PROTEIN IDENTIFICATION BY MASS SPECTROMETRY OBJECTIVES To become

PROTEIN IDENTIFICATION BY MASS SPECTROMETRY

OBJECTIVES • To become familiar with matrix assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS) • To become familiar with contemporary protein identification approaches • To identify unknown proteins from collected samples by peptide mass fingerprinting, using mass spectral data of tryptic peptides for database querying

What is mass spectrometry ? Most accurate technique available to determine the mass of an analyte (proteins, peptides, small molecules)
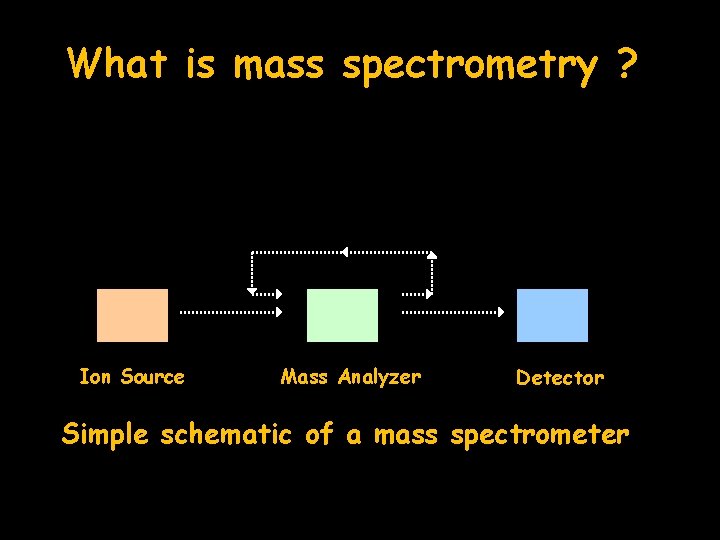
What is mass spectrometry ? Ion Source Mass Analyzer Detector Simple schematic of a mass spectrometer

Basics of mass spectrometry • Determination of mass to charge ratio (m/z) – note in MALDI-TOF MS, the charge is 1 so we essentially determine mass (i. e. m/1) • Components of MS - ionization and entrance to gas phase - ion (peptide) separation - mass analysis

Output from MS - mass-to-charge ratio (m/z) ratio of exact mass to net charge For example: 802 m/z ion (z=1; +1 charged) = 802 amu 500 m/z ion (z=2; +2 charged) = 1000 amu 327 m/z ion (z=3; +3 charged) = ? ? amu

Terminology • Isotopes • Ionization • Resolution
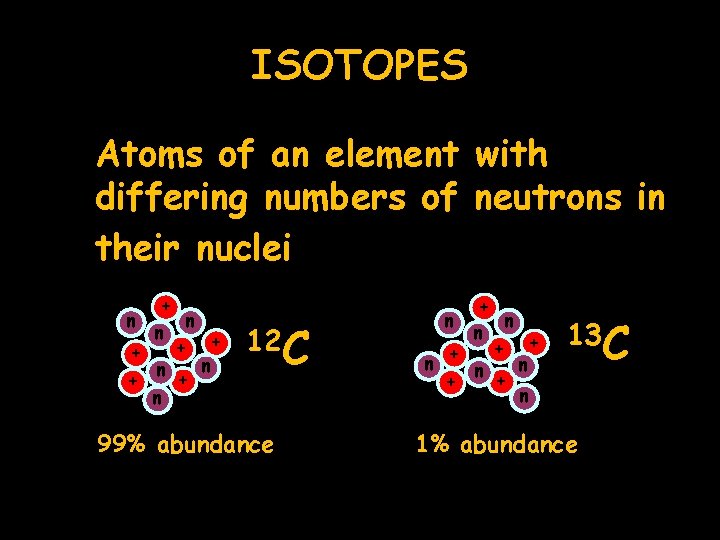
ISOTOPES Atoms of an element with differing numbers of neutrons in their nuclei + n n + + n n 12 C 99% abundance + n n + + + n n n + + n n 13 C 1% abundance
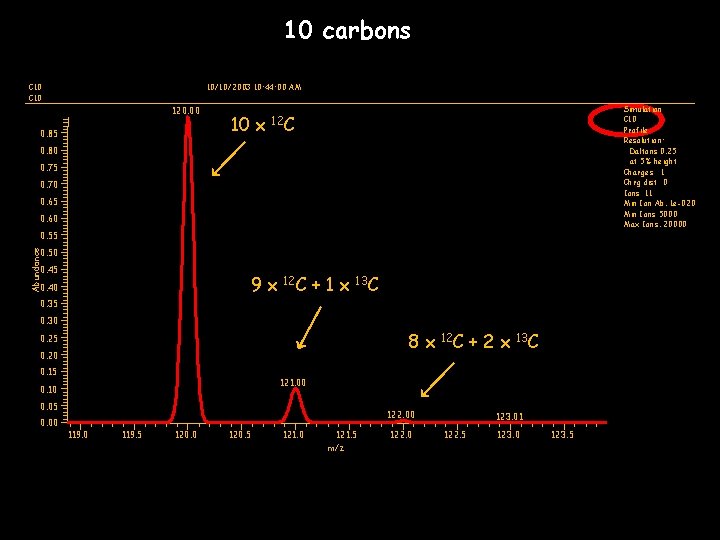
10 carbons C 10 10/10/2003 10: 44: 00 AM 120. 00 0. 85 10 x Simulation C 10 Profile Resolution: Daltons 0. 25 at 5% height Charges 1 Chrg dist 0 Ions 11 Min Ion Ab. 1 e-020 Min Ions 5000 Max Ions. 20000 12 C 0. 80 0. 75 0. 70 0. 65 0. 60 0. 55 Abundance 0. 50 0. 45 9 x 0. 40 12 C +1 x 13 C 0. 35 0. 30 8 x 0. 25 0. 20 0. 15 +2 x 13 C 121. 00 0. 10 0. 05 0. 00 12 C 122. 00 119. 5 120. 0 120. 5 121. 0 121. 5 m/z 122. 0 123. 01 122. 5 123. 0 123. 5

