Protein Feature Identification David Wishart Depts Computing Biological

Protein Feature Identification David Wishart Depts. Computing & Biological Science University of Alberta david. wishart@ualberta. ca

Proteins • Exhibit far more sequence and chemical complexity than DNA or RNA • Properties and structure are defined by the sequence and side chains of their constituent amino acids • The “engines” of life • >95% of all drugs target proteins • Favorite topic of post-genomic era

The Post-genomic Challenge • • How to rapidly identify a protein? How to rapidly purify a protein? How to identify post-trans modification? How to find information about function? How to find information about activity? How to find information about location? How to find information about structure? Answer: Look at Protein Features
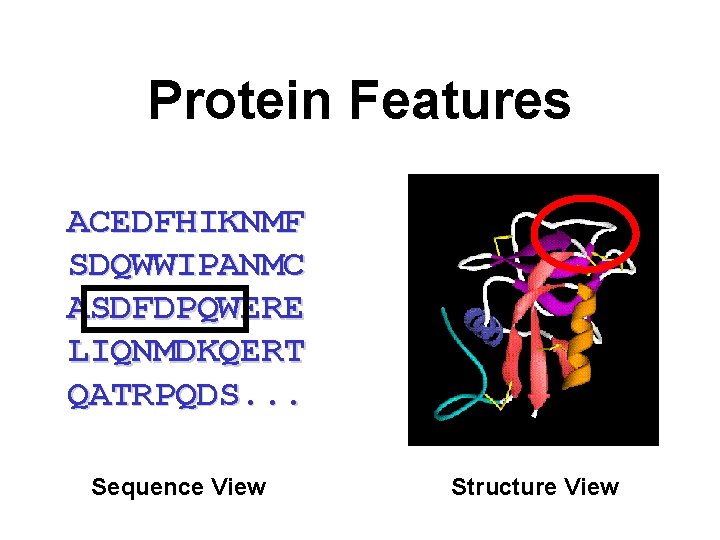
Protein Features ACEDFHIKNMF SDQWWIPANMC ASDFDPQWERE LIQNMDKQERT QATRPQDS. . . Sequence View Structure View

Different Types of Features • Composition Features – Mass, p. I, Absorptivity, Rg, Volume • Sequence Features – Active sites, Binding Sites, Targeting, Location, Property Profiles, 2 o structure • Structure Features – Supersecondary Structure, Global Fold, ASA, Volume

Where To Go http: //www. expasy. org/

Amino Acids (Review)
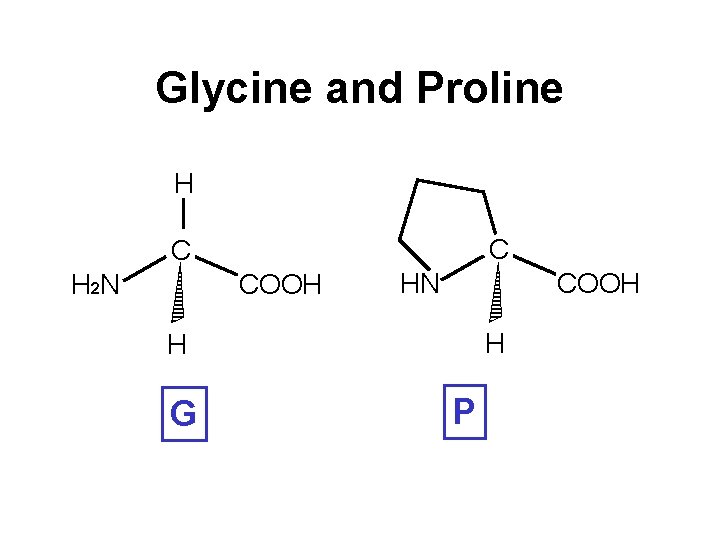
Glycine and Proline H C C H 2 N COOH H H G P
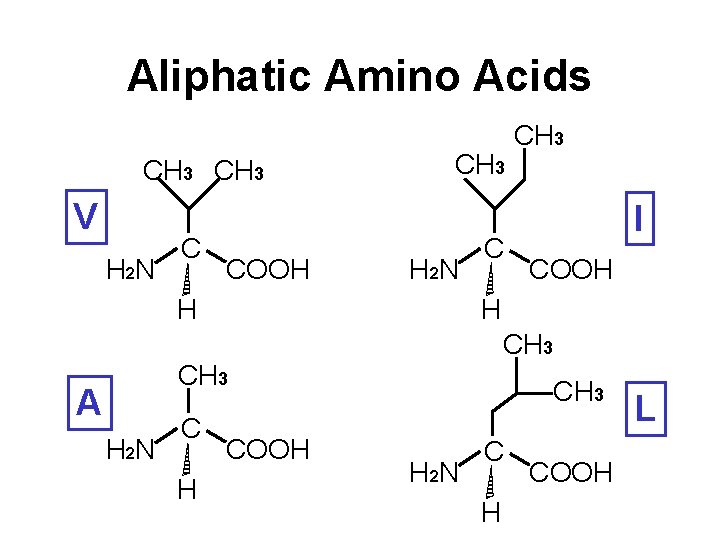
Aliphatic Amino Acids CH 3 V H 2 N C COOH CH 3 H 2 N H C H 2 N C H I COOH H CH 3 A CH 3 COOH CH 3 H 2 N C H COOH L
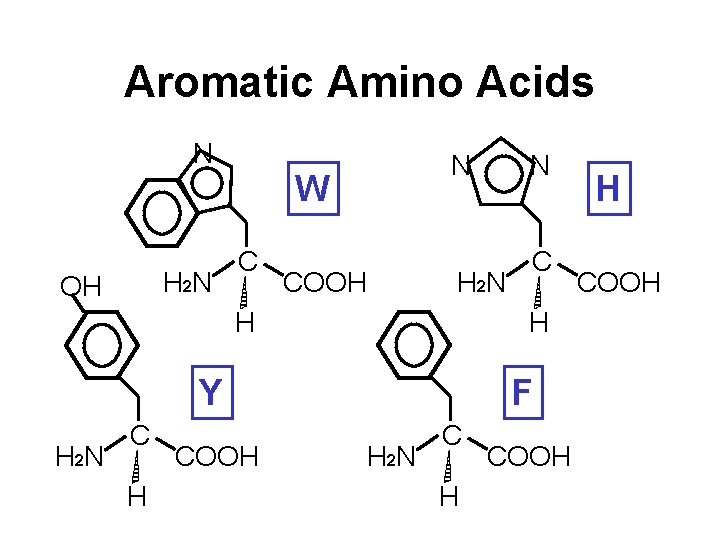
Aromatic Amino Acids N H 2 N OH N W C COOH H 2 N H COOH C H Y C N F H 2 N C H COOH
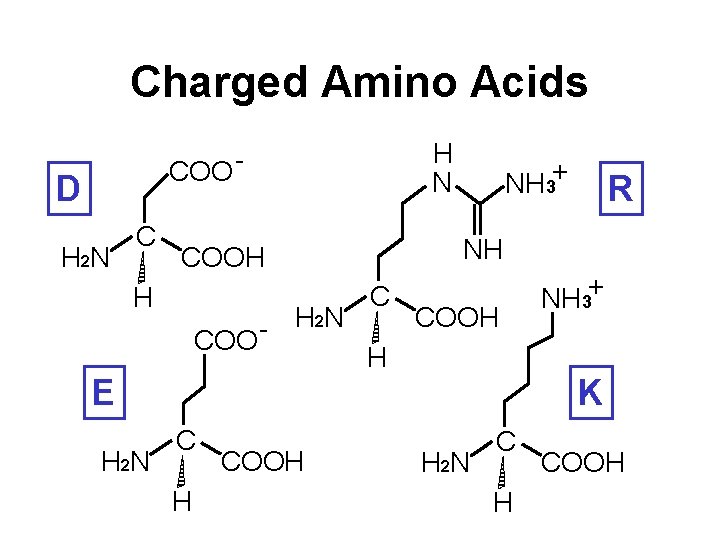
Charged Amino Acids H N COO - D H 2 N COO E H 2 N C H COOH R NH COOH H NH 3+ C COOH H NH 3+ K H 2 N C H COOH
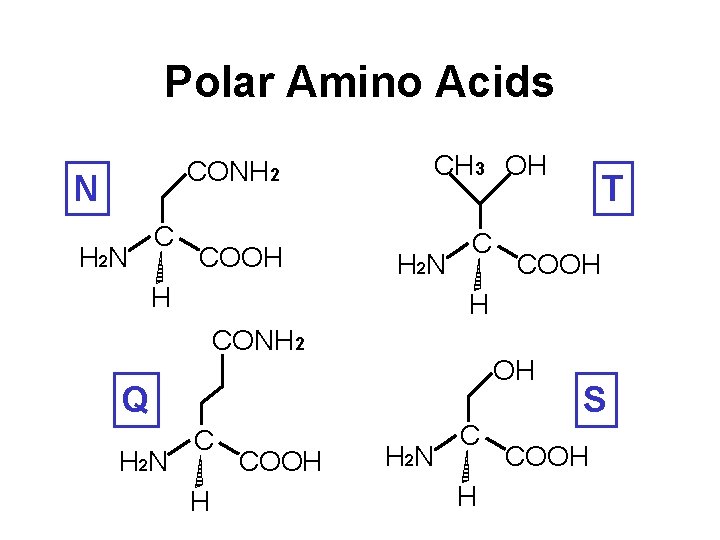
Polar Amino Acids CONH 2 N H 2 N C COOH CH 3 OH H 2 N H C COOH H CONH 2 OH Q H 2 N T C H COOH H 2 N C H S COOH
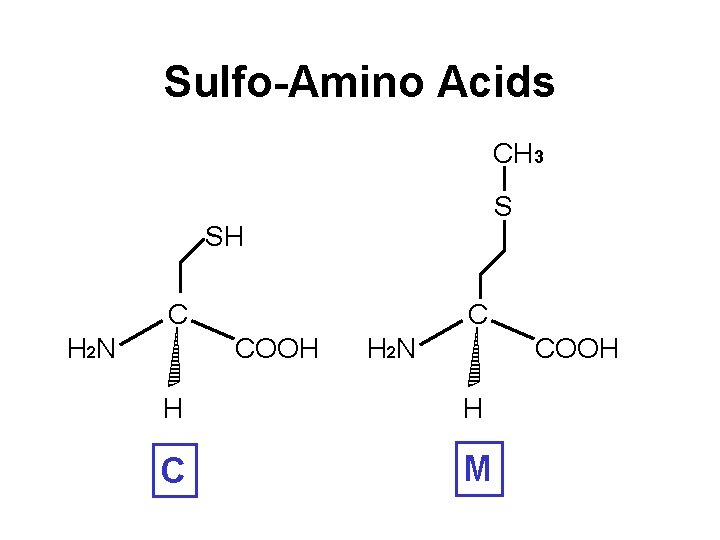
Sulfo-Amino Acids CH 3 S SH C H 2 N C COOH H 2 N COOH H H C M

