Progress on Biomarkers of Cancer Diagnosis and Prognosis

Progress on Biomarkers of Cancer Diagnosis and Prognosis William CS CHO Queen Elizabeth Hospital, Hong Kong May 22, 2010



Dual-specificity phosphatase 6 (DUSP 6), monocyte-to-macrophage differentiation associated protein (MMD), signal transducer and activator of transcription 1 (STAT 1), v-erb-b 2 avian erythroblastic leukemia viral oncogene homolog 3 (ERBB 3), lymphocyte-specific protein tyrosine kinase (LCK). Conclusions Our five-gene signature is closely associated with relapse-free and overall survival among patients with NSCLC.

Kaplan–Meier Estimates of Survival of Patients with NSCLC According to the Five-Gene Signatures as Measured by RT-PCR. Overall survival and relapse-free survival are shown for the 101 patients with NSCLC (Panel A and Panel B, respectively) and for the 59 patients with stage I or II disease (Panel C and Panel D, respectively). Overall survival is also shown for the independent cohort of 60 patients (Panel E), for the 42 patients in this cohort who had stage I or II disease (Panel F), and for the 86 patients described in an independent set of published NSCLC microarray data 10 (Panel G).
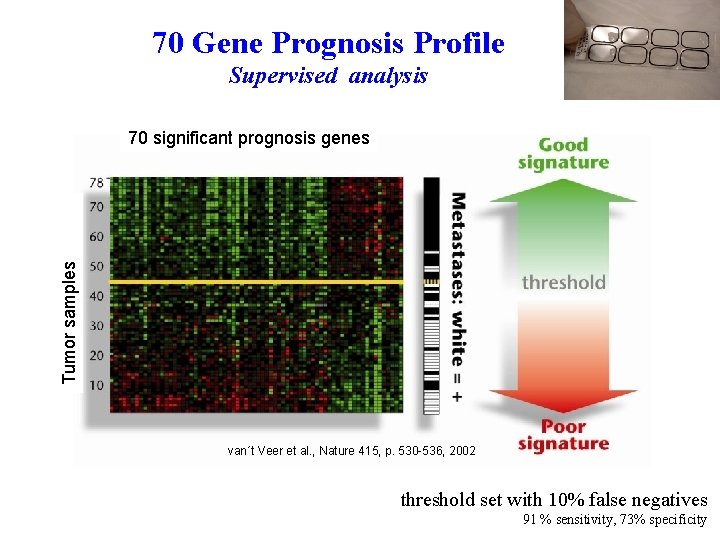
70 Gene Prognosis Profile Supervised analysis Tumor samples 70 significant prognosis genes van´t Veer et al. , Nature 415, p. 530 -536, 2002 threshold set with 10% false negatives 91 % sensitivity, 73% specificity
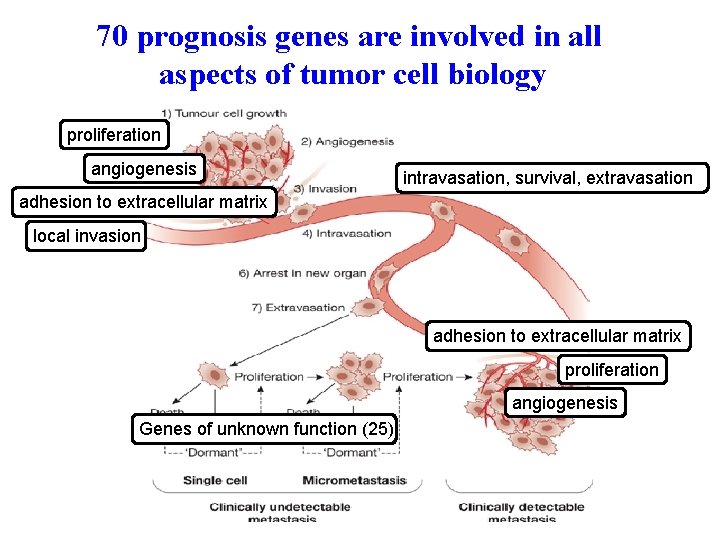
70 prognosis genes are involved in all aspects of tumor cell biology proliferation angiogenesis intravasation, survival, extravasation adhesion to extracellular matrix local invasion adhesion to extracellular matrix proliferation angiogenesis Genes of unknown function (25)

