Protein Fold recognition Morten Nielsen CBS Bio Centrum

Protein Fold recognition Morten Nielsen, CBS, Bio. Centrum, DTU

Objectives • Understand the basic concepts of fold recognition • Learn why even sequences with very low sequence similarity can be modeled – Understand why is %id such a terrible measure for reliability • See the beauty of sequence profiles – Position specific scoring matrices (PSSMs)

Protein Homology modeling? • Identify template(s) – initial alignment • Can give you protein function • Improve alignment • Can give you active site • Backbone generation • Loop modeling • Most difficult part • Side chains • Refinement • Validation

How to do it? Identify fold (template) for modeling – Find the structure in the PDB database that resembles your new protein the most – Can be used to predict function – And maybe active sites Align protein sequence to template – Simple alignment methods – Sequence profiles – Threading methods – Pseudo force fields Model side chains and loops
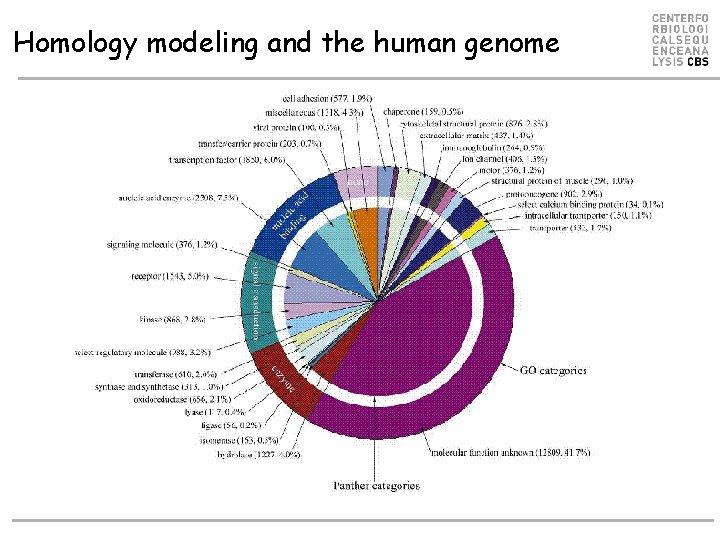
Homology modeling and the human genome
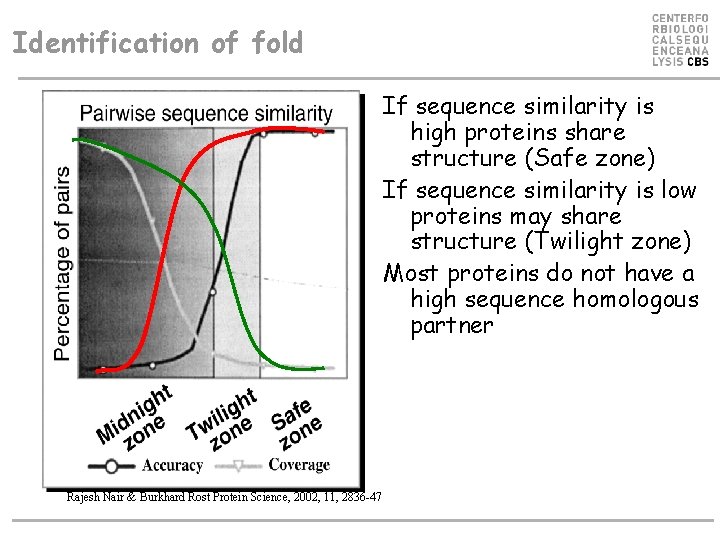
Identification of fold If sequence similarity is high proteins share structure (Safe zone) If sequence similarity is low proteins may share structure (Twilight zone) Most proteins do not have a high sequence homologous partner Rajesh Nair & Burkhard Rost Protein Science, 2002, 11, 2836 -47

Example. A post doc in our group did her Ph. D obtaining the structure of the sequence below >1 K 7 C. A TTVYLAGDSTMAKNGGGSGTNGWGEYLASYLSATVVNDAVAGRSARSYTREGRFENIADV VTAGDYVIVEFGHNDGGSLSTDNGRTDCSGTGAEVCYSVYDGVNETILTFPAYLENAAKL FTAKGAKVILSSQTPNNPWETGTFVNSPTRFVEYAELAAEVAGVEYVDHWSYVDSIYETL GNATVNSYFPIDHTHTSPAGAEVVAEAFLKAVVCTGTSLKSVLTTTSFEGTCL • What is the function • Where is the active site?