Compositional Features • • Molecular Weight Amino Acid Frequency Isoelectric Point UV Absorptivity Solubility, Size, Shape Radius of Gyration Free Energy of Folding
Molecular Weight

Molecular Weight • • • Useful for SDS PAGE and 2 D gel analysis Useful for deciding on SEC matrix Useful for deciding on MWC for dialysis Essential in synthetic peptide analysis Essential in peptide sequencing (classical or mass-spectrometry based) • Essential in proteomics and high throughput protein characterization

Molecular Weight • Crude MW calculation: MW = 110 X Numres • Exact MW calculation: MW = SAAi x MWi • Remember to add 1 water (18. 01 amu) after adding all res. • Note isotopic weights • Corrections for CHO, PO 4, Acetyl, CONH 2
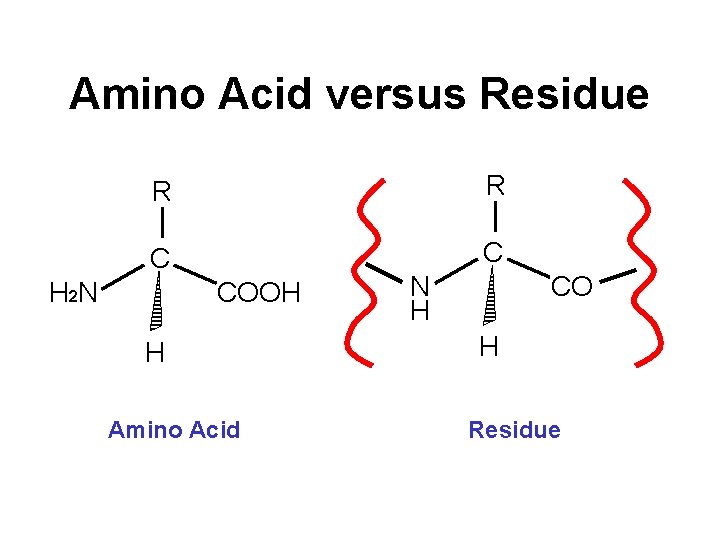
Amino Acid versus Residue R R C C H 2 N COOH H Amino Acid N H CO H Residue