RESOLUTION • How well two ions can be separated • Good resolution essential to distinguishing isotopes • Resolution usually decreases as the mass of the analyte increases
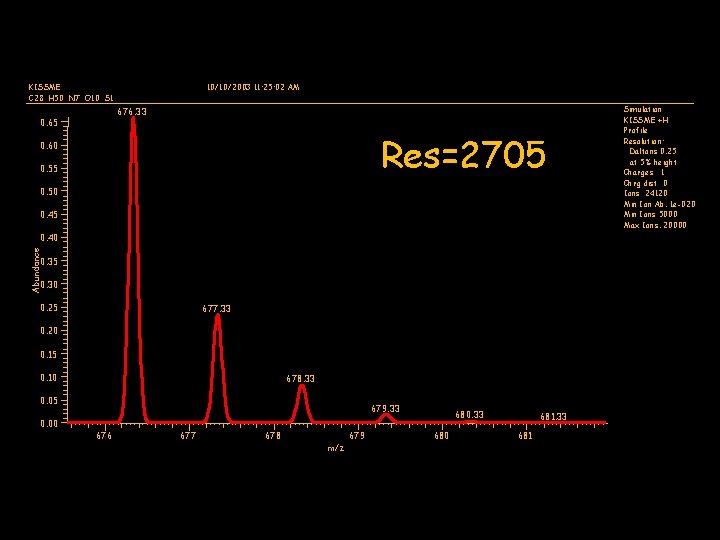
KISSME C 28 H 50 N 7 O 10 S 1 10/10/2003 11: 25: 02 AM 676. 33 0. 65 Res=2705 0. 60 0. 55 0. 50 0. 45 Abundance 0. 40 0. 35 0. 30 0. 25 677. 33 0. 20 0. 15 0. 10 678. 33 0. 05 0. 00 679. 33 676 677 678 679 m/z 680. 33 680 681. 33 681 Simulation KISSME +H Profile Resolution: Daltons 0. 25 at 5% height Charges 1 Chrg dist 0 Ions 24120 Min Ion Ab. 1 e-020 Min Ions 5000 Max Ions. 20000
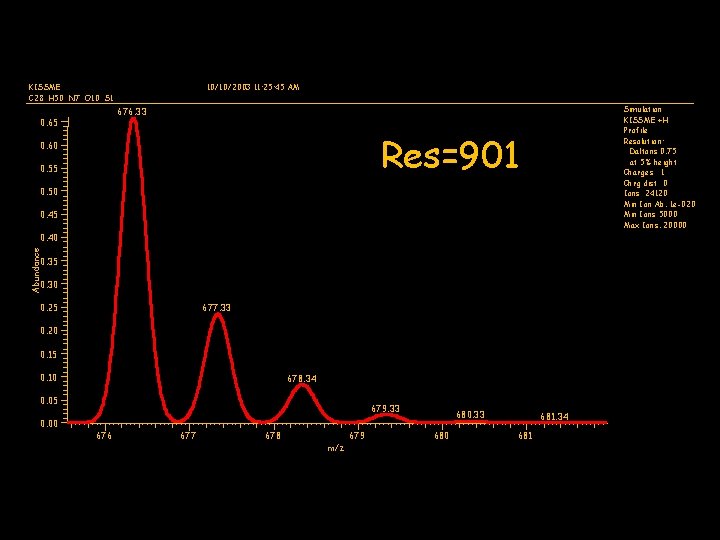
KISSME C 28 H 50 N 7 O 10 S 1 10/10/2003 11: 25: 45 AM Simulation KISSME +H Profile Resolution: Daltons 0. 75 at 5% height Charges 1 Chrg dist 0 Ions 24120 Min Ion Ab. 1 e-020 Min Ions 5000 Max Ions. 20000 676. 33 0. 65 Res=901 0. 60 0. 55 0. 50 0. 45 Abundance 0. 40 0. 35 0. 30 0. 25 677. 33 0. 20 0. 15 0. 10 678. 34 0. 05 0. 00 679. 33 676 677 678 679 m/z 680. 33 680 681. 34 681
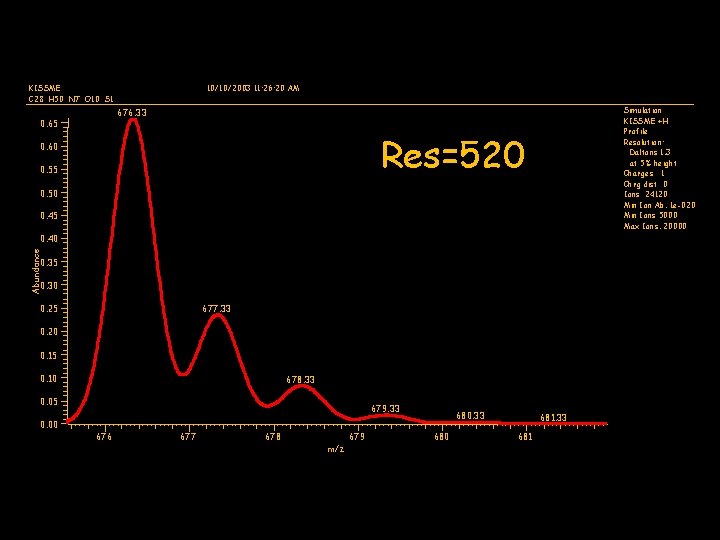
KISSME C 28 H 50 N 7 O 10 S 1 10/10/2003 11: 26: 20 AM Simulation KISSME +H Profile Resolution: Daltons 1. 3 at 5% height Charges 1 Chrg dist 0 Ions 24120 Min Ion Ab. 1 e-020 Min Ions 5000 Max Ions. 20000 676. 33 0. 65 Res=520 0. 60 0. 55 0. 50 0. 45 Abundance 0. 40 0. 35 0. 30 0. 25 677. 33 0. 20 0. 15 0. 10 678. 33 0. 05 0. 00 679. 33 676 677 678 679 m/z 680. 33 680 681. 33 681

