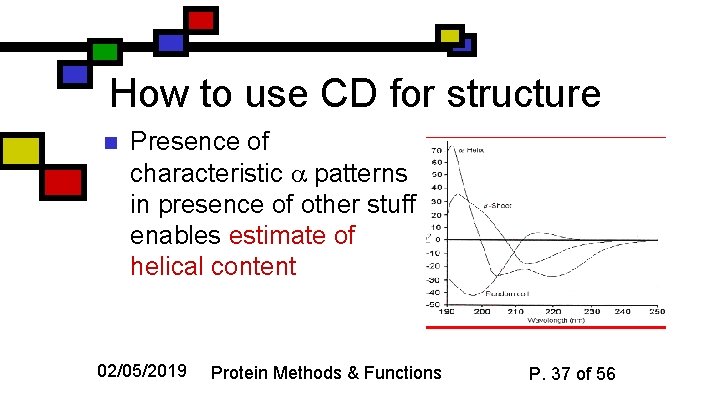
How to use CD for structure n Presence of characteristic patterns in presence of other stuff enables estimate of helical content 02/05/2019 Protein Methods & Functions P. 37 of 56

i. Clicker question 3 3. Which of these structures provides a n n more detailed and precise knowledge of n the structure of lactate n dehydrogenase? 1 T 2 F 5 W 8 J (a) PDB 1 T 2 F, Dmin=3Å (b) PDB 5 W 8 J, Dmin=1. 55Å (c) both are equally informative (d) insufficient information to tell 02/05/2019 Protein Methods & Functions P. 38 of 56
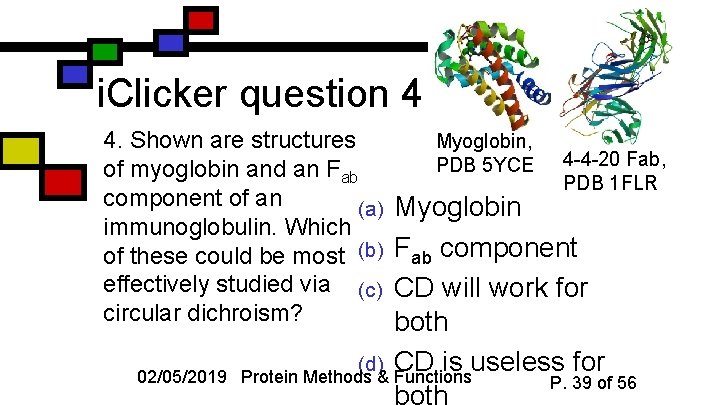
i. Clicker question 4 4. Shown are structures of myoglobin and an Fab component of an (a) immunoglobulin. Which of these could be most (b) effectively studied via (c) circular dichroism? 02/05/2019 Myoglobin, PDB 5 YCE 4 -4 -20 Fab, PDB 1 FLR Myoglobin Fab component CD will work for both (d) CD is useless for Protein Methods & Functions P. 39 of 56 both

Protein function n n We’ll devote most of the rest of the lecture to a discussion of the various functions performed by proteins We’ll do a quick run-through of the various functions, and then discuss PTM and allostery. 02/05/2019 Protein Methods & Functions P. 40 of 56

Classes of proteins n n Next segment of this lecture: small encyclopedia of the protein functions Reminder: proteins can take on more than one function 02/05/2019 Protein Methods & Functions Arginosuccinate lyase / Delta crystallin PDB 1 auw, 2. 5Å 206 k. Da tetramer P. 41 of 56

Moonlighting proteins n n n A protein may evolve for one purpose … then it gets co-opted for another Studies by C. Jeffery et al 02/05/2019 Protein Methods & Functions P. 42 of 56

Structural proteins n n Mechanical or scaffolding tasks Don’t do chemistry, unless this is a chemical reaction: (Person standing upright) (Person lying in a puddle on the floor) Examples: collagen, fibroin, keratin Often enzymes are recruited to perform structural roles 02/05/2019 Protein Methods & Functions Collagen model PDB 1 K 6 F P. 43 of 56

Enzymes n n Enzymes are biological catalysts, i. e. their job is to reduce the activation energy barrier between substrates and products Tend to be at least 12 k. Da (why? You need that much scaffolding) Usually but not always aqueous Usually organized with hydrophilic residues facing outward 02/05/2019 Protein Methods & Functions hen egg-white lysozyme 14. 2 k. Da monomer PDB 2 vb 1, 0. 65Å P. 44 of 56