Protein Identification via MW • MOWSE • http: //srs. hgmp. mrc. ac. uk/cgi-bin/mowse • Comb. Search • http: //ca. expasy. org/tools/Comb. Search/ • Mascot • http: //www. matrixscience. com/search_form _select. html • AAComp. Sim/AAComp. Ident • http: //ca. expasy. org/tools/
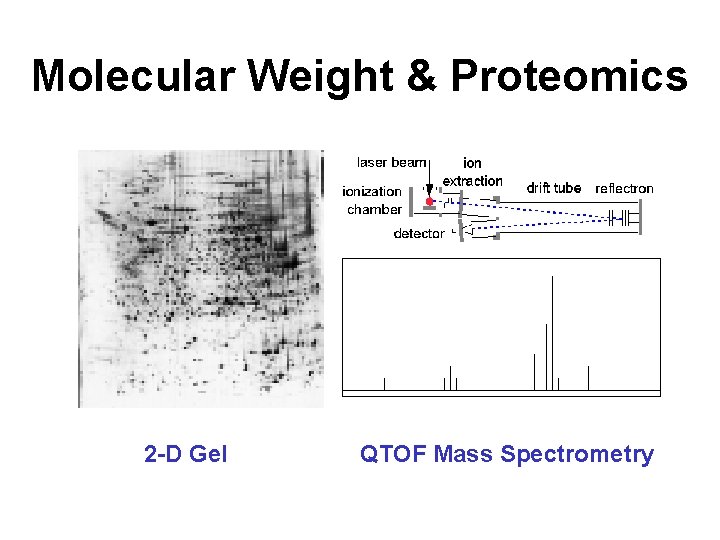
Molecular Weight & Proteomics 2 -D Gel QTOF Mass Spectrometry

Amino Acid Frequency • Deviations greater than 2 X average indicate something of interest • High K or R indicates possible nucleoprotein • High C’s indicate stable but hard-to-fold protein • High G, P, Q, or N says lack of stable structure
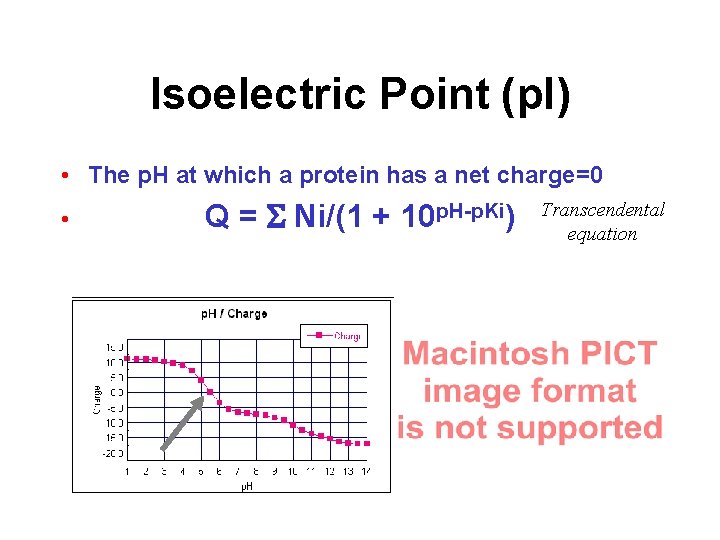
Isoelectric Point (p. I) • The p. H at which a protein has a net charge=0 • Q = S Ni/(1 + 10 p. H-p. Ki) Transcendental equation

Isoelectric Point • Calculation is only approximate (+/- 1 p. H) • Does not include 3 o structure interactions • Can be used in developing purification protocols via ion exchange chromatography • Can be used in estimating spot location for isoelectric focusing gels • Can be used to decide on best p. H to store or analyze protein

UV Spectroscopy

UV Absorptivity • UV (Ultraviolet light) has a wavelength of 200 to 400 nm • Most proteins and peptides (and all nucleic acids) absorb UV light quite strongly • UV spectroscopy is the most common form of spectroscopy performed today • UV spectra can be used to identify or classify some proteins or protein classes
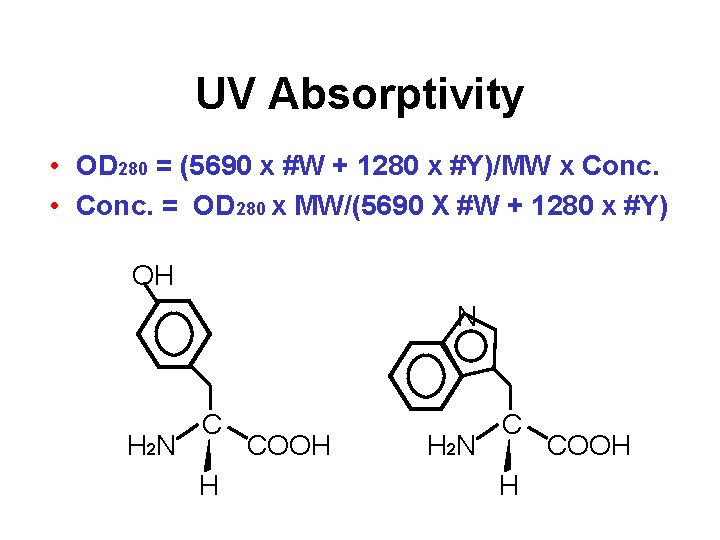
UV Absorptivity • OD 280 = (5690 x #W + 1280 x #Y)/MW x Conc. • Conc. = OD 280 x MW/(5690 X #W + 1280 x #Y) OH N H 2 N C H COOH

Hydrophobicity • Indicates Solubility • Indicates Stability • Indicates Location (membrane or cytoplasm) • Indicates Globularity or tendency to form spherical structure

Hydrophobicity • Average Hydrophobicity AH = S AAi x Hi • Hydrophobic Ratio RH = S H(-)/S H(+) • Hydrophobic % Ratio RHP = %philic/%phobic • Linear Charge Density LIND = (K+R+D+E+H+2)/# • Solubility SOL = RH + LIND - 0. 05 AH • Average AH = 2. 5 + 2. 5 Insol > 0. 1 Unstrc < -6 • Average RH = 1. 2 + 0. 4 Insol < 0. 8 Unstrc > 1. 9 • Average RHP = 0. 9 + 0. 2 Insol < 0. 7 Unstrc > 1. 4 • Average LIND = 0. 25 Insol < 0. 2 Unstrc > 0. 4 • Average SOL = 1. 6 + 0. 5 Insol < 1. 1 Unstrc > 2. 5

Protein Dimensions • Radius and Radius of Gyration • Molecular and Partial Specific Volume • Accessible Surface Area • Provides a size estimate of a protein • Used in analytical techniques such as neutron or X-ray scattering, analytical ultracentrifugation, light scattering

Radius & Radius of Gyration • RAD = 3. 875 x NUMRES 0. 333 (Folded) • RADG = 0. 41 x (110 x NUMRES) 0. 5 Radius (Unfolded) Radius of Gyration

Partial Specific Volume • Measured in m. L/g • Inverse measure of protein density (0. 70 -75) • Depends on protein’s composition and compactness • Measured via sedimentation analysis • PSV = S PSi x Wi