IONIZATION • Giving an electric charge to a neutral species A + H+ = (A+H)+ A + Na+ = (A+Na)+ A - H = (A-H)-

Soft-ionization methods for proteomics • MALDI – In MALDI-TOF MS, ionization occurs when the laser hits the sample • Electrospray – In electrospray, ionization occurs in the liquid as it exits a spray needle

MALDI = Matrix-Assisted Laser Desorption / Ionization • analyte is dried with matrix on target – matrix: CHCA, DHB, sinapinic acid – matrix absorbs laser energy and transfers to analyte • generates predominantly singly-charged ions
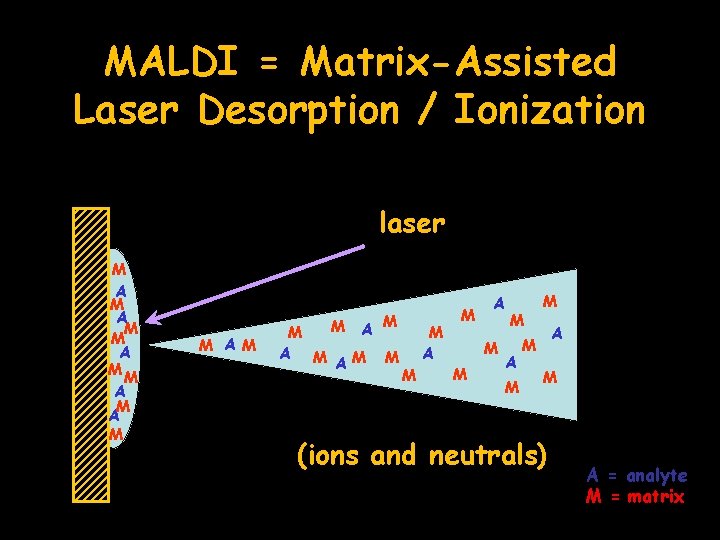
MALDI = Matrix-Assisted Laser Desorption / Ionization laser M A M A M M A M AM M A M M (ions and neutrals) A = analyte M = matrix

Mass Analyzers • • Time-Of-Flight (TOF) Quadrupole Ion Trap Ion Cyclotron Resonance

Mass to Charge Ratio (m/z) • Measure m/z not mass – C 7 H 7 mass is 91 Da – C 7 H 7+1; m/z = 91 – C 7 H 7+2; m/z = 45. 5
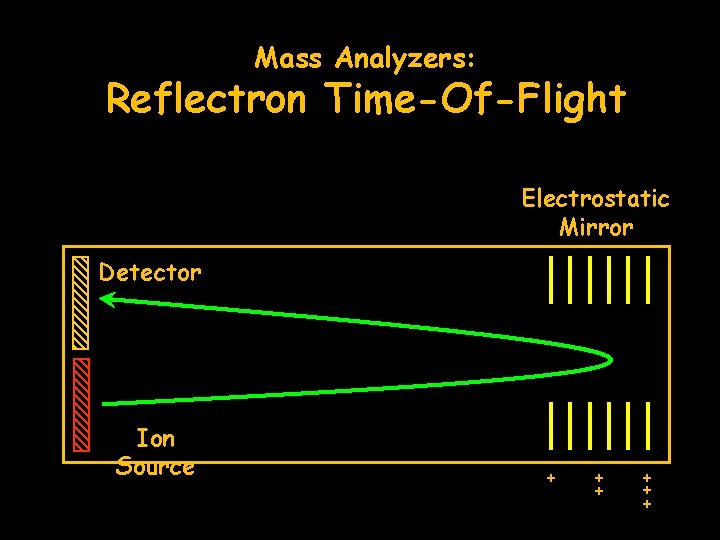
Mass Analyzers: Reflectron Time-Of-Flight Electrostatic Mirror Detector Ion Source + + +