Electron-transport proteins n Involved in oxidation-reduction reactions via n n n Incorporated metal ions Small organic moieties (NAD, FAD) Generally not enzymes: they’re ultimately altered by their reactions 02/05/2019 Protein Methods & Functions Recombinant human cytochrome c PDB 1 J 3 S NMR structure 11. 4 k. Da P. 45 of 56

But they can be parts of larger enzyme systems! n Some participate in larger enzyme complexes than can restore them to their original state 02/05/2019 Protein Methods & Functions P. 46 of 56

Sizes and characteristics n Some ET proteins: fairly small n n n Cytochrome c Some flavodoxins Others are multi-polypeptide complexes 02/05/2019 Protein Methods & Functions Anacystis flavodoxin PDB 1 czn 1. 7Å 18. 6 k. Da P. 47 of 56
Coenzymes and metals in electron-transport proteins n Coenzymes or metal ions may be closely associated (covalent in cytochromes) or more loosely bound 02/05/2019 Protein Methods & Functions P. 48 of 56

Storage and transport proteins n n n Hemoglobin, myoglobin classic examples “honorary enzymes”: share some characteristics with enzymes Sizes vary widely 02/05/2019 Protein Methods & Functions Sperm-whale myoglobin 18 k. Da monomer PDB 1 MTJ, 1. 7Å P. 49 of 56

Intracellular transporters n n Many transporters operate over much smaller size-scales than hemoglobin (µm vs. m): often involved in transport across membranes We’ll discuss intracellular transport a lot! 02/05/2019 Protein Methods & Functions P. 50 of 56

Why do we have storage proteins? n Many metabolites are toxic in the wrong places or at the wrong times n n Oxygen is nasty Too much Ca 2+ or Fe 3+ can be hazardous 02/05/2019 Protein Methods & Functions T. maritima ferritin 8*(18 k. Da) PDB 1 z 4 a P. 51 of 56

Why do we need these? n So storage proteins provide ways of encapsulating small molecules until they’re needed; then they’re released 02/05/2019 Protein Methods & Functions P. 52 of 56

Hormones n n n Transported signaling molecules, secreted by one tissue and detected by receptors in another tissue Signal noted by the receptor will trigger some kind of response in the second tissue. They’re involved in cell-cell or tissue-totissue communication. 02/05/2019 Protein Methods & Functions Human insulin PDB 1 t 1 k 3. 3+2. 3 k. Da P. 53 of 56

Some hormones aren’t proteins n Not all hormones are proteins n n n Some are organic, non-peptidic moieties Others: peptide oligomers, too small to be proteins But some hormones are in fact normalsized proteins. 02/05/2019 Protein Methods & Functions P. 54 of 56
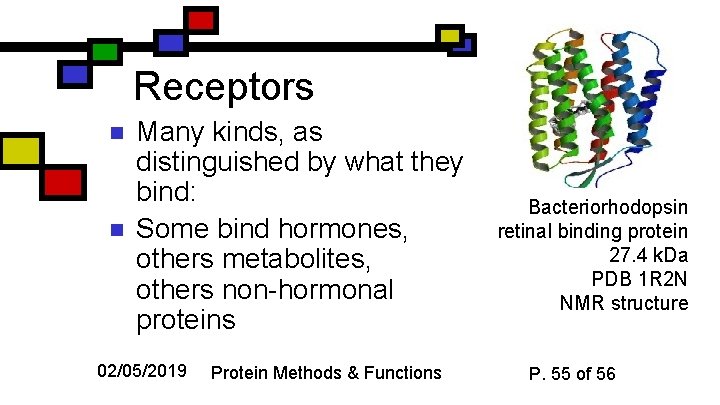
Receptors n n Many kinds, as distinguished by what they bind: Some bind hormones, others metabolites, others non-hormonal proteins 02/05/2019 Protein Methods & Functions Bacteriorhodopsin retinal binding protein 27. 4 k. Da PDB 1 R 2 N NMR structure P. 55 of 56

Receptors in membranes n Receptors are usually membraneassociated: n n a soluble piece sticking out Hydrophobic piece in the membrane sometimes another piece on the other side of the membrane Membrane part often helical: usually odd # of spanning helices (7? ) 02/05/2019 Protein Methods & Functions P. 56 of 56
- Slides: 56