Packing Volume Loose Packing Dense Packing Proteins are Densely Packed

Packing Volume (VP) • Determined via X-ray or NMR structure • “True” measure of volume occupied by protein • Approximate Value VP = 1. 245 x MW • Exact Value VP = S AAi x Vi

Different Types of Features • Composition Features – Mass, p. I, Absorptivity, Rg, Volume • Sequence Features – Active sites, Binding Sites, Targeting, Location, Property Profiles, 2 o structure • Structure Features – Supersecondary Structure, Global Fold, ASA, Volume
![Sequence Features AHGQSDFILDEADGMMKSTVPN… HGFDSAAVLDEADHILQWERTY… GGGNDEYIVDEADSVIASDFGH… *[LIVM]DEAD*[LIVM]* (EIF 4 A ATP DEPENDENT HELICASE) Sequence Features AHGQSDFILDEADGMMKSTVPN… HGFDSAAVLDEADHILQWERTY… GGGNDEYIVDEADSVIASDFGH… *[LIVM]DEAD*[LIVM]* (EIF 4 A ATP DEPENDENT HELICASE)](http://slidetodoc.com/presentation_image/3c84c3ec745422e7cd581fa52b310ace/image-35.jpg)
Sequence Features AHGQSDFILDEADGMMKSTVPN… HGFDSAAVLDEADHILQWERTY… GGGNDEYIVDEADSVIASDFGH… *[LIVM]DEAD*[LIVM]* (EIF 4 A ATP DEPENDENT HELICASE)

Probability & Seq. Features • Expectation value (e) is the expected number of hits for a given sequence pattern or motif • e = N x f 1 x f 2 x f 3 x. . fk • N is the number of residues in DB (108) • fi is the frequency of a given amino acid(s)

Example #1 ACIDS e = 108*0. 088*0. 021*0. 054*0. 059*0. 065 e = 38. 3 #Found in OWL database = 14
![Example #2 A*ACI[DEN]S e = 108*0. 088*1. 000*0. 088*0. 021*0. 054 *{0. 059 + Example #2 A*ACI[DEN]S e = 108*0. 088*1. 000*0. 088*0. 021*0. 054 *{0. 059 +](http://slidetodoc.com/presentation_image/3c84c3ec745422e7cd581fa52b310ace/image-38.jpg)
Example #2 A*ACI[DEN]S e = 108*0. 088*1. 000*0. 088*0. 021*0. 054 *{0. 059 + 0. 046}*0. 065 e = 9. 4 #Found in OWL database = 9

Minimum Pattern Lengths f = 0. 08 e = 108*0. 088 = 0. 17 min = 8 f = 0. 05 e = 108*0. 057 = 0. 08 min = 7 f = 0. 03 e = 108*0. 036 = 0. 07 min = 6

How Long Should a Sequence Motif or Sequence Block Be? • How many matching segments of length “l” could be found in comparing a query of length M to a DB of N ? • Answer: n(l) = M x N x fl • Assume f = 0. 05, M = 300, N = 100, 000

Rule of Thumb Make your protein sequence motifs at least 8 residues long

Sites that Support Pattern Queries • OWL Database – http: //bioinf. man. ac. uk/dbbrowser/OWL/ • PIR Website – http: //pir. georgetown. edu/pirwww/search/patmatch. html • SCNPSITE at EXPASY – http: //ca. expasy. org/tools/scanprosite/ • FPAT (Regular Expression Query) – http: //stateslab. bioinformatics. med. umich. edu/service/fpat/
![Regular Expressions • C[ACG]T - Matches CAT, CCT and CGT only • C. T Regular Expressions • C[ACG]T - Matches CAT, CCT and CGT only • C. T](http://slidetodoc.com/presentation_image/3c84c3ec745422e7cd581fa52b310ace/image-43.jpg)
Regular Expressions • C[ACG]T - Matches CAT, CCT and CGT only • C. T - Matches CAT, Ca. T, C 1 T, CXT, not CT • CA? T - Matches CT or CAT only • C+T - Matches CT, CCCT, CCCCT… • C(HE)? A[TP] - Matches CHEAT, CHEAP, CAP • S[A-I, L-Q, T-Z]? LK[A-I, L-Q, T-Z]? A - Matches S*LK*A
![PROSITE Pattern Expressions C - [ACG] - T - Matches CAT, CCT and CGT PROSITE Pattern Expressions C - [ACG] - T - Matches CAT, CCT and CGT](http://slidetodoc.com/presentation_image/3c84c3ec745422e7cd581fa52b310ace/image-44.jpg)
PROSITE Pattern Expressions C - [ACG] - T - Matches CAT, CCT and CGT only C - X -T - Matches CAT, CCT, CDT, CET, etc. C - {A} -T - Matches every CXT except CAT C - (1, 3) - T - Matches CT, CCCT C - A(2) - [TP] - Matches CAAT, CAAP [LIV] - [VIC] - X(2) - G - [DENQ] - X - [LIVFM] (2) -G

Sequence Feature Databases • PROSITE - http: //ca. expasy. org/prosite/ • BLOCKS - http: //www. blocks. fhcrc. org/ • DOMO - http: //www. infobiogen. fr/services/domo/ • PFAM - http: //pfam. wustl. edu • PRINTS - http: //www. bioinf. man. ac. uk/dbbrowser/PRINTS/ • SEQSITE - Pep. Tool
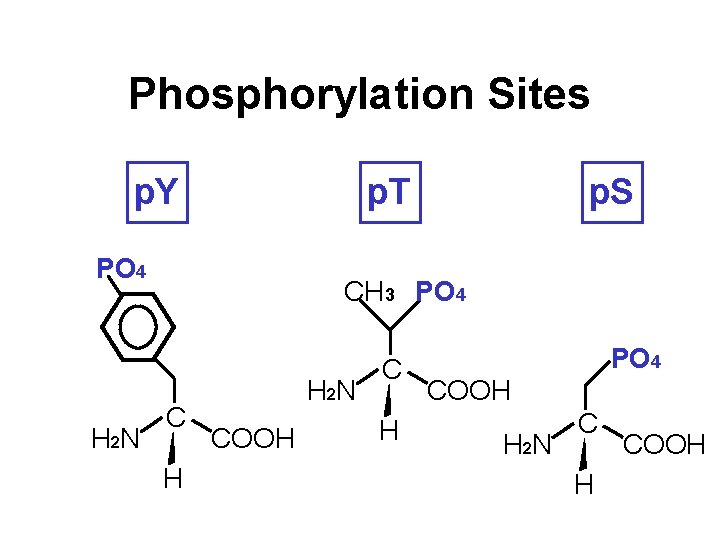
Phosphorylation Sites p. Y p. T PO 4 H 2 N p. S CH 3 PO 4 C H H 2 N COOH C H PO 4 COOH H 2 N C H COOH