What would you do? • Function • Run Blast against PDB • No significant hits • Run Blast against NR (Sequence database) • Function is Acetylesterase? • Where is the active site?
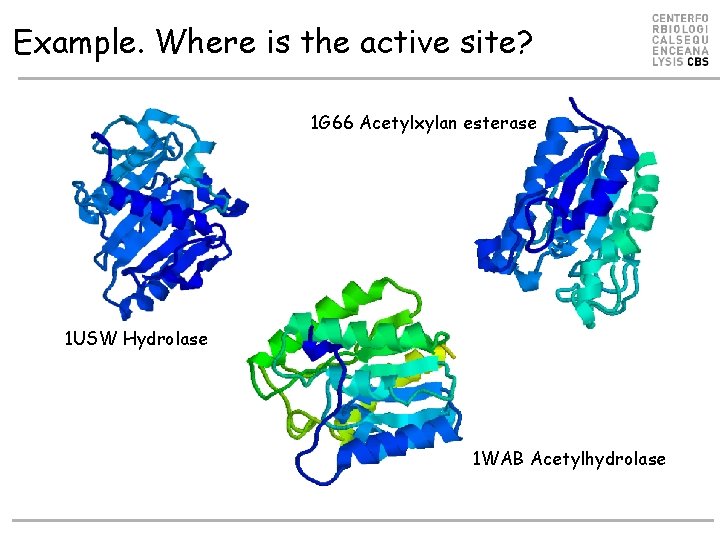
Example. Where is the active site? 1 G 66 Acetylxylan esterase 1 USW Hydrolase 1 WAB Acetylhydrolase

Example. Where is the active site? • Align sequence against structures of known acetylesterase, like • 1 WAB, 1 FXW, … • Cannot be aligned. Too low sequence similarity 1 K 7 C. A 1 WAB. _ RMSD 11. 2397 QAL 1 K 7 C. A 71 GHNDGGSLSTDNGRTDCSGTGAEVCYSVYDGVNETILTF DAL 1 WAB. _ 160 GHPRAHFLDADPGFVHSDGTISH--HDMYDYLHLSRLGY

!!! Is it really impossible? W RO N G! !!! Protein homology modeling is only possible if %id greater than 30 -50%
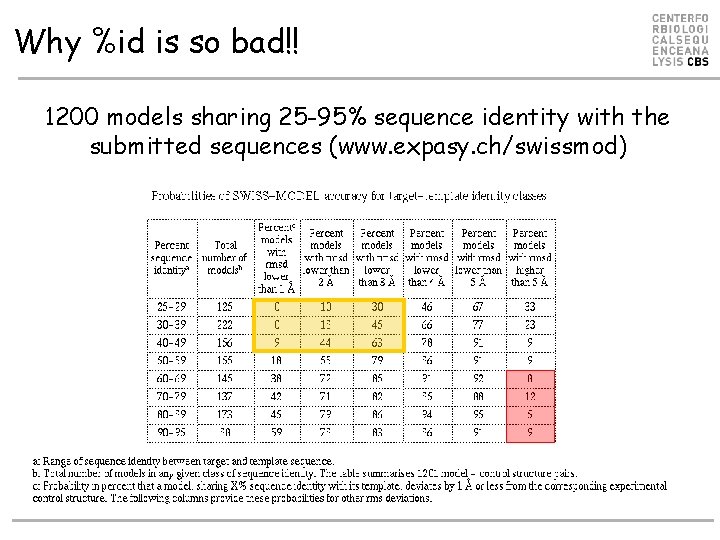
Why %id is so bad!! 1200 models sharing 25 -95% sequence identity with the submitted sequences (www. expasy. ch/swissmod)