Independent validation: Buyse et al. (2006) JNCI. 98, 1183 -1192. 307 patients
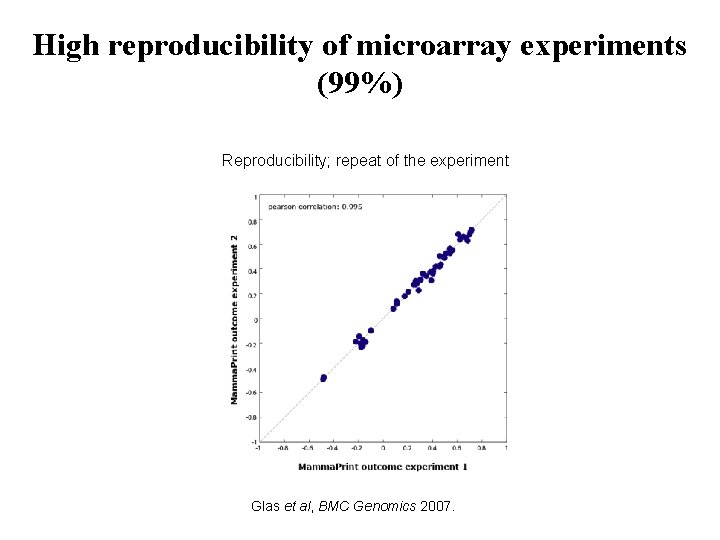
High reproducibility of microarray experiments (99%) Reproducibility; repeat of the experiment Glas et al, BMC Genomics 2007.
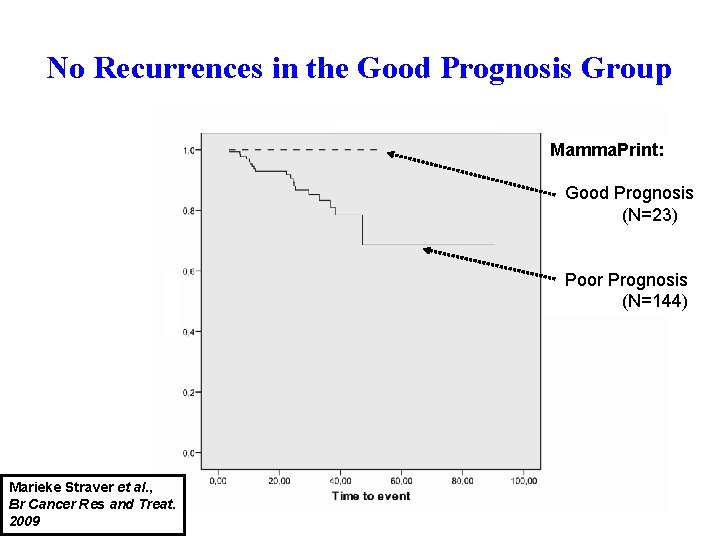
No Recurrences in the Good Prognosis Group Mamma. Print: Good Prognosis (N=23) Poor Prognosis (N=144) Marieke Straver et al. , Br Cancer Res and Treat. 2009
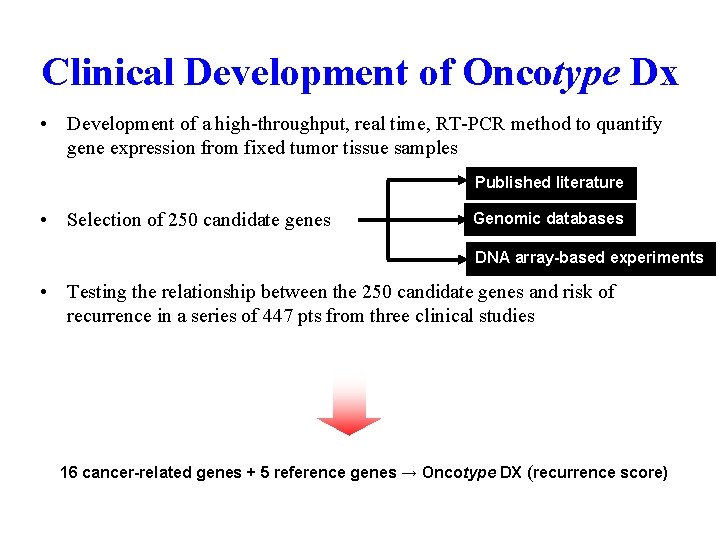
Clinical Development of Oncotype Dx • Development of a high-throughput, real time, RT-PCR method to quantify gene expression from fixed tumor tissue samples Published literature • Selection of 250 candidate genes Genomic databases DNA array-based experiments • Testing the relationship between the 250 candidate genes and risk of recurrence in a series of 447 pts from three clinical studies 16 cancer-related genes + 5 reference genes → Oncotype DX (recurrence score) Paik et al. NEJM. 2004.
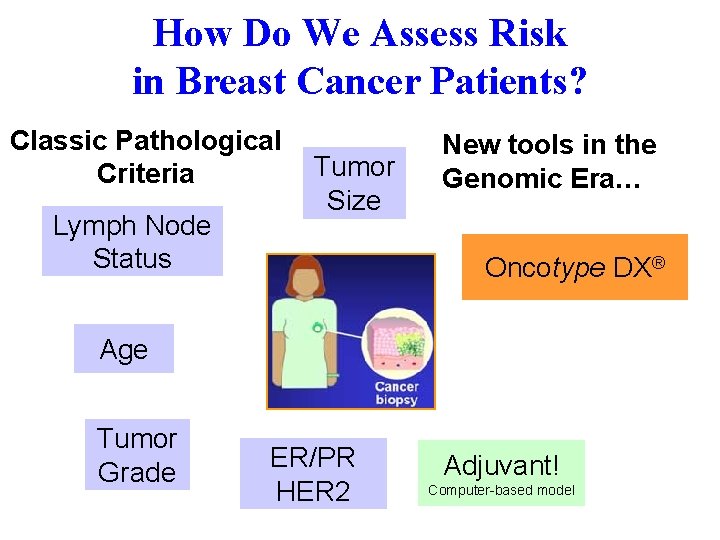
How Do We Assess Risk in Breast Cancer Patients? Classic Pathological Criteria Lymph Node Status Tumor Size New tools in the Genomic Era… Oncotype DX® Age Tumor Grade ER/PR HER 2 Adjuvant! Computer-based model

Oncotype DX 21 -gene recurrence score 16 cancer genes and 5 reference genes make up the Oncotype DX gene panel. The expression of these genes is used to calculate the recurrence score: PROLIFERATION Ki-67 STK 15 Survivin Cyclin B 1 MYBL 2 REFERENCE Beta-actin GAPDH RPLPO GUS TFRC ESTROGEN ER PR Bcl 2 SCUBE 2 RS = BAG 1 GSTM 1 INVASION Stromelysin 3 Cathepsin L 2 CD 68 HER 2 GRB 7 HER 2 + 0. 47 x HER 2 Group Score - 0. 34 x ER Group Score + 1. 04 x Proliferation Group Score + 0. 10 x Invasion Group Score + 0. 05 x CD 68 - 0. 08 x GSTM 1 Paik et al. N Engl J Med. - 0. 07 x BAG 1 2004; 351: 2817 -26.
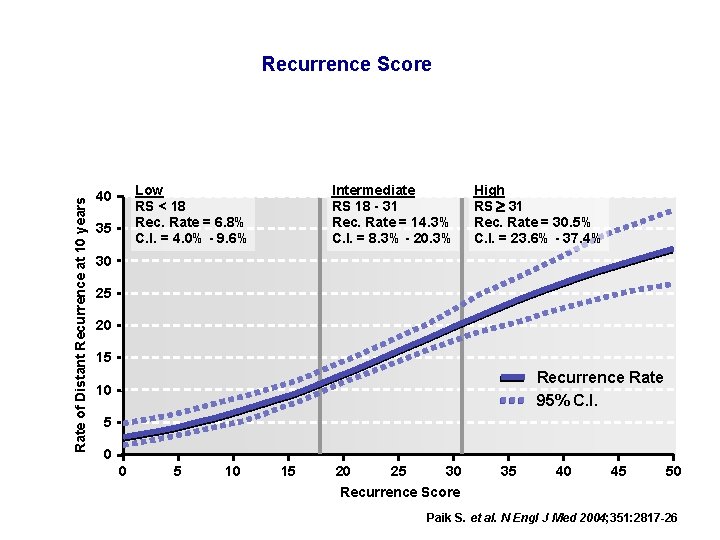
Rate of Distant Recurrence at 10 years Recurrence Score Low RS < 18 Rec. Rate = 6. 8% C. I. = 4. 0% - 9. 6% 40 35 Intermediate RS 18 - 31 Rec. Rate = 14. 3% C. I. = 8. 3% - 20. 3% High RS 31 Rec. Rate = 30. 5% C. I. = 23. 6% - 37. 4% 30 25 20 15 Recurrence Rate 95% C. I. 10 5 0 0 5 10 15 20 25 30 Recurrence Score 35 40 45 50 Paik S. et al. N Engl J Med 2004; 351: 2817 -26

Oncotype DXTM Low RS associated with minimal chemotherapy benefit; High RS associated with large chemotherapy benefit. The Oncotype DX Recurrence Score provides precise, quantitative information for individual patients on prognosis across and statistically independent of information on patient age, tumor size, and tumor grade.