Phosphorylation Sites
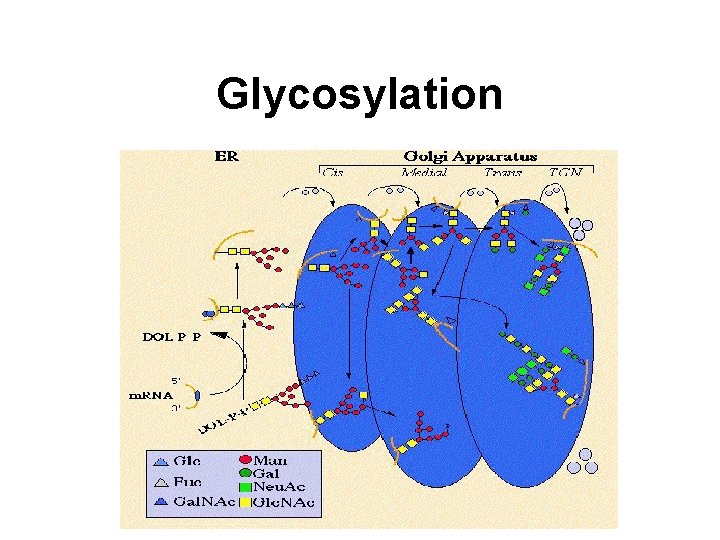
Glycosylation

Glycosylation Sites
Signaling

Signaling Sites
Protease Cut Sites

Protease Cut Sites

Binding Sites

Family Signature Sequences

Enzyme Active Sites

T-Cell Epitopes • • Type I peptides are 8 - 10 amino acids Type II peptides are 12 - 20 amino acids Type I are endogenous, Type II exogenous Suggestion they are amphipathic helices • HLA-A 1 *[ED]P****[YF] • A 2. 1 ***[AVILF][AVILF]*** • HLA-DR 1 b[YF]**[ML]*[GA]**L

Better Methods for Sequence Feature ID • Sequence Profiles/Scoring Matrices • Neural Networks • Hidden Markov Models • Bayesian Belief Nets • Reference Point Logistics

A Sample Sequence Profile <e>i = log 2(qi/pi)
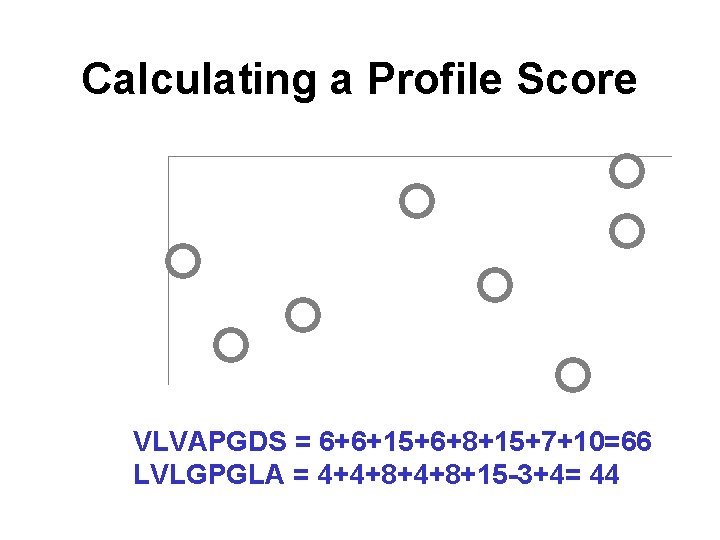
Calculating a Profile Score VLVAPGDS = 6+6+15+6+8+15+7+10=66 LVLGPGLA = 4+4+8+15 -3+4= 44
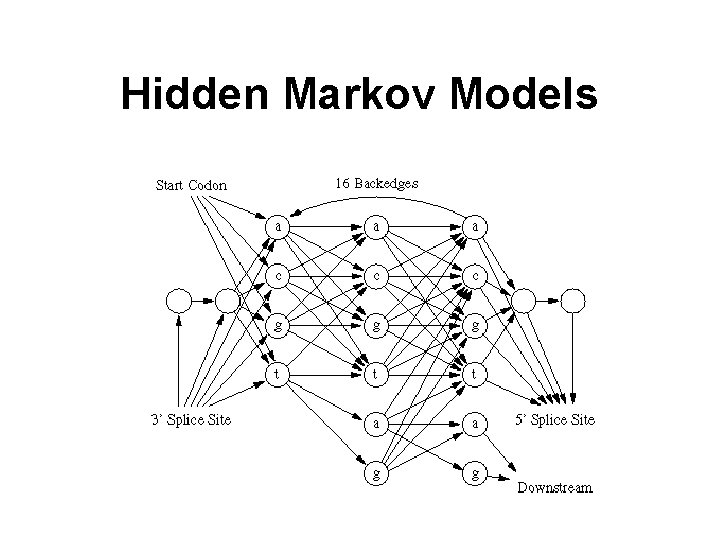
Hidden Markov Models
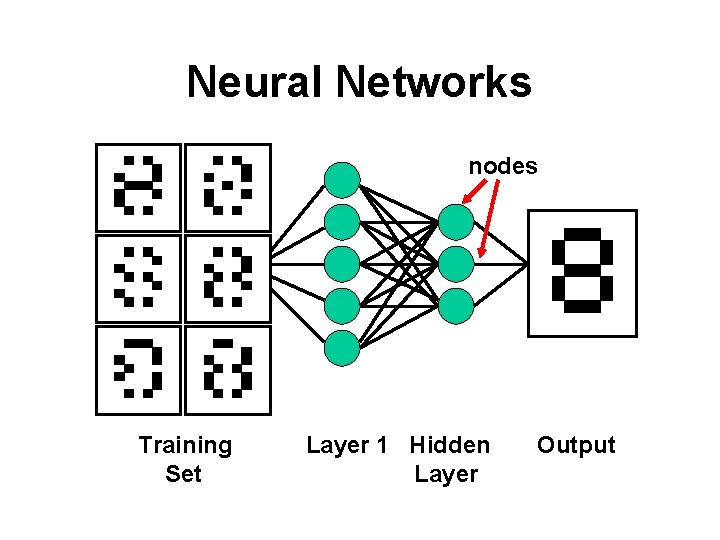
Neural Networks nodes Training Set Layer 1 Hidden Layer Output

What Can Be Predicted? • • • O-Glycosylation Sites Phosphorylation Sites Protease Cut Sites Nuclear Targeting Sites Mitochondrial Targ Sites Chloroplast Targ Sites Signal Sequence Cleav. Peroxisome Targ Sites • • • ER Targeting Sites Transmembrane Sites Tyrosine Sulfation Sites GPInositol Anchor Sites PEST sites Coil-Coil Sites T-Cell/MHC Epitopes Protein Lifetime A whole lot more….

