Protease Enzyme Nomenclature Add ase to the name





































- Slides: 42

Protease

Enzyme Nomenclature • Add –ase to the name of the substrate – Urease Phosphatase • Class, subclass, subsubclass, serial no. EC 3. 4. 21. 5

The Six Classes of Enzymes 1. Oxidoreductases (dehydrogenases) Catalyze oxidation-reduction reactions 2. Transferases Catalyze group transfer reactions 3. Hydrolases 4. Catalyze hydrolysis reactions where water is the acceptor of the transferred group

4. Lyases Catalyze lysis of a substrate, generating a double bond in a nonhydrolytic, nonoxidative elimination (Synthases catalyze the addition to a double bond, the reverse reaction of a lyase) 5. Isomerases Catalyze isomerization reactions 6. Ligases (synthetases) - Catalyze ligation, or joining of two substrates - Require chemical energy (e. g. ATP)

Coenzyme • Many enzymes require nonprotein components to carry out their catalytic function- cofactors • Cofactors may be metal ions or organic molecules (coenzyme) • Cofactor: metal ion + coenzyme • Many coenzymes are vitamins or contain vitamins as part of their structure • Tightly bound coenzymes are referred as prosthetic groups

Holoenzyme and Apoenzyme • Holoenzyme – Complex of protein and prosthetic groups – Catalytically active • Apoenzyme – The enzyme without the prosthetic groups – Catalytically inactive

Oxidation-reduction reactions • Electrons are transferred between two species • Oxidizing agent gains electrons (is reduced) • Reducing agent donates electrons (is oxidized)

Acid-Base Catalysis • Reaction acceleration is achieved by catalytic transfer of a proton • A general base (B: ) can act as a proton acceptor to remove protons from OH, NH, CH or other XH

General base catalysis reactions • A general base (B: ) can remove a proton from water and thereby generate the equivalent of OH- in neutral solution • A general acid (BH+) can donate protons

Covalent Catalysis • All or part of a substrate is bound covalently to the enzyme to form a reactive intermediate • Group X can be transferred from A-X to B in two steps via the covalent ES complex X-E A-X + E X-E + A X-E + B B-X + E

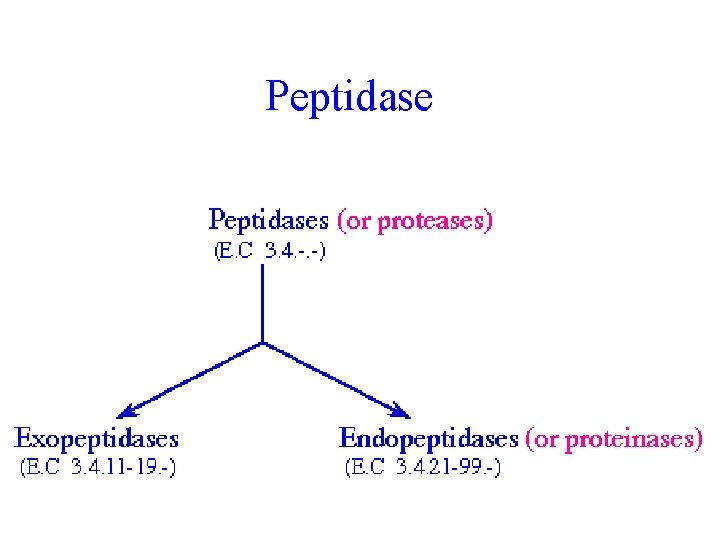
Peptidase

Protease (Peptide hydrolase, EC 3. 4. ) • Exopeptidase (EC 3. 4. 11 -19) – Act on N- or C-terminus of the peptide • Endopeptidase (EC 3. 4. 21 -24) – Classification based on the catalytic residues in the active site

Endopeptidase • • • Serine proteinases (EC 3. 4. 21) Cysteine proteinases (EC 3. 4. 22) Aspartic proteinases (EC 3. 4. 23) Metallo-proteinases (EC 3. 4. 24) EC 3. 4. 99 – A new, temporary subclass – Unknown catalytic mechanism