Identification of correct fold • % ID is a poor measure – Many evolutionary related proteins share low sequence homology – A short alignment of 5 amino acids can share 100% id, what does this mean? • Alignment score even worse – Many sequences will score high against every thing (hydrophobic stretches) • P-value or E-value more reliable
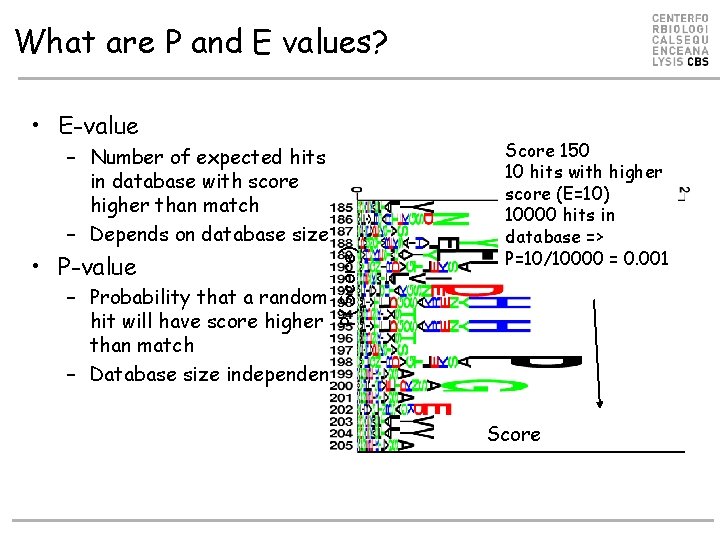
What are P and E values? – Number of expected hits in database with score higher than match – Depends on database size • P-value P(Score) • E-value Score 150 10 hits with higher score (E=10) 10000 hits in database => P=10/10000 = 0. 001 – Probability that a random hit will have score higher than match – Database size independent Score

What goes wrong when Blast fails? • Conventional sequence alignment uses a (Blosum) scoring matrix to identify amino acids matches in the two protein sequences
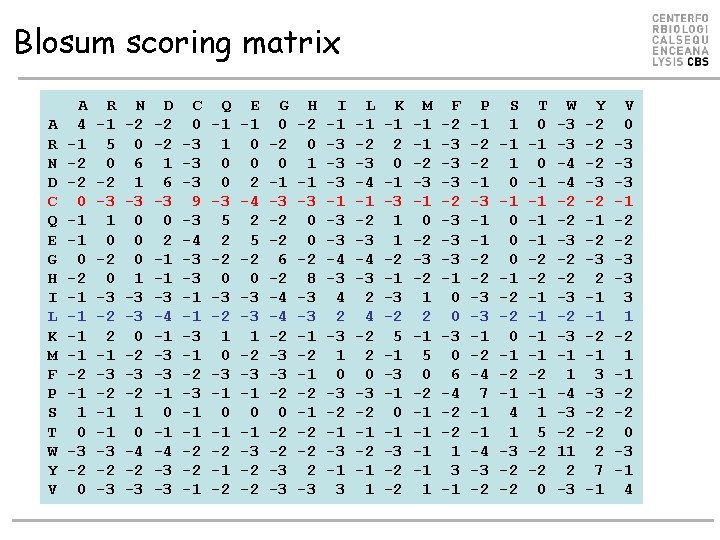