MALDI-TOF mass spectrometers

MALDI-TOF MS matrix-assisted laser desorption / ionizationtime of flight mass spectrometer Peptides put on to plate surface Voyager DE-PRO
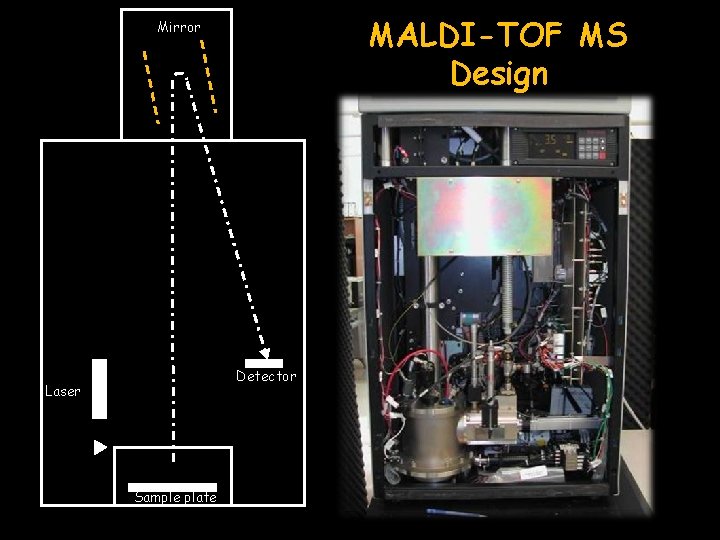
MALDI-TOF MS Design Mirror Detector Laser Sample plate
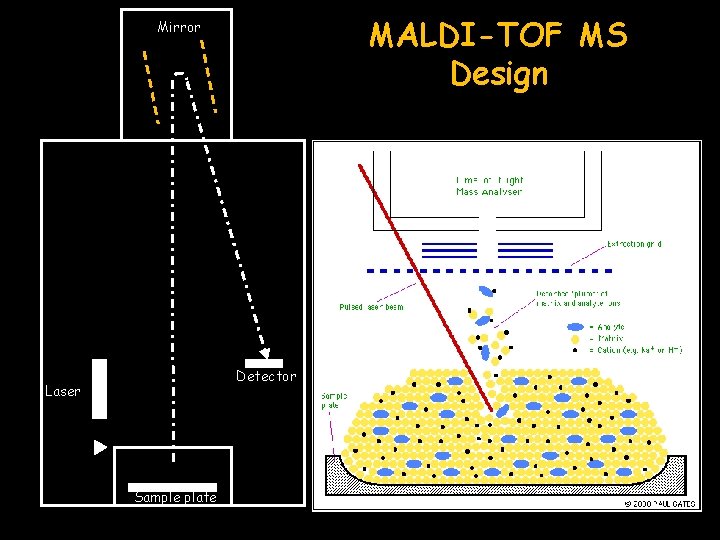
MALDI-TOF MS Design Mirror Detector Laser Sample plate
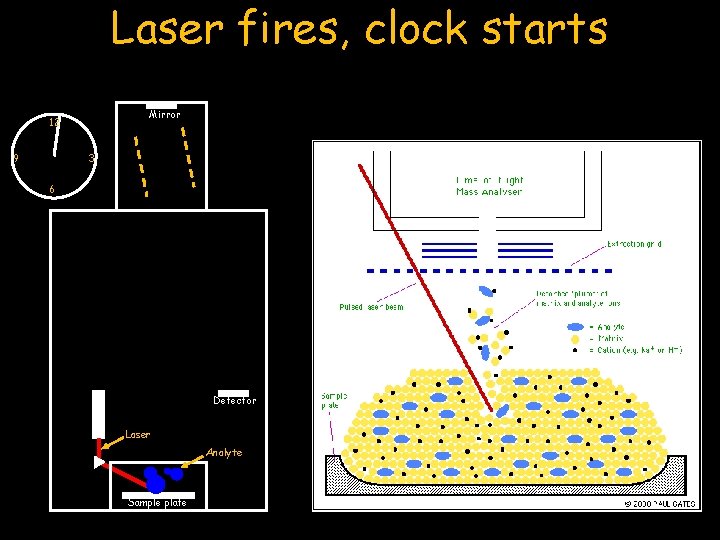
Laser fires, clock starts Mirror 12 9 3 6 Laser fires…Clock Starts Detector Laser Analyte Sample plate
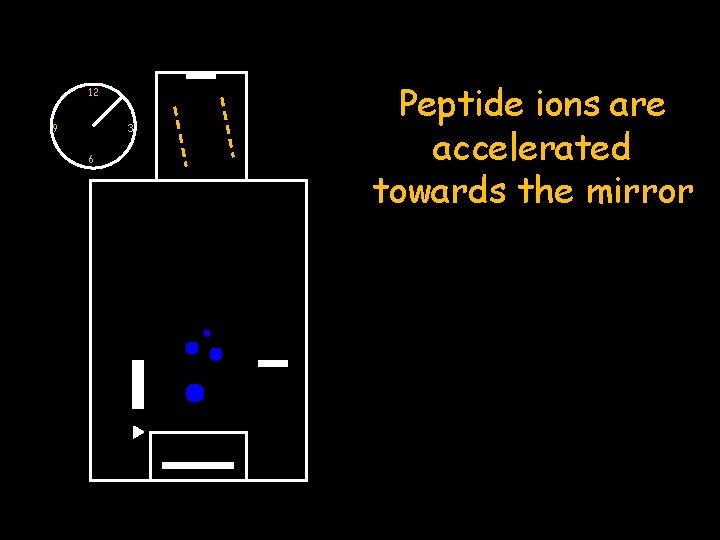
12 9 3 6 Peptide ions are accelerated towards the mirror
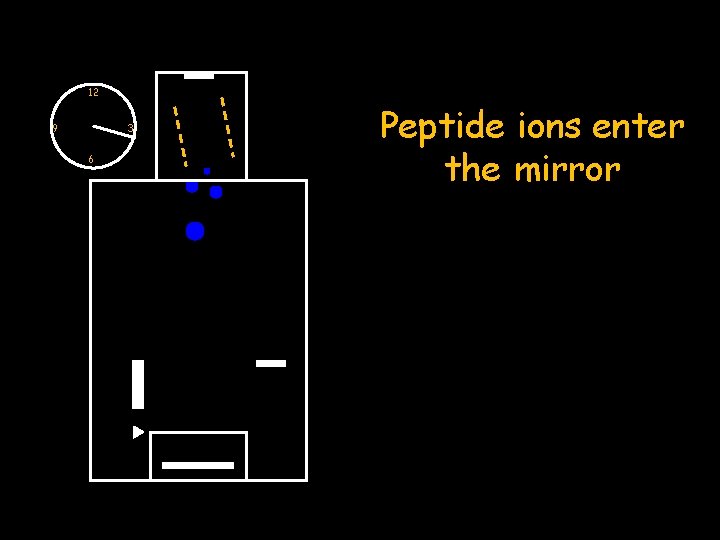
12 9 3 6 Peptide ions enter the mirror
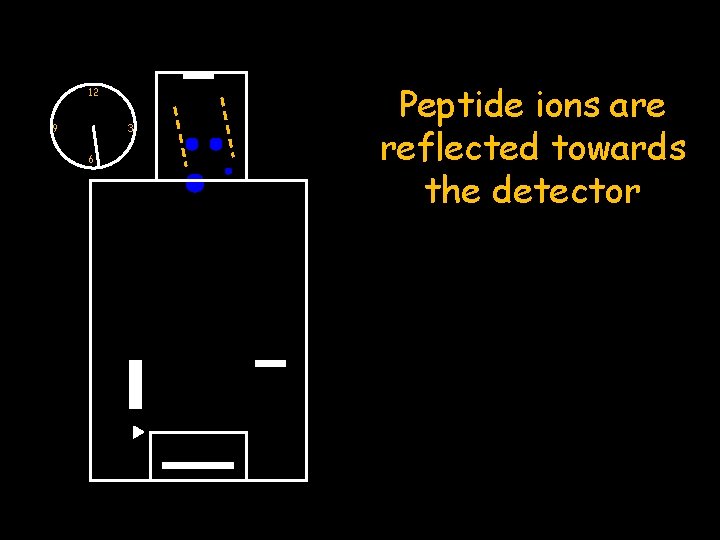
12 9 3 6 Peptide ions are reflected towards the detector
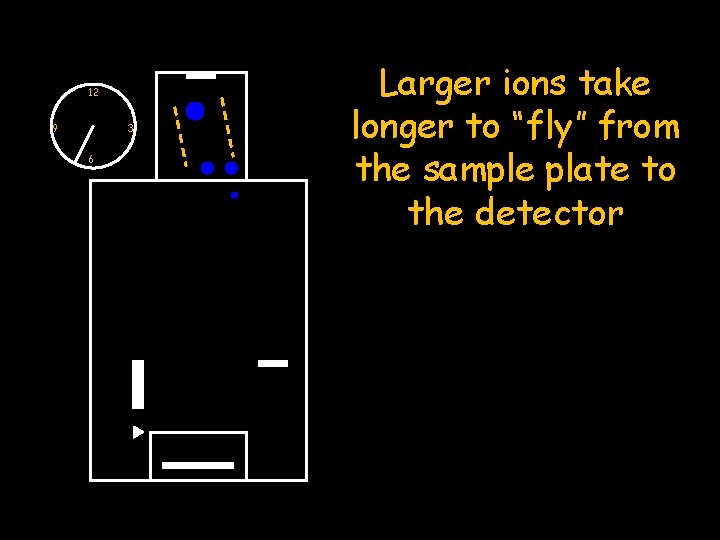