Serine proteinases (EC 3. 4. 21) • Contains serine at the active site • Catalytic triad • Covalent binding of substrate to Ser

Inhibitors of serine proteinases • Inhibited by diisopropyl fluorophosphate (DIF, DIFP) and diisopropyl phosphofluoridate (DIPF) • Most inhibited by phenylmethanesulfonyl fluoride (PMSF) • Some inhibited by chloromethyl ketone – TLCK (N-p-tosyl-L-lysine chloromethyl ketone) – TPCK (L-1 -tosylamido-2 -phenylethyl chloromethyl ketone)

Inhibition of serine protease with DFP • Diisopropyl fluorophosphate (DFP) is an organic phosphate that inactivates serine proteases • DFP reacts with the active site serine (Ser-195) of chymotrypsin to form DFP-chymotrypsin

Organophosphorous inhibitors • Such organophosphorous inhibitors are used as insecticides or for enzyme research • These inhibitors are toxic because they inhibit acetylcholinesterase (a serine protease that hydrolyzes the neurotransmitter acetylcholine)

Serine proteinases (EC 3. 4. 21) • Two superfamily – Chymotrypsin family • Trypsin • Chymotrypsin • Elastase • kallikrein and subtilism family • Subtilisin

Binding sites of chymotrypsin, and elastase • Substrate specificities are due to relatively small structural differences in active-site binding cavities

Serine Proteases • a-Chymotrypsin active site groups include: Ser-195, His-57 , Asp-102 Catalytic triad of chymotrypsin

Cysteine proteinases • Contains cysteine at the active site • Covalent binding of substrate to Cys • In cytosol or in lysosome

Inhibitors of cysteine proteinases • Inhibited by low concentration of p. HMB (phydroxymercuribenzoate), p. CMB (the hydrolysis product of pchloromercuribenzoate) • Inhibited by alkylating agents – Iodoacetate – Iodoacetamide – N-ethylmaleimide

Inhibitors of cysteine proteinases • Sulfhydryl reagents are not specific for Cys at the active site – Many proteinases contain –SH in their structures • Active site-specific cysteine reagent – E-64 [L-trans-epoxysuccinyl-leucylamido (4 guanidino) butane]

Cysteine proteinases • Papain- MW 23, 000, a single polypeptide • Lysosomal cysteine proteinase – Cathepsin B, H, L, S • Calpain – A cytosolic enzyme • Metal-dependent cysteine proteinase

Aspartic proteinases (Acid proteinases, EC 3. 4. 23) • Contains Asp at active site • General acid-base catalysis • No formation of covalent enzyme-substrate complex • Exists in secretory granules, membranes, endosomes, or lysosomes • Present in eukaryotes, not in prokaryotes

Inhibitors of aspartic proteinases • Inhibited by pepstatins • Inhibited by diazoacetyl compounds – Diazoacetyl-L-phe-methyl ester

Aspartic proteinases • Pepsin family – Digestive enzymes: pepsin and chymosin – Lysosomal cathepsins D – Processing enzymes: renin – Certain fungal proteases: penicillopepsin, rhizopuspepsin, endothiapepsin • Viral proteinases – Protease from the AIDS virus (HIV), also called retropepsin

Metallo-proteinases (EC 3. 4. 24) • Contains metal ions at the active center • The metal ions are an integral part of their structures, and enhance the nucleophilicity of H 2 O and polarize the peptide bond to be cleaved prior to nucleophiolic attack

Nucleophilic species are electron rich Electrophilic species are electron poor

Many Enzymes Require Inorganic Cations • Metal-activated enzymes • Metalloenzymes

Metal-activated enzymes • Have an absolute requirement or are stimulated by metal ions (examples: K+, Ca 2+, Mg 2+) • Binds metals loosely • Contains Cys or Ser at the active site

Metallo-enzymes • Contain firmly bound metal ions at the enzyme active sites (examples: iron, zinc, copper, cobalt ) • Widely distrubuted in prokaryotes and eukaryotes • Most of the exopeptidases are metalloproteins