Blosum scoring matrix A R N D C Q E G H I L K M F P S T W Y V A 4 -1 -2 -2 0 -1 -1 0 -2 -1 -1 -2 -1 1 0 -3 -2 0 R -1 5 0 -2 -3 1 0 -2 0 -3 -2 2 -1 -3 -2 -1 -1 -3 -2 -3 N -2 0 6 1 -3 0 0 0 1 -3 -3 0 -2 -3 -2 1 0 -4 -2 -3 D -2 -2 1 6 -3 0 2 -1 -1 -3 -4 -1 -3 -3 -1 0 -1 -4 -3 -3 C 0 -3 -3 -3 9 -3 -4 -3 -3 -1 -1 -3 -1 -2 -3 -1 -1 -2 -2 -1 Q -1 1 0 0 -3 5 2 -2 0 -3 -2 1 0 -3 -1 0 -1 -2 E -1 0 0 2 -4 2 5 -2 0 -3 -3 1 -2 -3 -1 0 -1 -3 -2 -2 G 0 -2 0 -1 -3 -2 -2 6 -2 -4 -4 -2 -3 -3 -2 0 -2 -2 -3 -3 H -2 0 1 -1 -3 0 0 -2 8 -3 -3 -1 -2 -2 2 -3 I -1 -3 -3 -3 -1 -3 -3 -4 -3 4 2 -3 1 0 -3 -2 -1 -3 -1 3 L -1 -2 -3 -4 -3 2 4 -2 2 0 -3 -2 -1 1 K -1 2 0 -1 -3 1 1 -2 -1 -3 -2 5 -1 -3 -1 0 -1 -3 -2 -2 M -1 -1 -2 -3 -1 0 -2 -3 -2 1 2 -1 5 0 -2 -1 -1 1 F -2 -3 -3 -3 -1 0 0 -3 0 6 -4 -2 -2 1 3 -1 P -1 -2 -2 -1 -3 -1 -1 -2 -2 -3 -3 -1 -2 -4 7 -1 -1 -4 -3 -2 S 1 -1 1 0 -1 0 0 0 -1 -2 -2 0 -1 -2 -1 4 1 -3 -2 -2 T 0 -1 -1 -2 -2 -1 -1 -2 -1 1 5 -2 -2 0 W -3 -3 -4 -4 -2 -2 -3 -1 1 -4 -3 -2 11 2 -3 Y -2 -2 -2 -3 -2 -1 -2 -3 2 -1 -1 -2 -1 3 -3 -2 -2 2 7 -1 V 0 -3 -3 -3 -1 -2 -2 -3 -3 3 1 -2 1 -1 -2 -2 0 -3 -1 4