Cutting Edge Sequence Feature Servers • Membrane Helix Prediction – http: //www. cbs. dtu. dk/services/TMHMM-2. 0/ • T-Cell Epitope Prediction – http: //syfpeithi. bmiheidelberg. com/scripts/MHCServer. dll/home. htm • O-Glycosylation Prediction – http: //www. cbs. dtu. dk/services/Net. OGlyc/ • Phosphorylation Prediction – http: //www. cbs. dtu. dk/services/Net. Phos/ • Protein Localization Prediction – http: //psort. nibb. ac. jp/

Subcellular Localization http: //www. cs. ualberta. ca/~bioinfo/PA/Sub/

Profiles & Motifs are Useful • Helped identify active site of HIV protease • Helped identify SH 2/SH 3 class of STP’s • Helped identify important GTP oncoproteins • Helped identify hidden leucine zipper in HGA • Used to scan for lectin binding domains • Regularly used to predict T-cell epitopes

Amino Acid Property Profiles

Amino Acid Property Profiles • Intent is to predict protein’s physical properties directly from sequence as opposed to composition or wet chemistry • Offers a more detailed, graphical view of sequence-specific properties than compositional analysis (more powerful? ) • Underlying assumption is: amino acid properties are additive

Property Profile Algorithm • Assign each residue a numeric value corresponding to the physical property • Choose an odd numbered window (5 or 7) and calculate the average value • Assign the average value to the middle residue in the window • Move the window down by one residue and repeat steps 1 to 4 until finished - PLOT

Common Property Profiles • Hydrophobicity (Watch Scales!) • Helical Wheel (Not a True Profile) • Hydrophobic Moments (Helix & Beta sheet) • Flexibility (Thermal B Factors) • Surface Accessibility (ASA) • Antigenicity (B-cell epitopes/T-cell epitopes)

Hydrophobicity Profile • Plotted using: <H>i = S Hn/(2 k + 1) • Shows location of membrane spanning regions, epitopes, surface exposed AA’s, etc.

Helical Wheel • Used to identify disposition of AA side chains around a helix, looking end-on • Identifies Helical Amphipathicity
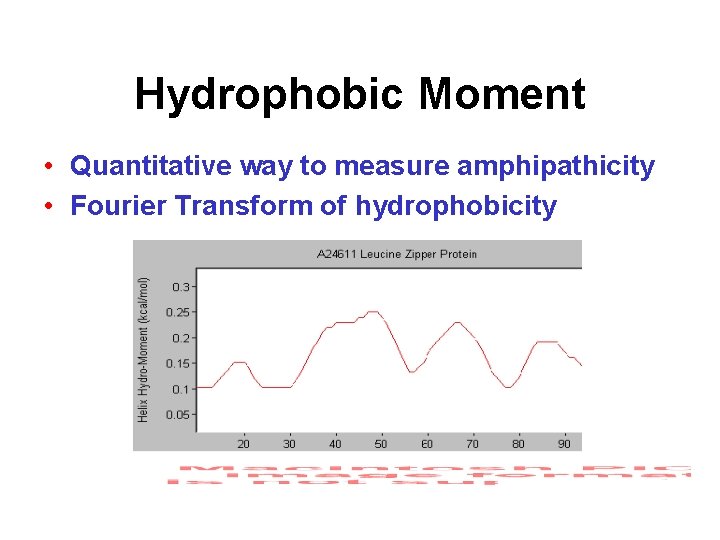
Hydrophobic Moment • Quantitative way to measure amphipathicity • Fourier Transform of hydrophobicity

Flexibility • B factors from X-ray crystallography • Potentially identifies antigenic and active sites from sequence data alone
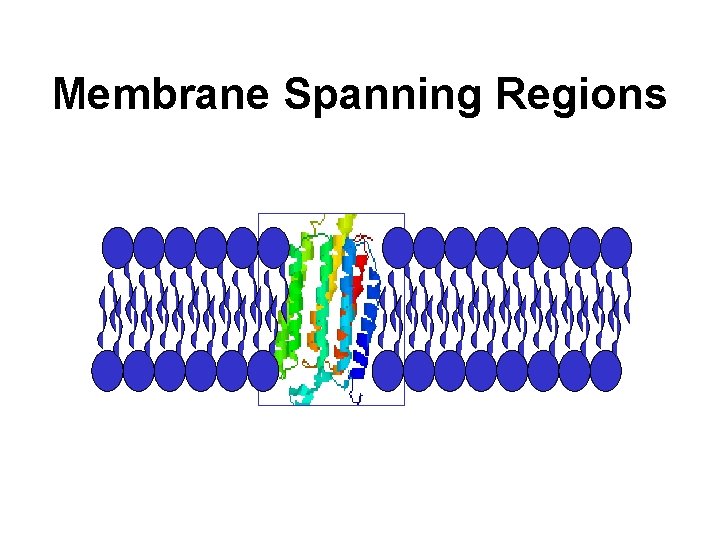
Membrane Spanning Regions