Metallo-proteinase • Exists in ER, plasma membrane, mitochondria, cytosol • Inhibited by chealting agents- EDTA, EGTA • Metal chelator may inhibit metal-activated proteinases in addition to metalloproteinases

Regulation of proteinases activity • Compartmentation – Cathepsin in lysosome – Attachment of proteinases to membranes results in a loss of freedom • Synthesis and degradation • Inhibitors and activators • Regulation by metabolites – Ca 2+ , nucleotide • Covalent modification

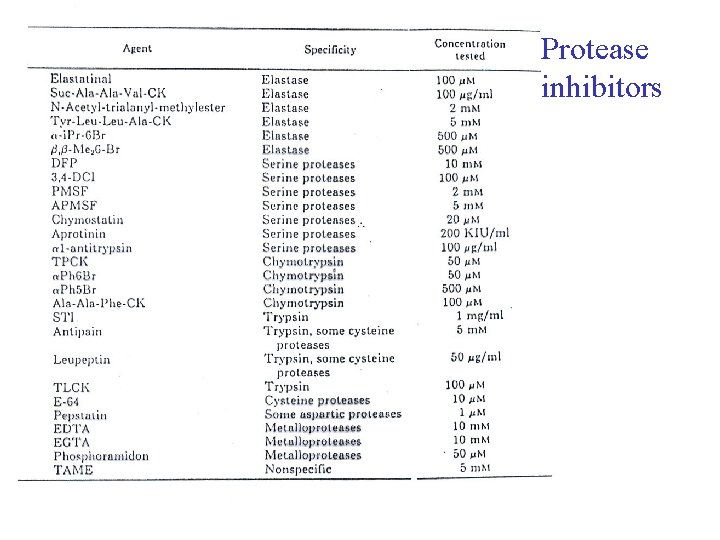
Protease inhibitors

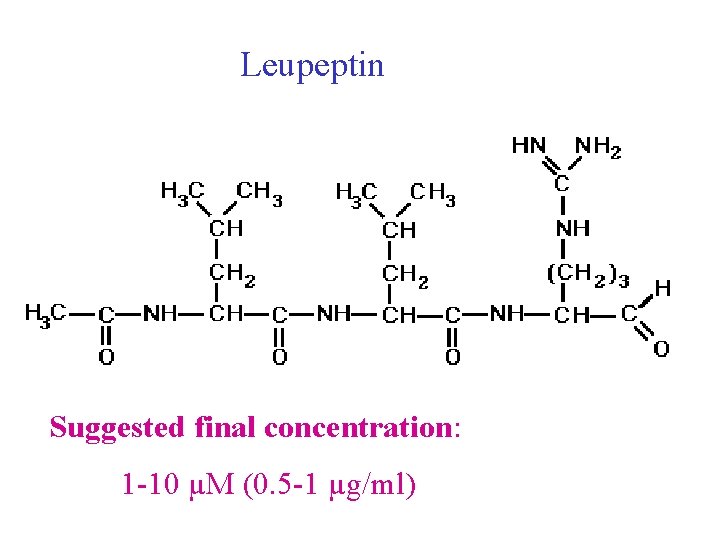
Leupeptin • C 20 H 38 N 6 O 4 (1/2 H 2 SO 4(H 2 O) • Formula Weight: 493. 62 • A protease inhibitor, will strongly inhibit trypsin, papain, plasmin, thrombokinase, kallikrein and cathepsin B. • The half-maximal inhibitory concentration ranges from 0. 5 to 75µg/ml, depending on the enzyme and the substrate. • Leupeptin does not inhibit chymotrypsin, elastase, renin, or pepsin. • Storage: -20°C

Leupeptin • Structure: Acetyl-leucyl-arginal • Inhibition spectrum: inhibits serine (trypsin (Ki=13 µM), plasmin, porcine kallikrein) and cysteine proteinases (papain, cathepsin B). Does not inhibit chymotrypsin and thrombin. • Mechanism of action: Competitive and reversible inhibitor. Inhibition may be relieved by an excess of substrate. • Properties: Soluble in water, ethanol, acetic acid and DMF (Stock solution: 10 m. M) MW: leupeptin: 426. 6; leupeptin hemisulphate monohydrate: 542. 7

Leupeptin Suggested final concentration: 1 -10 µM (0. 5 -1 µg/ml)

PMSF • A widely used serine protease • Half-life in water: 15 -60 minutes (depending on your reference source). • PMSF is not very soluble in water and should be kept at -20°C in dry methanol/propanol.