What goes wrong when Blast fails? • Conventional sequence alignment uses a (Blosum) scoring matrix to identify amino acids matches in the two protein sequences • This scoring matrix is identical at all positions in the protein sequence! EVVFIGDSLVQLMHQC A G D S. G G G D S X X X

Alignment accuracy. Scoring functions • Blosum 62 score matrix. Fg=1. Ng=0? L A G D S D F 0 -2 -3 -3 -2 -3 I 2 1 -4 -3 -2 -3 G -4 0 6 -1 0 -1 D -4 -2 -1 6 0 6 S -2 1 0 0 4 0 L 4 -1 -4 -4 -2 -4 • Score =2+6+6+4 -1=17 • Alignment LAGDS I-GDS
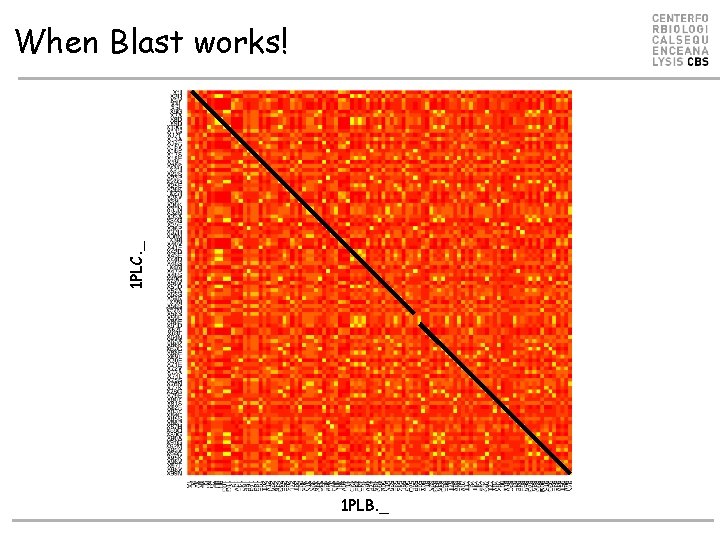
1 PLC. _ When Blast works! 1 PLB. _
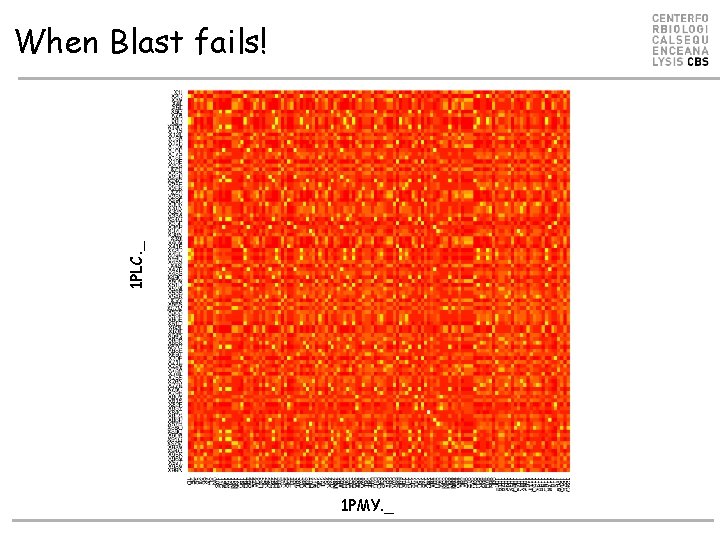
1 PLC. _ When Blast fails! 1 PMY. _