Predicting via Hydrophobicity Bacteriorhodopsin Omp. A Bacteriorhodoposin Omp. A

Predicting via Hydrophobicity 7
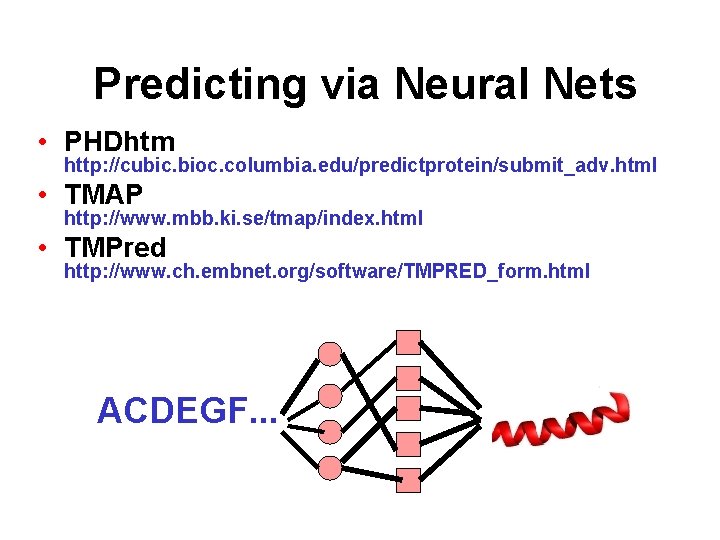
Predicting via Neural Nets • PHDhtm http: //cubic. bioc. columbia. edu/predictprotein/submit_adv. html • TMAP http: //www. mbb. ki. se/tmap/index. html • TMPred http: //www. ch. embnet. org/software/TMPRED_form. html ACDEGF. . .
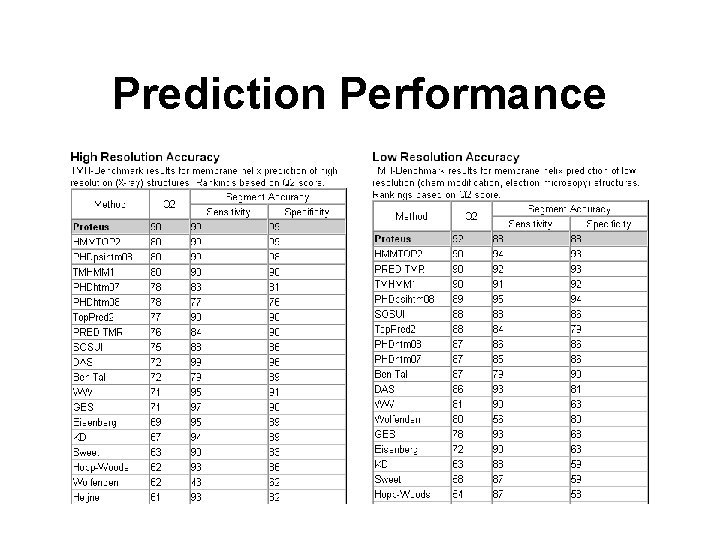
Prediction Performance
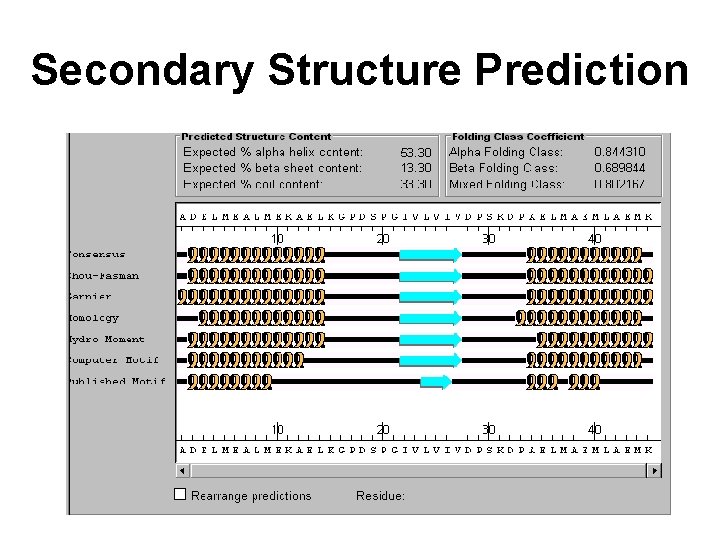
Secondary Structure Prediction

Secondary Structure Prediction • • Statistical (Chou-Fasman, GOR) Homology or Nearest Neighbor (Levin) Physico-Chemical (Lim, Eisenberg) Pattern Matching (Cohen, Rooman) Neural Nets (Qian & Sejnowski, Karplus) Evolutionary Methods (Barton, Niemann) Combined Approaches (Rost, Levin, Argos)

Chou-Fasman Statistics

The Ph. D Approach PRFILE. . .

Prediction Performance

Best of the Best • Predict. Protein-PHD (72%) – http: //cubic. bioc. columbia. edu/predictprotein • Jpred (73 -75%) – http: //www. compbio. dundee. ac. uk/~www-jpred/ • SABLE (75%) – http: //sable. chmcc. org/ • PSIpred (77%) – http: //bioinf. cs. ucl. ac. uk/psipred/ • Proteus (78 -90%) – http: //wishart. biology. ualberta. ca/proteus/index. shtml
The Proteus Server
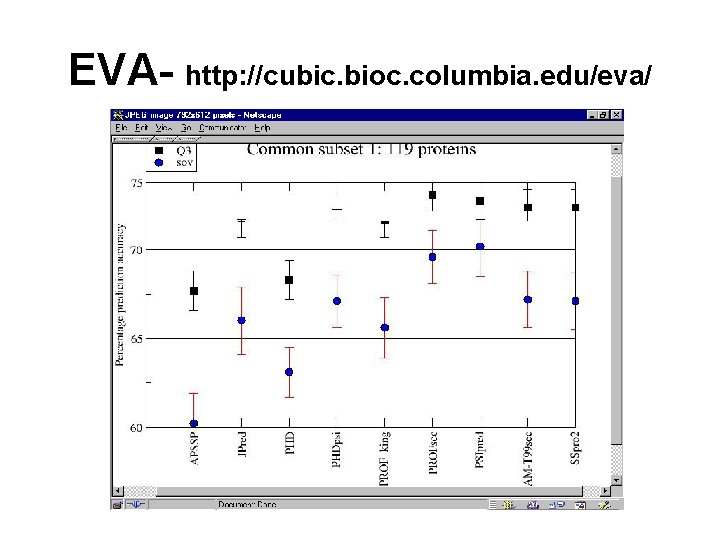
EVA- http: //cubic. bioc. columbia. edu/eva/

Different Types of Features • Composition Features – Mass, p. I, Absorptivity, Rg, Volume • Sequence Features – Active sites, Binding Sites, Targeting, Location, Property Profiles, 2 o structure • Structure Features – Supersecondary Structure, Global Fold, ASA, Volume

3 D Protein Features
Secondary Structure Table 10 - -
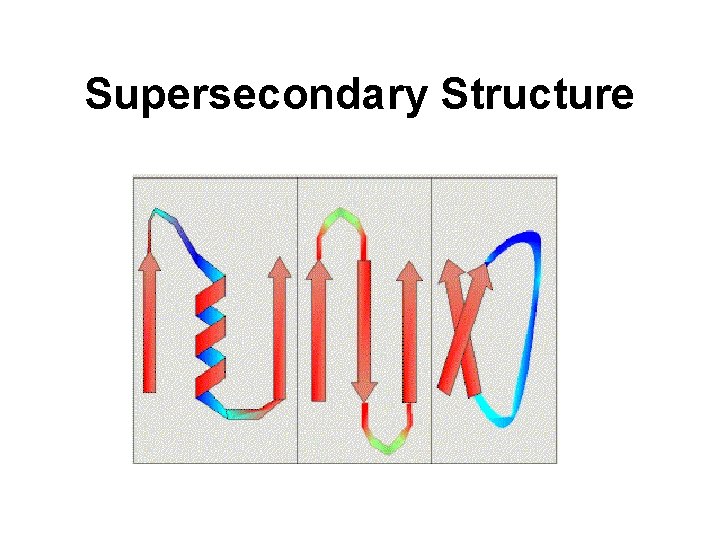
Supersecondary Structure
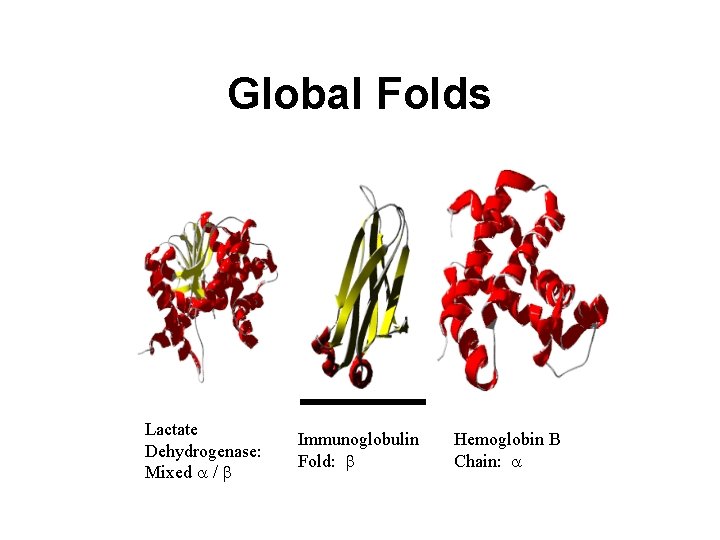
Global Folds Lactate Dehydrogenase: Mixed a / b Immunoglobulin Fold: b Hemoglobin B Chain: a