12 9 3 6 Larger ions take longer to “fly” from the sample plate to the detector
12 9 3 6 Peptide ions approach the detector
9 3 6 Smallest peptide ion reaches the detector Intensity scale 12 Mass scale
12 9 3 intensity 6 mass
9 3 6 “Mass Spectrum” shows distribution and intensity of peptides in the sample intensity 12 mass
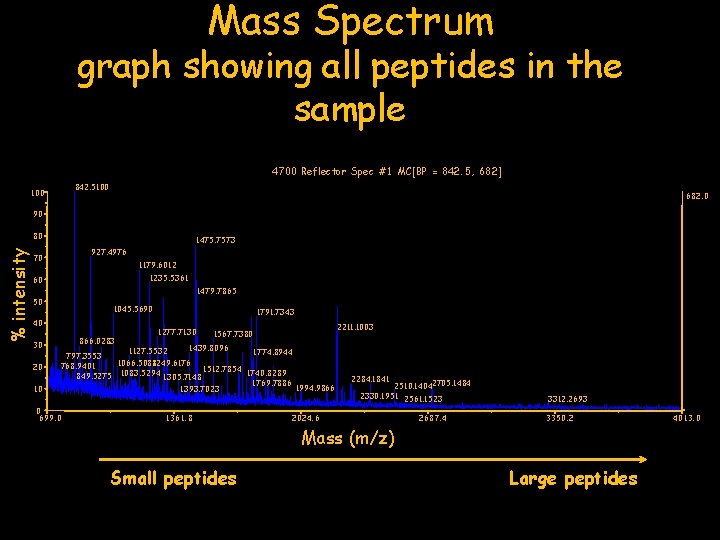
Mass Spectrum graph showing all peptides in the sample 4700 Reflector Spec #1 MC[BP = 842. 5, 682] 842. 5100 682. 0 90 % intensity 80 1475. 7573 927. 4976 70 1179. 6012 1235. 5361 60 1479. 7865 50 1045. 5690 40 30 20 10 1791. 7343 1277. 7130 2211. 1003 1567. 7380 866. 0283 1439. 8096 1127. 5532 1774. 8944 797. 3553 1066. 5088 1 249. 6176 768. 9401 1512. 7854 1740. 8289 849. 5275 1083. 5294 1305. 7148 1769. 7886 1994. 9866 1393. 7023 0 699. 0 1361. 8 2284. 1841 2510. 14042705. 1484 2330. 1951 2561. 1523 2024. 6 2687. 4 3312. 2693 3350. 2 Mass (m/z) Small peptides Large peptides 4013. 0