When Blast fails, use sequence profiles!
1 PLC. _ When Blast fails, use sequence profiles! 1 PMY. _

Sequence profiles • In reality not all positions in a protein are equally likely to mutate • • • Some amino acids (active cites) are highly conserved, and the score for mismatch must be very high Other amino acids can mutate almost for free, and the score for mismatch should be lower than the BLOSUM score Sequence profiles can capture these differences
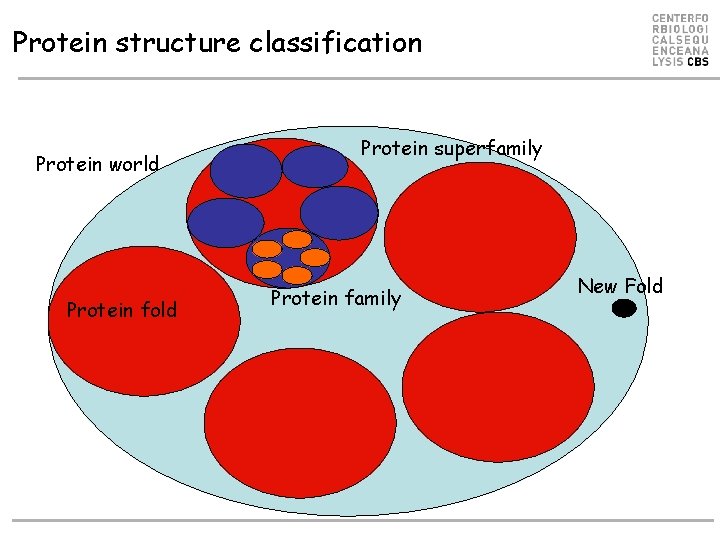
Protein structure classification Protein world Protein fold Protein superfamily Protein family New Fold
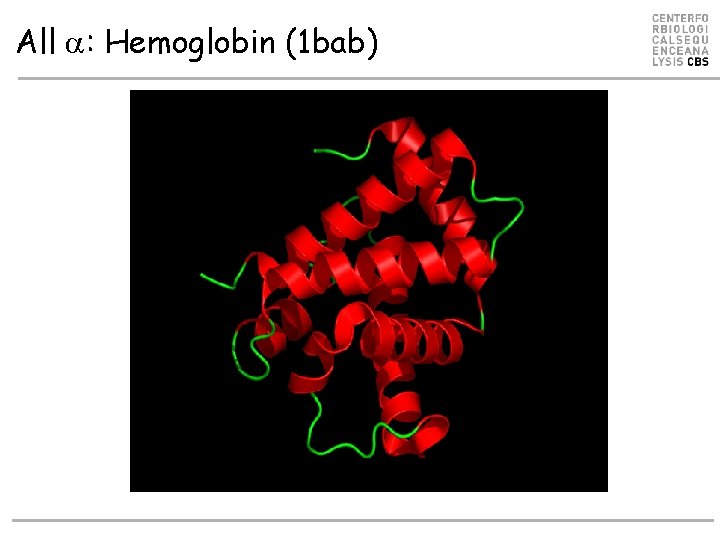
All a: Hemoglobin (1 bab)
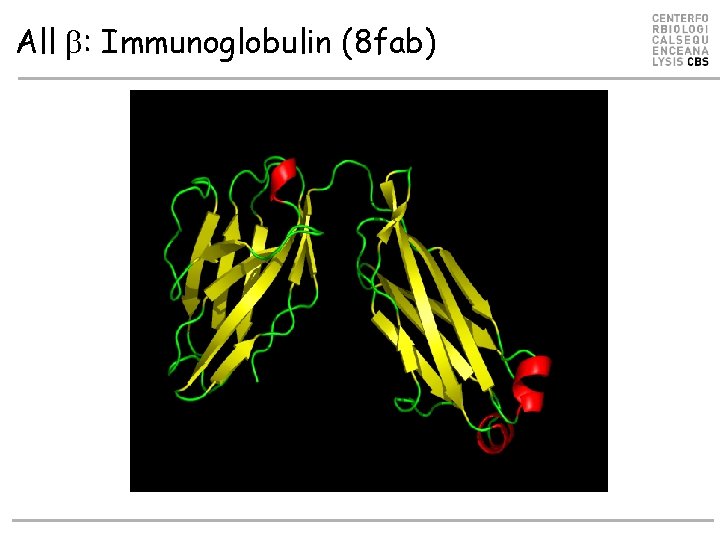
All b: Immunoglobulin (8 fab)
a/b: Triose phosphate isomerase (1 hti)
a+b: Lysozyme (1 jsf)