3 D Structure • Allows direct identification and/or location of cofactors, ligands, crevices, protrusions and other features • Allows one to identify possible function (through 3 D homology) • Allows protein to be classified into a folding family

3 D Structure Classifiers • CATH – http: //www. biochem. ucl. ac. uk/bsm/cath/ • VAST – http: //www. ncbi. nlm. nih. gov/Structure/VAST/va stsearch. html/ • Combinatorial Extension (CE) – http: //cl. sdsc. edu/ce. html • FSSP/Dali – http: //www. ebi. ac. uk/dali/Interactive. html
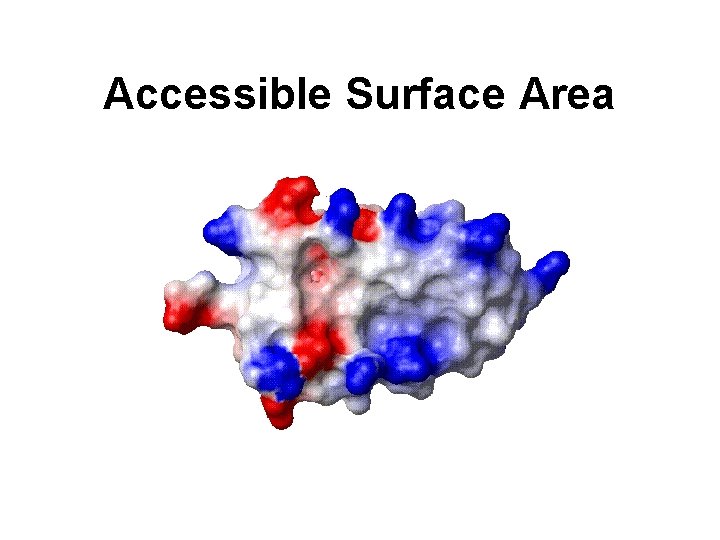
Accessible Surface Area
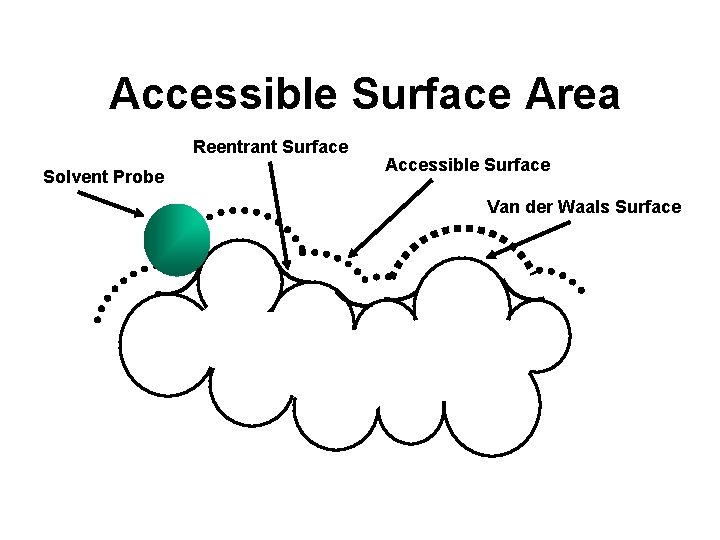
Accessible Surface Area Reentrant Surface Solvent Probe Accessible Surface Van der Waals Surface

ASA -- A Powerful Tool • Provides a picture of how water or other small molecules “see” the protein • Allows identification of exterior features from interior features • Allows identification of protrusions or crevices (i. e. active sites or binding sites)
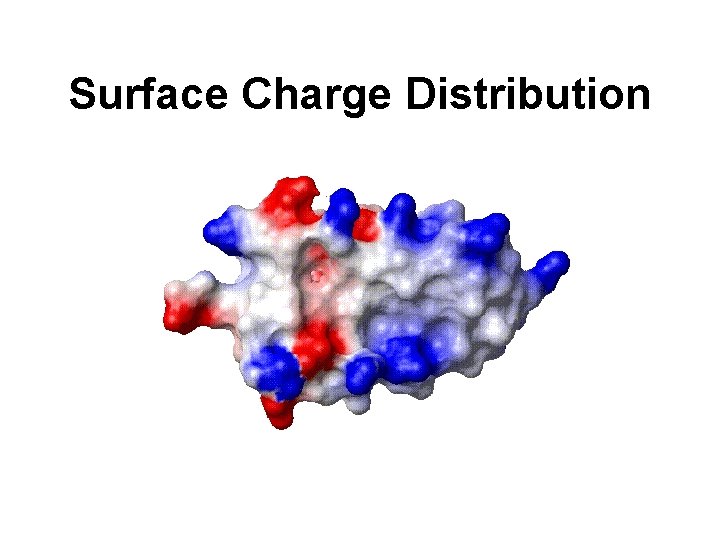
Surface Charge Distribution

Surface Charge • Allows positively and negatively charged structural features (protrusions, crevices) to be identified • Can be used to ID possible active sites or probably character of ligands • Key to many drug design efforts

Structure Features • • • Secondary Structure Supersecondary Structure Folding Class Polar/Nonpolar ASA Hydrogen Bond Parameters Stereochemistry Packing Defects Surface Charge Distribution Surface Roughness
http: //redpoll. pharmacy. ualberta. ca

Conclusion • Composition Features – Mass, p. I, Absorptivity, Rg, Volume • Sequence Features – Active sites, Binding Sites, Targeting, Location, Property Profiles, 2 o structure • Structure Features – Supersecondary Structure, Global Fold, ASA, Volume
- Slides: 102