Nobel Prize in Physiology or Medicine 2006 Andrew Z. Fire Craig C. Mello Cho WC. Micro. RNAs in cancer - from research to therapy. Biochim Biophys Acta - Rev Cancer 2010; 1805(2): 209 -217. C. elegans
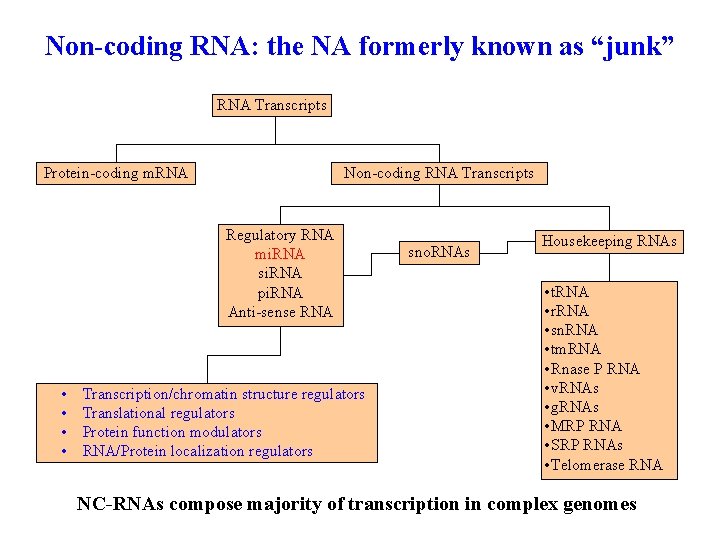
Non-coding RNA: the NA formerly known as “junk” RNA Transcripts Protein-coding m. RNA Non-coding RNA Transcripts Regulatory RNA mi. RNA si. RNA pi. RNA Anti-sense RNA • • Transcription/chromatin structure regulators Translational regulators Protein function modulators RNA/Protein localization regulators sno. RNAs Housekeeping RNAs • t. RNA • r. RNA • sn. RNA • tm. RNA • Rnase P RNA • v. RNAs • g. RNAs • MRP RNA • SRP RNAs • Telomerase RNA NC-RNAs compose majority of transcription in complex genomes
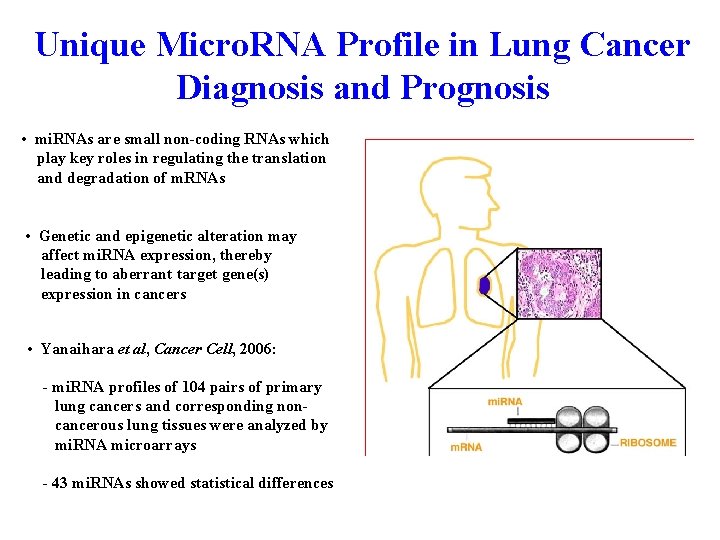
Unique Micro. RNA Profile in Lung Cancer Diagnosis and Prognosis • mi. RNAs are small non-coding RNAs which play key roles in regulating the translation and degradation of m. RNAs • Genetic and epigenetic alteration may affect mi. RNA expression, thereby leading to aberrant target gene(s) expression in cancers • Yanaihara et al, Cancer Cell, 2006: - mi. RNA profiles of 104 pairs of primary lung cancers and corresponding noncancerous lung tissues were analyzed by mi. RNA microarrays - 43 mi. RNAs showed statistical differences
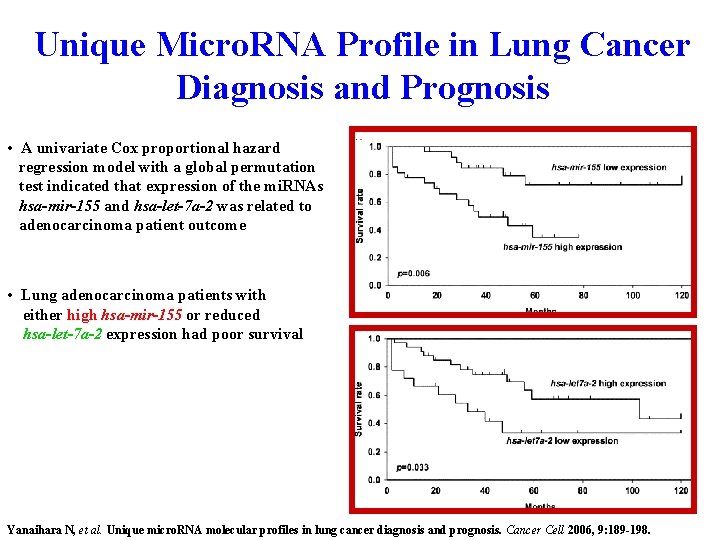
Unique Micro. RNA Profile in Lung Cancer Diagnosis and Prognosis • A univariate Cox proportional hazard regression model with a global permutation test indicated that expression of the mi. RNAs hsa-mir-155 and hsa-let-7 a-2 was related to adenocarcinoma patient outcome • Lung adenocarcinoma patients with either high hsa-mir-155 or reduced hsa-let-7 a-2 expression had poor survival Yanaihara N, et al. Unique micro. RNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 2006, 9: 189 -198.


The role of micro. RNAs in cancer diagnosis • With the application of in situ RT-PCR, it was shown that the aberrantly expressed mi. R-221, mi. R-301 and mi. R-376 a were localized to pancreatic cancer cells but not to stroma or normal acini or ducts. • Aberrant mi. RNA expression offered new clues to pancreatic tumorigenesis and might provide diagnostic biomarkers for pancreatic cancer. Lee EJ, et al. Expression profiling identifies micro. RNA signature in pancreatic cancer. Int J Cancer 2007, 120: 1046 -1054. Cho WC. Micro. RNAs: potential biomarkers for cancer diagnosis, prognosis and targets for therapy. Int J Biochem Cell Biol 2010. Cho WC. Micro. RNAs in cancer - from research to therapy. Biochim Biophys Acta - Rev Cancer 2010; 1805(2): 209 -217.

The role of micro. RNAs in cancer prognosis • Expression of let-7 mi. RNA was frequently reduced in human lung cancers, and that reduced let-7 mi. RNA expression was significantly associated with shorter postoperative survival. • Overexpression of let-7 mi. RNA in A 549 lung adenocarcinoma cell line inhibited lung cancer cell growth in vitro. Takamizawa J, et al. Reduced expression of the let-7 micro. RNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 2004, 64: 3753 -3756.