Sequence profiles d ed erv s n Co e erv ns o n-c No ADDGSLAFVPSEF--SISPGEKIVFKNNAGFPHNIVFDEDSIPSGVDASKISMSEEDLLN TVNGAI--PGPLIAERLKEGQNVRVTNTLDEDTSIHWHGLLVPFGMDGVPGVSFPG---I -TSMAPAFGVQEFYRTVKQGDEVTVTIT-----NIDQIED-VSHGFVVVNHGVSME---I IE--KMKYLTPEVFYTIKAGETVYWVNGEVMPHNVAFKKGIV--GEDAFRGEMMTKD---TSVAPSFSQPSF-LTVKEGDEVTVIVTNLDE------IDDLTHGFTMGNHGVAME---V ASAETMVFEPDFLVLEIGPGDRVRFVPTHK-SHNAATIDGMVPEGVEGFKSRINDE---TVNGQ--FPGPRLAGVAREGDQVLVKVVNHVAENITIHWHGVQLGTGWADGPAYVTQCPI TKAVVLTFNTSVEICLVMQGTSIV----AAESHPLHLHGFNFPSNFNLVDPMERNTAGVP Matching any thing but G => large negative score Any thing can match

How to make sequence profiles • • Align (BLAST) sequence against large sequence database (Swiss-Prot) Select significant alignments and make profile (weight matrix) using techniques for sequence weighting and pseudo counts Use weight matrix to align against sequence database to find new significant hits Repeat 2 and 3 (normally 3 times!)
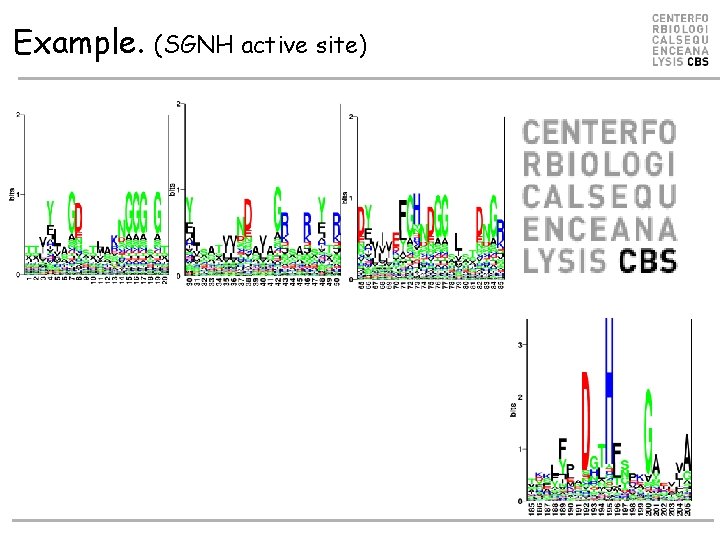
Example. (SGNH active site)
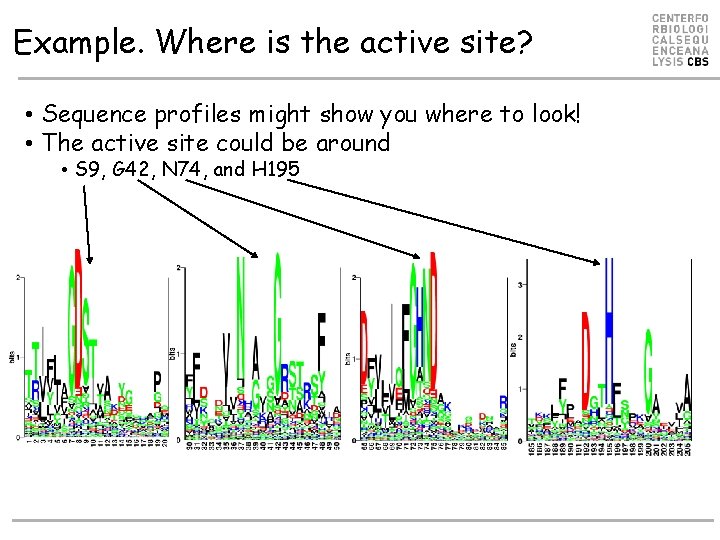
Example. Where is the active site? • Sequence profiles might show you where to look! • The active site could be around • S 9, G 42, N 74, and H 195
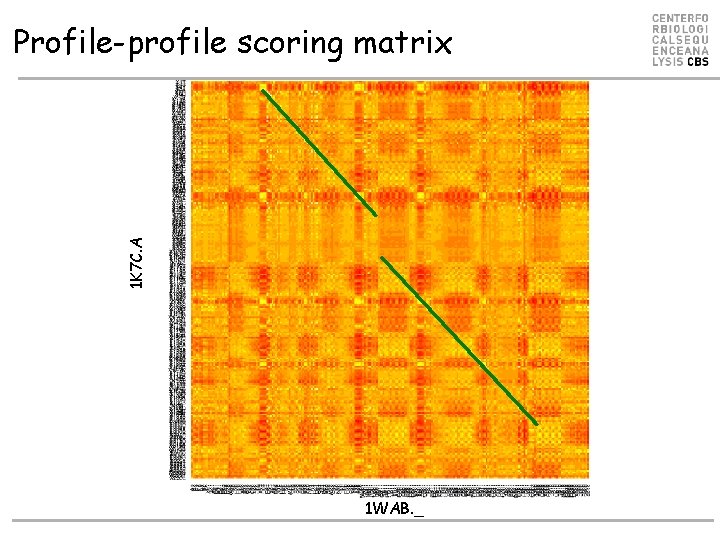
1 K 7 C. A Profile-profile scoring matrix 1 WAB. _