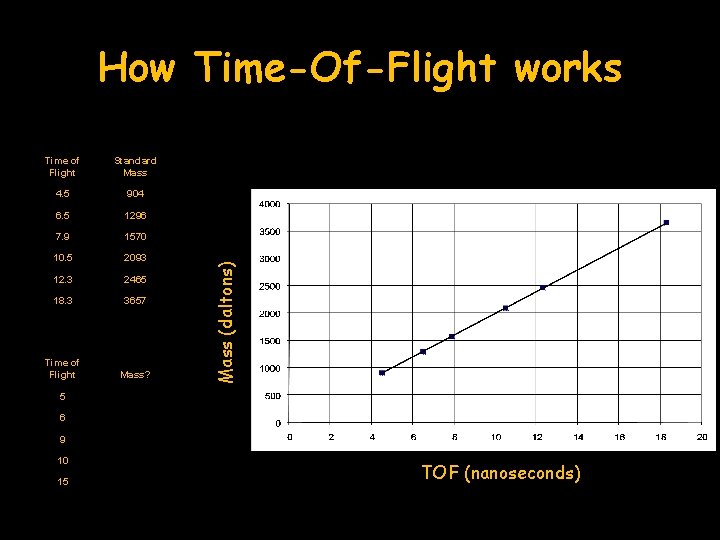
Time of Flight Standard Mass 4. 5 904 6. 5 1296 7. 9 1570 10. 5 2093 12. 3 2465 18. 3 3657 Time of Flight Mass? Mass (daltons) How Time-Of-Flight works 5 6 9 10 15 TOF (nanoseconds)
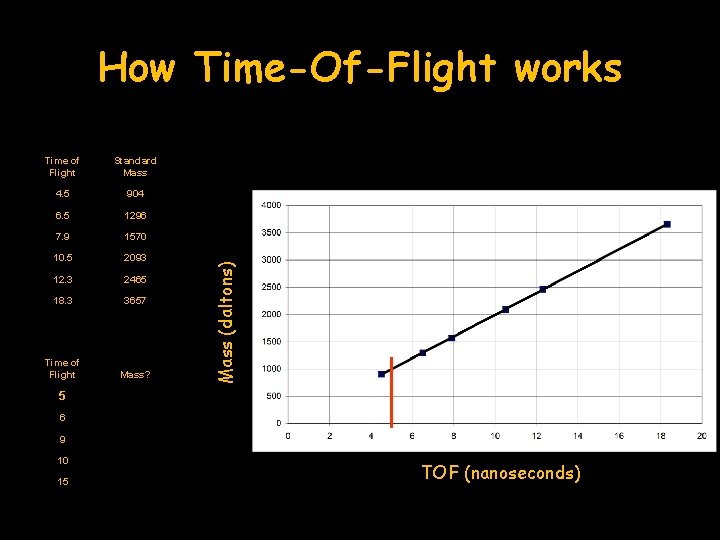
Time of Flight Standard Mass 4. 5 904 6. 5 1296 7. 9 1570 10. 5 2093 12. 3 2465 18. 3 3657 Time of Flight Mass? Mass (daltons) How Time-Of-Flight works 5 6 9 10 15 TOF (nanoseconds)
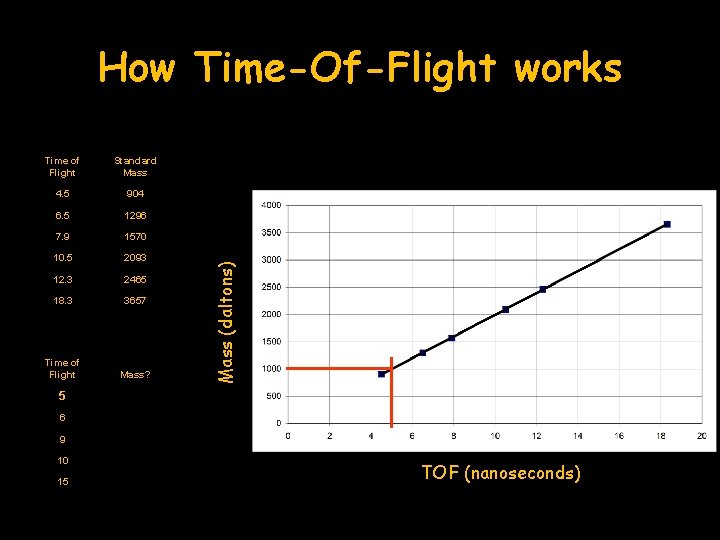
Time of Flight Standard Mass 4. 5 904 6. 5 1296 7. 9 1570 10. 5 2093 12. 3 2465 18. 3 3657 Time of Flight Mass? Mass (daltons) How Time-Of-Flight works 5 6 9 10 15 TOF (nanoseconds)
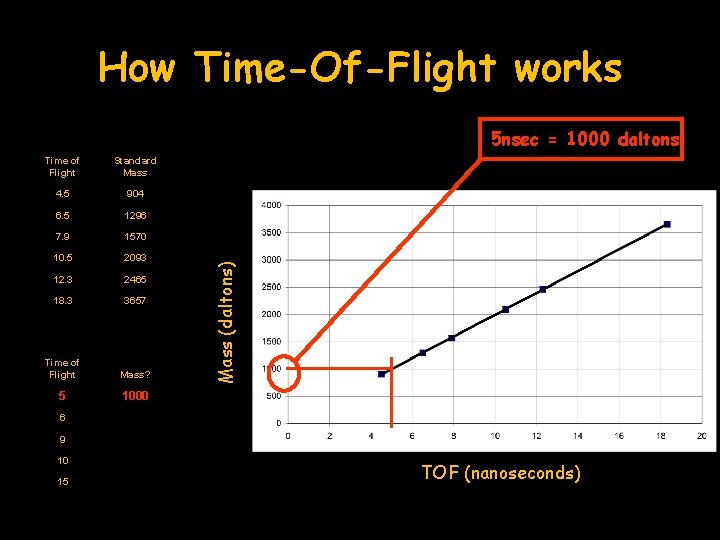