The role of micro. RNAs in cancer prognosis • The expression pattern of mi. RNAs in pancreatic cancer were compared with those of normal pancreas and chronic pancreatitis using mi. RNA microarrays. • Differentially expressed mi. RNAs were identified which could differentiate pancreatic cancer from normal pancreas, chronic pancreatitis, or both. • High expression of mi. R-196 a-2 was found to predict poor survival of more than 24 months. Bloomston M, et al. Micro. RNA expression patterns to differentiate pancreatic adenocarcinoma from normal pancreas and chronic pancreatitis. JAMA 2007, 297: 1901 -1908.
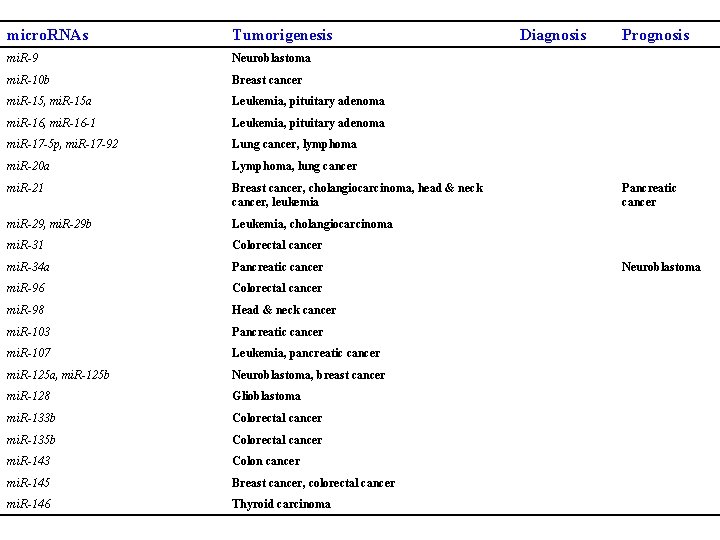
micro. RNAs Tumorigenesis mi. R-9 Neuroblastoma mi. R-10 b Breast cancer mi. R-15, mi. R-15 a Leukemia, pituitary adenoma mi. R-16, mi. R-16 -1 Leukemia, pituitary adenoma mi. R-17 -5 p, mi. R-17 -92 Lung cancer, lymphoma mi. R-20 a Lymphoma, lung cancer mi. R-21 Breast cancer, cholangiocarcinoma, head & neck cancer, leukemia mi. R-29, mi. R-29 b Leukemia, cholangiocarcinoma mi. R-31 Colorectal cancer mi. R-34 a Pancreatic cancer mi. R-96 Colorectal cancer mi. R-98 Head & neck cancer mi. R-103 Pancreatic cancer mi. R-107 Leukemia, pancreatic cancer mi. R-125 a, mi. R-125 b Neuroblastoma, breast cancer mi. R-128 Glioblastoma mi. R-133 b Colorectal cancer mi. R-135 b Colorectal cancer mi. R-143 Colon cancer mi. R-145 Breast cancer, colorectal cancer mi. R-146 Thyroid carcinoma Diagnosis Prognosis Pancreatic cancer Neuroblastoma
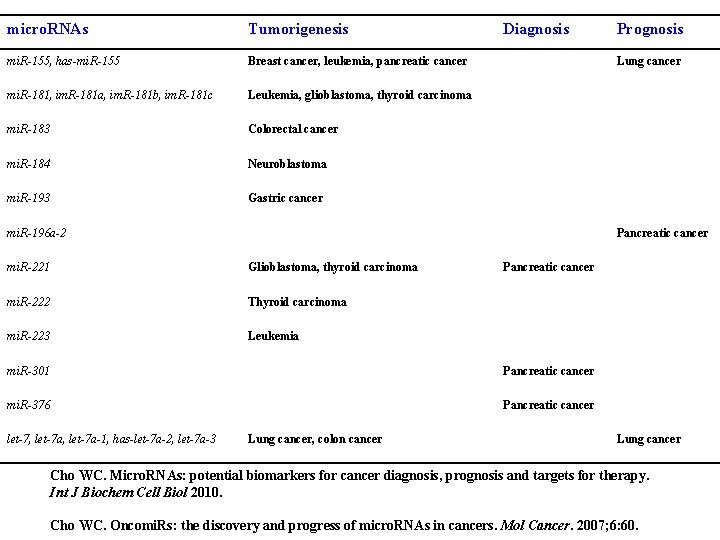
micro. RNAs Tumorigenesis mi. R-155, has-mi. R-155 Breast cancer, leukemia, pancreatic cancer mi. R-181, im. R-181 a, im. R-181 b, im. R-181 c Leukemia, glioblastoma, thyroid carcinoma mi. R-183 Colorectal cancer mi. R-184 Neuroblastoma mi. R-193 Gastric cancer Diagnosis Lung cancer mi. R-196 a-2 Pancreatic cancer mi. R-221 Glioblastoma, thyroid carcinoma mi. R-222 Thyroid carcinoma mi. R-223 Leukemia Pancreatic cancer mi. R-301 Pancreatic cancer mi. R-376 Pancreatic cancer let-7, let-7 a-1, has-let-7 a-2, let-7 a-3 Prognosis Lung cancer, colon cancer Lung cancer Cho WC. Micro. RNAs: potential biomarkers for cancer diagnosis, prognosis and targets for therapy. Int J Biochem Cell Biol 2010. Cho WC. Oncomi. Rs: the discovery and progress of micro. RNAs in cancers. Mol Cancer. 2007; 6: 60.

Beyond the genome Same genome Different proteome
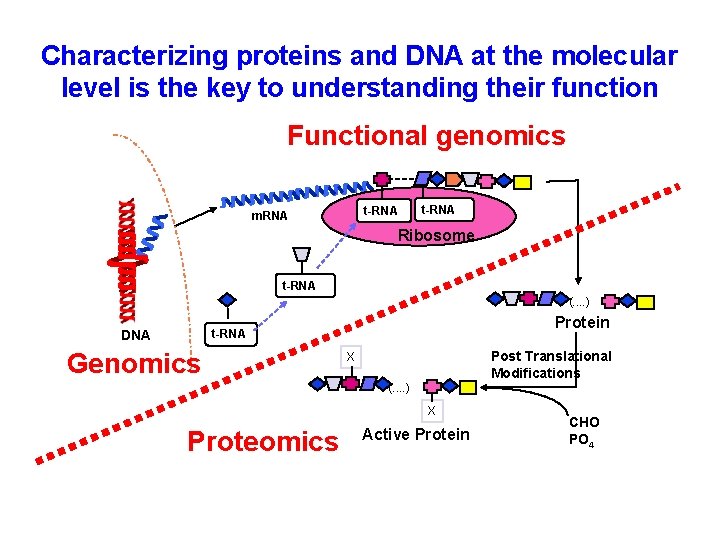
Characterizing proteins and DNA at the molecular level is the key to understanding their function Functional genomics t-RNA m. RNA t-RNA Ribosome t-RNA (. . ) DNA Protein t-RNA Genomics Post Translational Modifications X (. . ) X Proteomics Active Protein CHO PO 4

Proteomics: leading biological science in the 21 st century • Proteomics represents the effort to establish the identities, quantities, structures, biochemical and cellular functions of all proteins in an organism, organ, or organelle • and how these properties vary in space, time, or physiological state. Cho WC. Proteomics – Leading biological science in the 21 st century. Science J, 2004; 56(5): 14 -17. Cho WC, Cheng CH. Oncoproteomics: current trends and future perspectives. Expert Rev Proteomics 2007; 4(3): 401 -410.