Example. Where is the active site? Align using sequence profiles ALN 1 K 7 C. A 1 WAB. _ RMSD = 5. 29522. 14% ID 1 K 7 C. A TVYLAGDSTMAKNGGGSGTNGWGEYLASYLSATVVNDAVA GRSARSYTREGRFENIADVVTAGDYVIVEFGH NDGGSLSTDN S G N 1 WAB. _ EVVFIGDSLVQLMHQCE---IWRELFS---PLHALNFGIG GDSTQHVLW--RLENGELEHIRPKIVVVWVGT NNHG-----1 K 7 C. A GRTDCSGTGAEVCYSVYDGVNETILTFPAYLENAAKLFTAK--GAKVILSSQTPNNPWETGTFVNSPTRFVEYAEL-AAEVA 1 WAB. _ -----------HTAEQVTGGIKAIVQLVNERQPQARVVVLGLLPRGQ-HPNPLREKNRRVNELVRAALAGHP 1 K 7 C. A GVEYVDHWSYVDSIYETLGNATVNSYFPIDHT HTSPAGAEVVAEAFLKAVVCTGTSL H 1 WAB. _ RAHFLDADPG---FVHSDG--TISHHDMYDYL HLSRLGYTPVCRALHSLLLRL---L
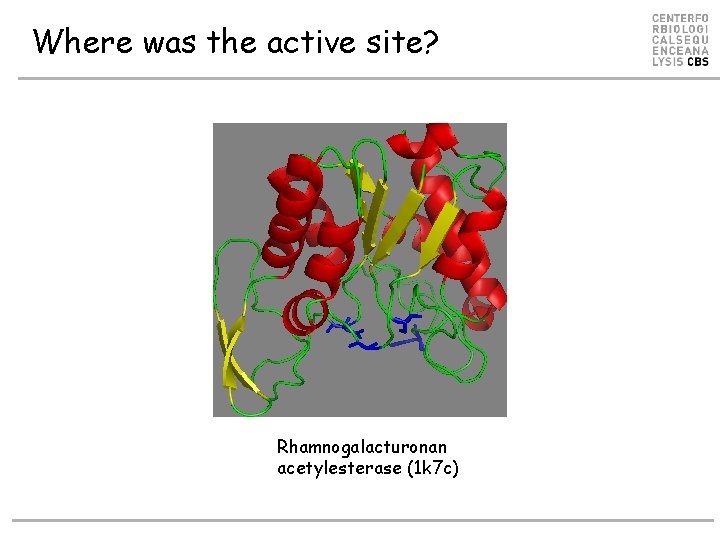
Where was the active site? Rhamnogalacturonan acetylesterase (1 k 7 c)

How good are we?

Alignment accuracy

AUC performance measure Query 1 CJ 0. A 1 CJ 0. A Templ 1 B 78. A 1 B 8 A. A 1 B 8 B. A 1 B 8 G. A 1 B 9 H. A 1 BAR. A 1 BAV. C Score Hit/nonhit 0. 170963 0 -0. 040029 0 -0. 012789 0 12. 342823 1 13. 394361 1 -1. 281068 0 -1. 091305 0 Query 1 CJ 0. A 1 CJ 0. A Templ 1 B 8 G. A 1 DTY. A 1 DGD. _ 1 GTX. A 2 GSA. A 1 BW 9. A 1 AUP. _ 1 GTM. A Score Hit/nonhit 12. 342823 1 11. 867786 1 11. 271914 1 11. 010288 1 10. 958170 1 2. 651775 0 2. 507336 1 2. 444512 0 AUC (area under the ROC curve)

Fold recognition performance
Outlook • Include position dependent gap penalties • The conventional alignment methods use equal gap penalties through out the scoring matrix • In real proteins placement of insertions and deletions is highly structure dependent • No gaps in secondary structure elements • Gaps most frequent in loops • Distance dependency

Take home message • Identifying the correct fold is only a small step towards successful homology modeling • Do not trust % ID or alignment score to identify the fold. Use P-values • You can do reliable fold recognition AND homology modeling when for low sequence homology • Use sequence profiles and local protein structure to align sequences

What are (some of) the different available methods? • Simple sequence based methods – Align (BLAST) sequence against sequence of proteins with known structure (PDB database) • Sequence profile based methods – Align sequence profile (Psi-BLAST) against sequence of proteins with known structure (PDB, FUGUE) – Align sequence profile against profile of proteins with known structure (FFAS) • Sequence and structure based methods – Align profile and predicted secondary structure against proteins with known structure (3 D-PSSM, Phyre) • Sequence profiles and structure based methods – HHpred
- Slides: 42