How Time-Of-Flight works Time of Flight Standard Mass 4. 5 904 6. 5 1296 7. 9 1570 10. 5 2093 12. 3 2465 18. 3 3657 Time of Flight Mass? 5 1000 Mass (daltons) 5 nsec = 1000 daltons 6 9 10 15 TOF (nanoseconds)
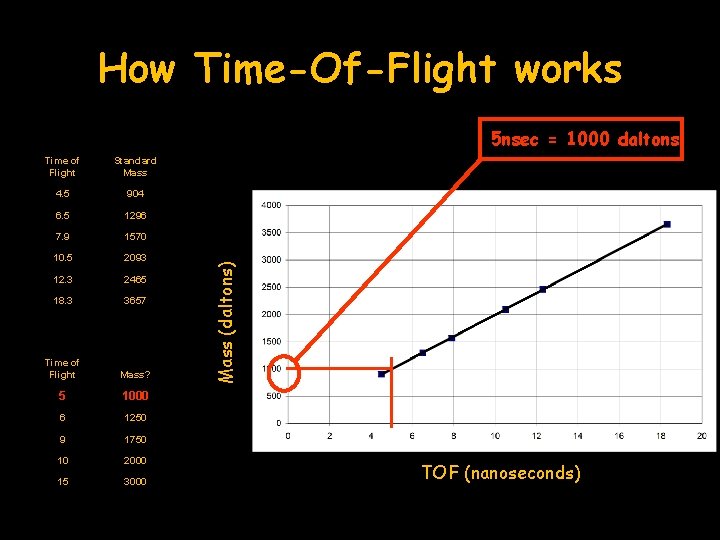
How Time-Of-Flight works Time of Flight Standard Mass 4. 5 904 6. 5 1296 7. 9 1570 10. 5 2093 12. 3 2465 18. 3 3657 Time of Flight Mass? 5 1000 6 1250 9 1750 10 2000 15 3000 Mass (daltons) 5 nsec = 1000 daltons TOF (nanoseconds)
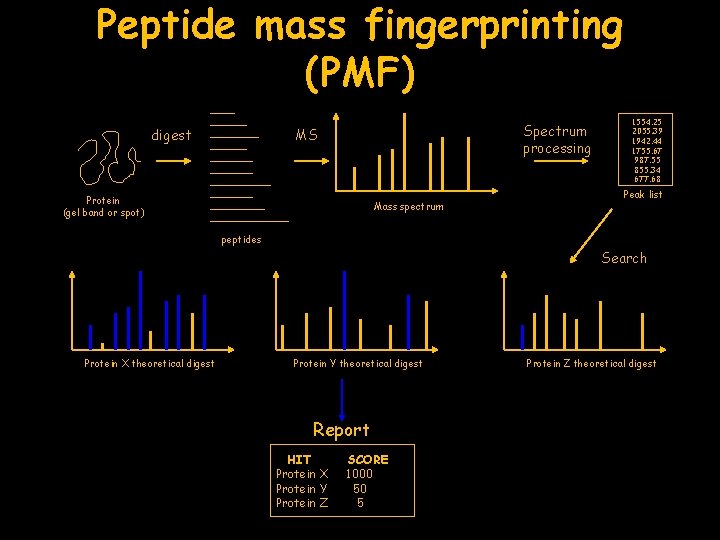
Peptide mass fingerprinting (PMF) digest Spectrum processing MS Protein (gel band or spot) Mass spectrum 1554. 25 2055. 39 1942. 44 1755. 67 987. 55 855. 34 677. 68 Peak list peptides Search Protein X theoretical digest Protein Y theoretical digest Report HIT Protein X Protein Y Protein Z SCORE 1000 50 5 Protein Z theoretical digest