Traditional vs High-throughput approach
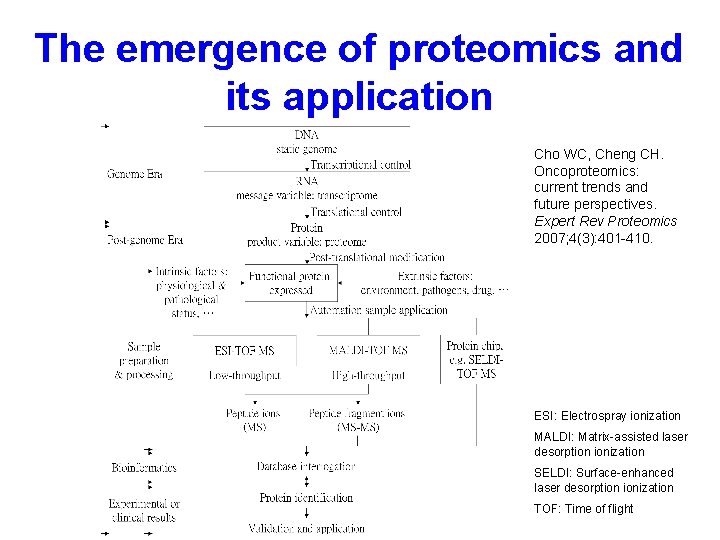
The emergence of proteomics and its application Cho WC, Cheng CH. Oncoproteomics: current trends and future perspectives. Expert Rev Proteomics 2007; 4(3): 401 -410. ESI: Electrospray ionization MALDI: Matrix-assisted laser desorption ionization SELDI: Surface-enhanced laser desorption ionization TOF: Time of flight
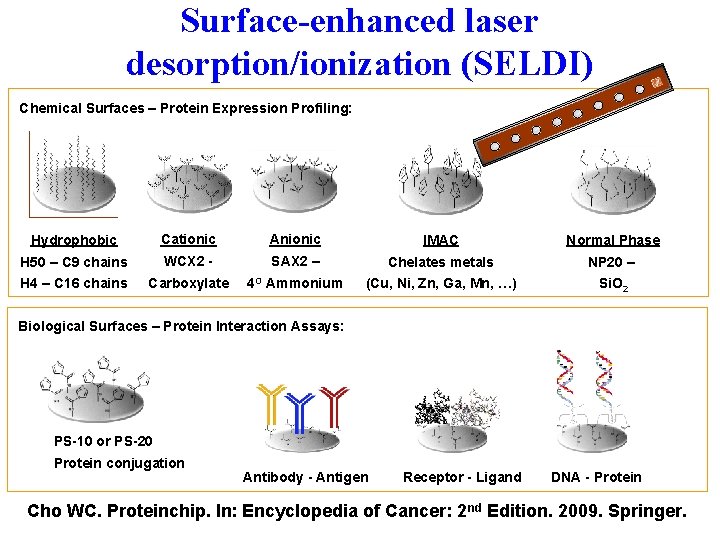
Surface-enhanced laser desorption/ionization (SELDI) Chemical Surfaces – Protein Expression Profiling: Hydrophobic Cationic Anionic IMAC Normal Phase H 50 – C 9 chains WCX 2 - SAX 2 – Chelates metals NP 20 – H 4 – C 16 chains Carboxylate 4 O Ammonium (Cu, Ni, Zn, Ga, Mn, …) Si. O 2 Biological Surfaces – Protein Interaction Assays: PS-10 or PS-20 Protein conjugation Antibody - Antigen Receptor - Ligand DNA - Protein Cho WC. Proteinchip. In: Encyclopedia of Cancer: 2 nd Edition. 2009. Springer.

HTP automation Biomek 2000 (Beckman) Programmed protocols for highly reproducible sample processing Proteinchip System PCS 4000 Aquarius (Tecan)

Sample fractionation, chip binding and data acquisition in SELDI-TOF MS Cho WC, et al. Clin Cancer Res 2004; 10: 43 -52. Cho WC. Chin J Biotech 2006; 22(6): 871 -876. Cho WC, et al. J Cell Biochem 2006; 99(1): 256 -68. Cho WC, et al. Dis Markers 2006; 22(3): 153 -66. Cho WC, et al. J Ethnopharmacol 2006; 108(2): 272 -9. Cho WC, et al. Clin Chem 2007; 53(2): 241 -250.
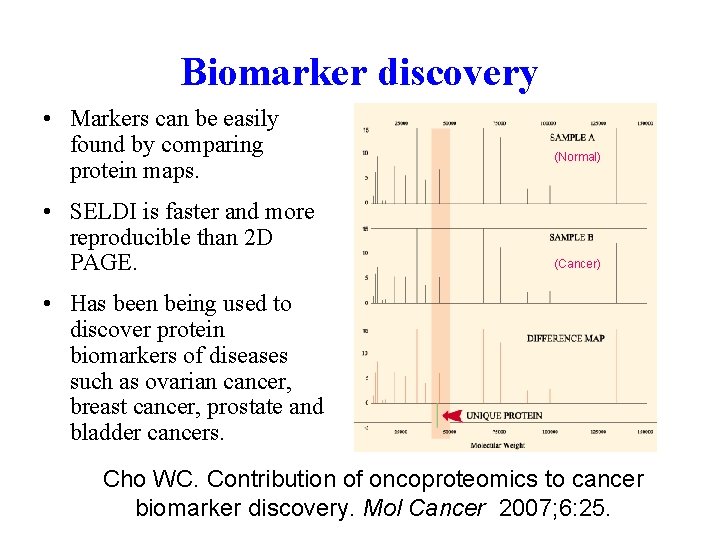
Biomarker discovery • Markers can be easily found by comparing protein maps. • SELDI is faster and more reproducible than 2 D PAGE. (Normal) (Cancer) • Has been being used to discover protein biomarkers of diseases such as ovarian cancer, breast cancer, prostate and bladder cancers. Cho WC. Contribution of oncoproteomics to cancer biomarker discovery. Mol Cancer 2007; 6: 25.