Proteases cocktail • PMSF – Stock 100 m. M (keep 4°C methanol), – Use at 1 m. M – Inhibit serine proteases • E 64 – Stock 1 m. M (keep -20°C, H 20) – Use at 10 m. M – Inhibit cysteine proteases (papain, calpain, cathepsin B, cathepsin L) • EDTA – Stock 500 m. M (keep 4°C) – Use at 5 m. M – Inhibit metallo-proteases

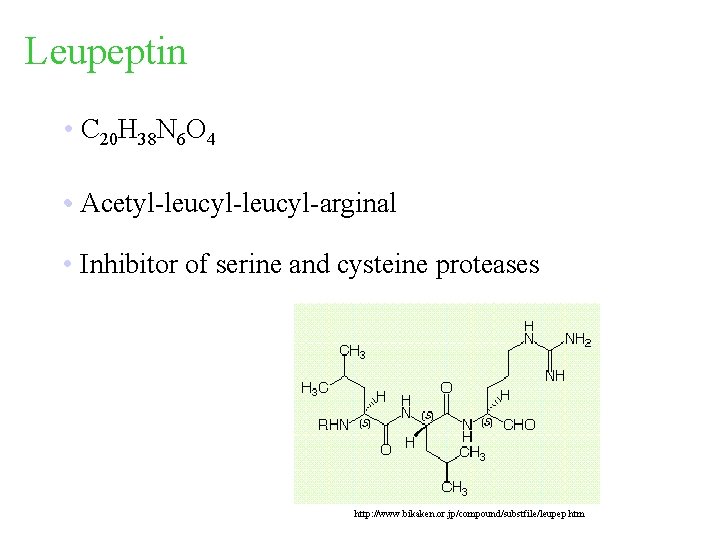
Leupeptin • C 20 H 38 N 6 O 4 • Acetyl-leucyl-arginal • Inhibitor of serine and cysteine proteases http: //www. bikaken. or. jp/compound/substfile/leupep. htm

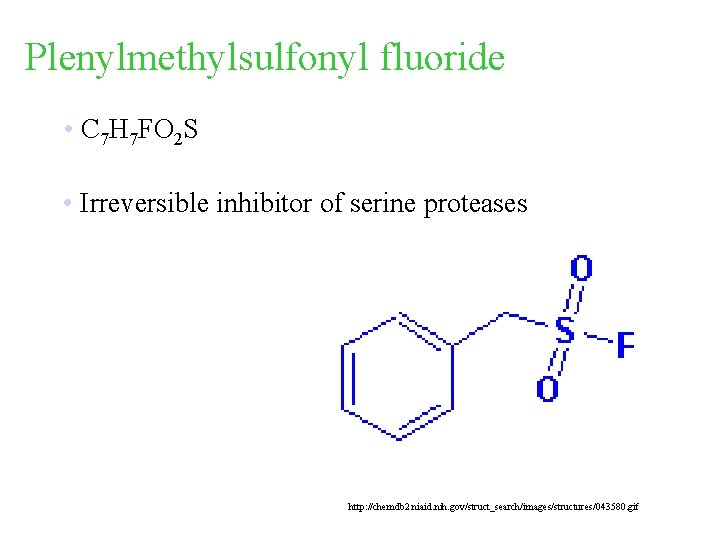
Plenylmethylsulfonyl fluoride • C 7 H 7 FO 2 S • Irreversible inhibitor of serine proteases http: //chemdb 2. niaid. nih. gov/struct_search/images/structures/043580. gif