Generate mass list of all peptides • Computer program used to convert graph data (from mass spectrum) to table data (list of peptide masses) 972. 5954 1007. 467 973. 4202 1023. 462 973. 4202 1036. 481 975. 6104 1066. 493 977. 4627 1082. 488 982. 4603 1105. 601 983. 4991 1153. 524 998. 4552 1172. 581
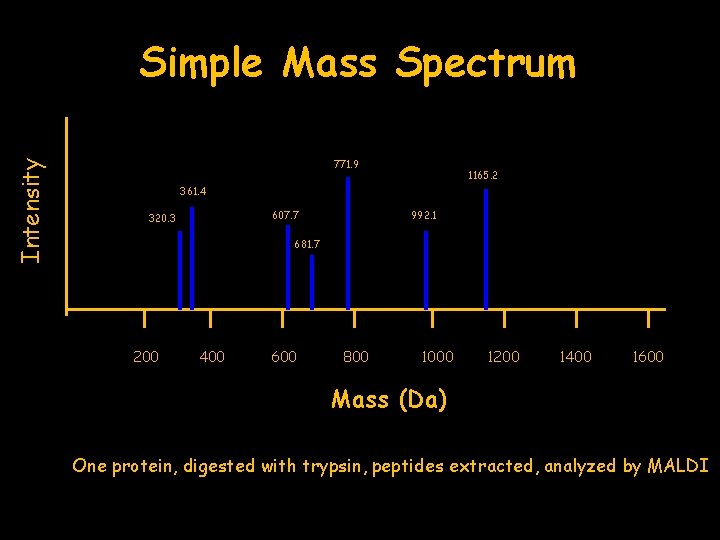
Intensity Simple Mass Spectrum 771. 9 1165. 2 361. 4 607. 7 320. 3 992. 1 681. 7 200 400 600 800 1000 1200 1400 1600 Mass (Da) One protein, digested with trypsin, peptides extracted, analyzed by MALDI

320. 3 361. 4 607. 7 681. 7 771. 9 992. 1 1165. 2 List of masses Database search using computer algorithm e. g. MASCOT EAT EGR ISPYK EMETR EMANYK PLEASEMAK EATSTHEYAR Sequence matches PLEASEMAKEMANYKRISPYKREMETREATSTHEYAREGREAT

QUESTIONS?
- Slides: 45