Proteins as biomarkers The protein composition may be associated with disease processes in the organism and thus have potential utility as diagnostic markers. • Proteins are closer to the actual disease process, in most cases, than parent genes • Proteins are ultimate regulators of cellular function • Most cancer markers are proteins • The vast majority of drug targets are proteins Cho WC. Cancer biomarkers (an overview). In Hayat MA (ed): Methods of cancer diagnosis, therapy and prognosis. Volume 7. New York, NY: Springer, 5 Jan 2010.
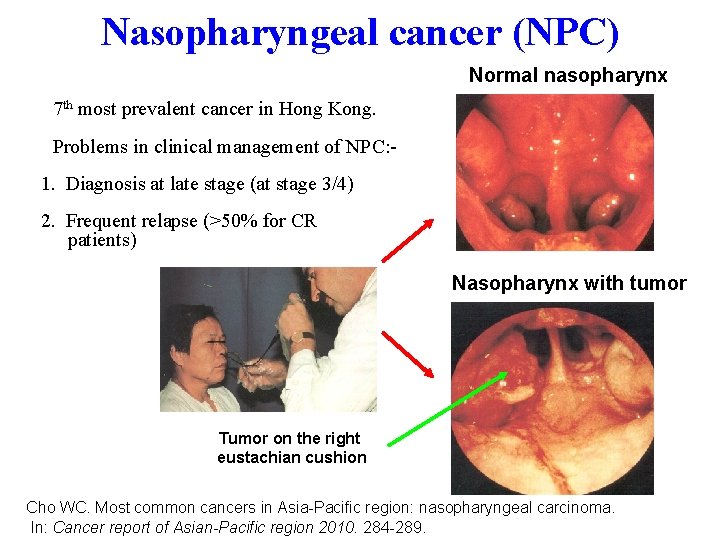
Nasopharyngeal cancer (NPC) Normal nasopharynx • 7 th most prevalent cancer in Hong Kong. • Problems in clinical management of NPC: 1. Diagnosis at late stage (at stage 3/4) 2. Frequent relapse (>50% for CR patients) Nasopharynx with tumor Tumor on the right eustachian cushion Cho WC. Most common cancers in Asia-Pacific region: nasopharyngeal carcinoma. In: Cancer report of Asian-Pacific region 2010. 284 -289.

Proteinchip application: nasopharyngeal carcinoma biomarkers discovery • Serum samples from 149 NPC patients (undifferentiated carcinoma of the nasopharyngeal type or poorly differentiated squamous cell type) • 35 normal individuals


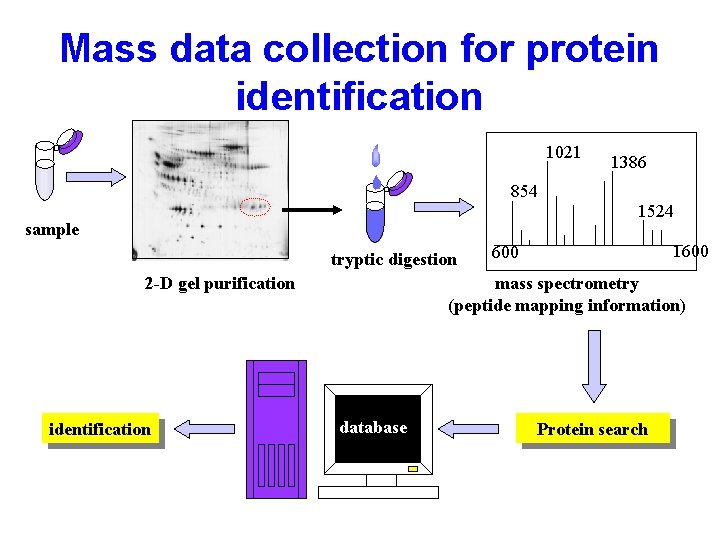
Mass data collection for protein identification 1021 854 sample tryptic digestion 2 -D gel purification identification 1386 1524 1600 mass spectrometry (peptide mapping information) database Protein search

Identification of marker by MS/MS 34/37 ions matched with Serum Amyloid A
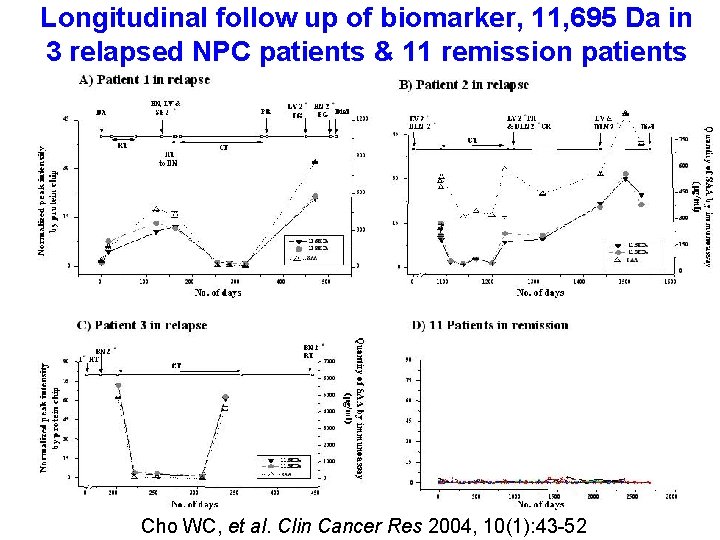
Longitudinal follow up of biomarker, 11, 695 Da in 3 relapsed NPC patients & 11 remission patients Cho WC, et al. Clin Cancer Res 2004, 10(1): 43 -52

Serum biomarkers with changes before and after chemotherapy in relapsed NPC patients EP: Biomarker: 7, 659 Da EP, Etoposide and Cisplatinum; GC: Biomarker: 7, 765 Da GC, Gemcitabine and Cisplatinum. Cho WC et al. Protein. Chip array profiling for identification of disease- and chemotherapy-associated biomarkers of nasopharyngeal carcinoma. Clin Chem. 2007; 53(2): 241 -50.

Basic statistics of ovarian cancer • Prevalence 40/100, 000 (1 in 2500) • 23, 000 new cases diagnosed annually • 14, 000 deaths annually • Overall 5 year survival 20 -30% • 75% of cases are diagnosed in late stage (stage III/IV) • 90% cure rate in stage I/IIa • Therefore, detection in earlier stages critical in improving overall survival
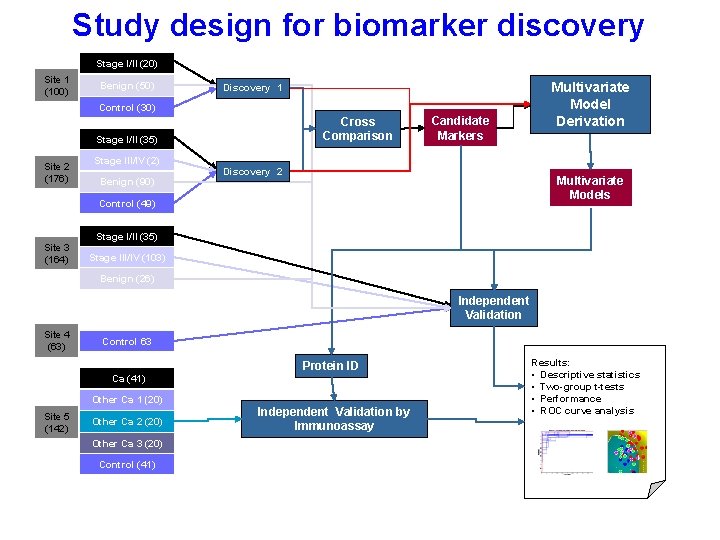
Study design for biomarker discovery Stage I/II (20) Site 1 (100) Benign (50) Discovery 1 Control (30) Cross Comparison Stage I/II (35) Site 2 (176) Stage III/IV (2) Benign (90) Candidate Markers Discovery 2 Multivariate Models Control (49) Site 3 (164) Multivariate Model Derivation Stage I/II (35) Stage III/IV (103) Benign (26) Independent Validation Site 4 (63) Control 63 Protein ID Ca (41) Other Ca 1 (20) Site 5 (142) Other Ca 2 (20) Other Ca 3 (20) Control (41) Independent Validation by Immunoassay Results: • Descriptive statistics • Two-group t-tests • Performance • ROC curve analysis

Summary of performance • Markers for Stage I/II ovarian cancer discovered using Protein. Chip system • 503 samples from 5 institutions • Rigorous cross-validation and independent validation study design • Fixed specificity (97%) • 3 marker panel (Apolipoprotein A 1, inter alpha trypsin inhibitor IV and Transthyretin) : 74% sensitivity • CA 125: 65% sensitivity • Fixed sensitivity (83%) • 3 marker panel: 94% specificity • CA 125: 54% specificity
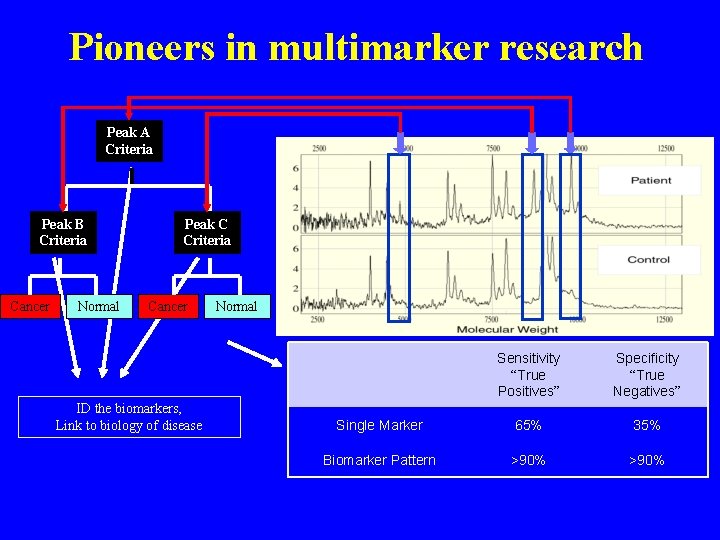
Pioneers in multimarker research Peak A Criteria Peak B Criteria Cancer Normal Peak C Criteria Cancer ID the biomarkers, Link to biology of disease Normal Sensitivity “True Positives” Specificity “True Negatives” Single Marker 65% 35% Biomarker Pattern >90%

FDA Cleared the OVA 1 Test on Sep 11, 2009 ● Translating biomarker discovery from lab to clinic ● Based on a prospective double-blind clinical trial involved 516 patients from 27 institutions • 269 patients were evaluated by pre-surgical information alone • 247 patients were evaluated by pre-surgical information with OVA 1 results ● OVA 1 identified additional patients with potential malignancies ● Help to guide surgical decisions

OVA 1 ● First FDA-cleared protein-based in vitro diagnostic multivariate index assay ● First FDA-cleared prognostic test for ovarian cancer in the pre- and post-surgical setting ● Test 5 proteins in blood sample • β 2 -microglobulin, transferrin, transthyretin identified by SELDI apolipoprotein • CA 125 ● Indicate the likelihood of benign or malignant A 1,
Scientific American Cho WC. Proteomic approaches to cancer target identification. Drug Discov Today: Ther Strategies 2007; 4(4): 245 -250.
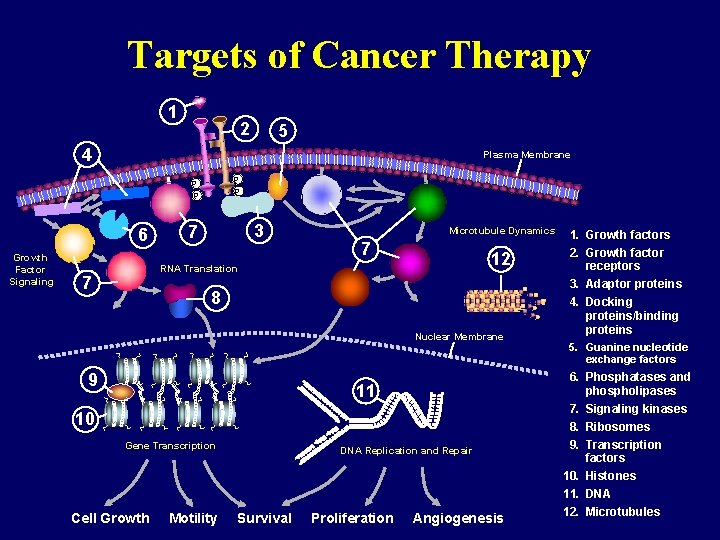
Targets of Cancer Therapy 1 2 5 4 Plasma Membrane P P Growth Factor Signaling PDK 1, 2 6 3 7 Microtubule Dynamics 7 12 RNA Translation 7 8 Nuclear Membrane 9 11 10 Gene Transcription Cell Growth Motility DNA Replication and Repair Survival Proliferation Angiogenesis 1. Growth factors 2. Growth factor receptors 3. Adaptor proteins 4. Docking proteins/binding proteins 5. Guanine nucleotide exchange factors 6. Phosphatases and phospholipases 7. Signaling kinases 8. Ribosomes 9. Transcription factors 10. Histones 11. DNA 12. Microtubules

In colon cancer KRAS mutation determines response to EGFR therapy Mutant KRAS +EGFR -EGFR Wild type KRAS +EGFR -EGFR 51 Amado et al. J Clin Oncol; 26: 1626 -1634 2008
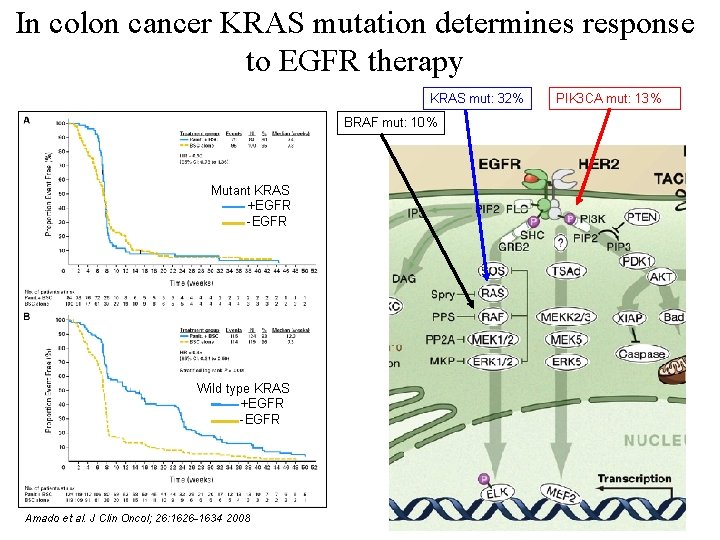
In colon cancer KRAS mutation determines response to EGFR therapy KRAS mut: 32% PIK 3 CA mut: 13% BRAF mut: 10% Mutant KRAS +EGFR -EGFR Wild type KRAS +EGFR -EGFR 52 Amado et al. J Clin Oncol; 26: 1626 -1634 2008

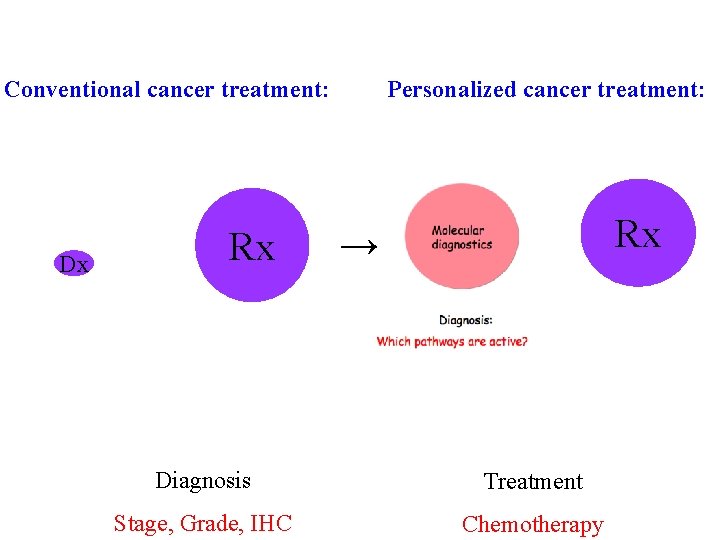
Conventional cancer treatment: Dx Rx Personalized cancer treatment: Rx → Diagnosis Treatment Stage, Grade, IHC Chemotherapy
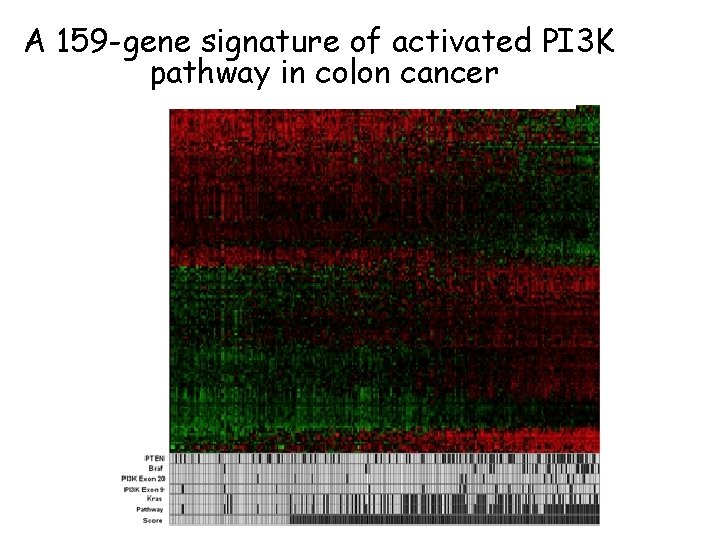
A 159 -gene signature of activated PI 3 K pathway in colon cancer
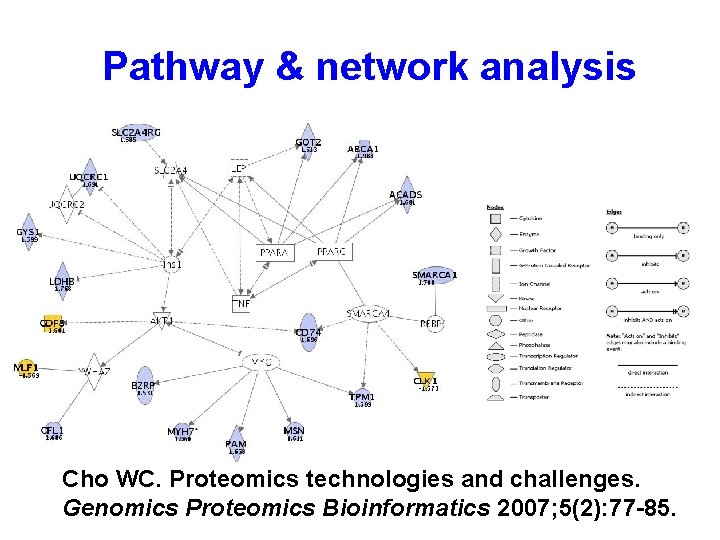
Pathway & network analysis Cho WC. Proteomics technologies and challenges. Genomics Proteomics Bioinformatics 2007; 5(2): 77 -85.
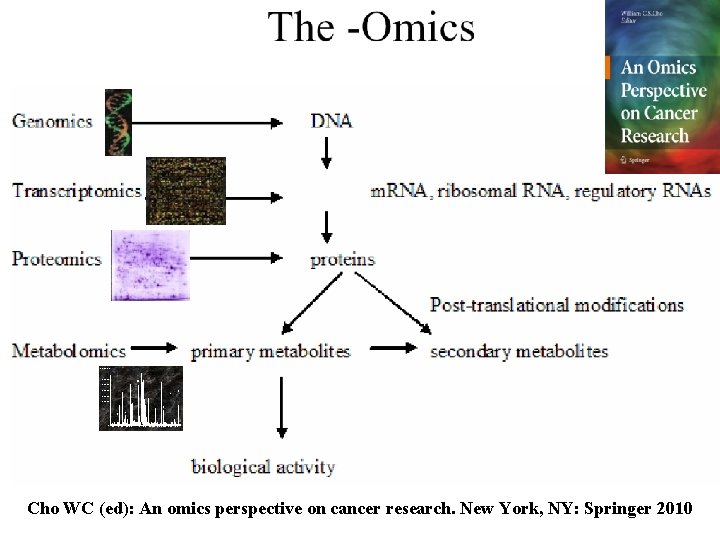
Cho WC (ed): An omics perspective on cancer research. New York, NY: Springer 2010

Thank You E-mail: chocs@ha. org. hk
- Slides: 59