Program Normalization exercise from last week Dimension reduction













- Slides: 31

Program • Normalization exercise (from last week) • Dimension reduction theory (PCA/Clustering) • Dimension reduction exercise DNA Microarray Bioinformatics - #27611

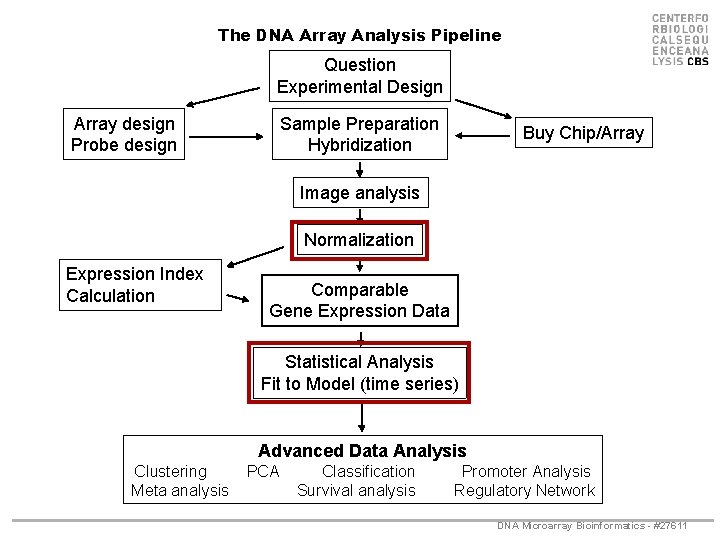
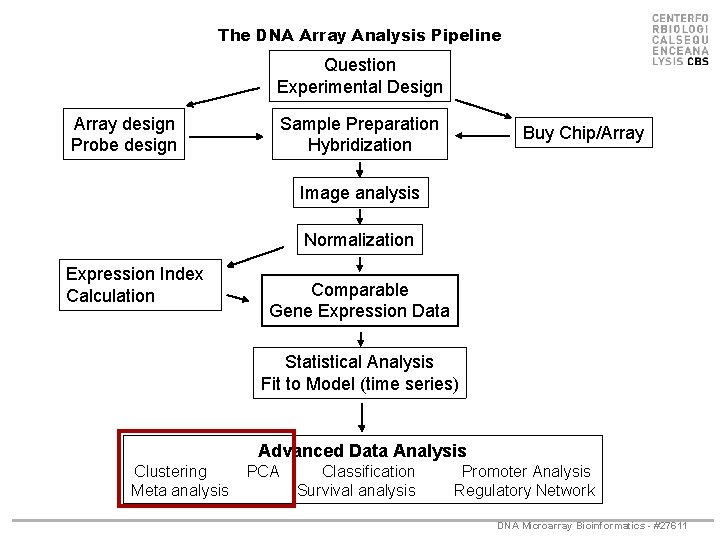
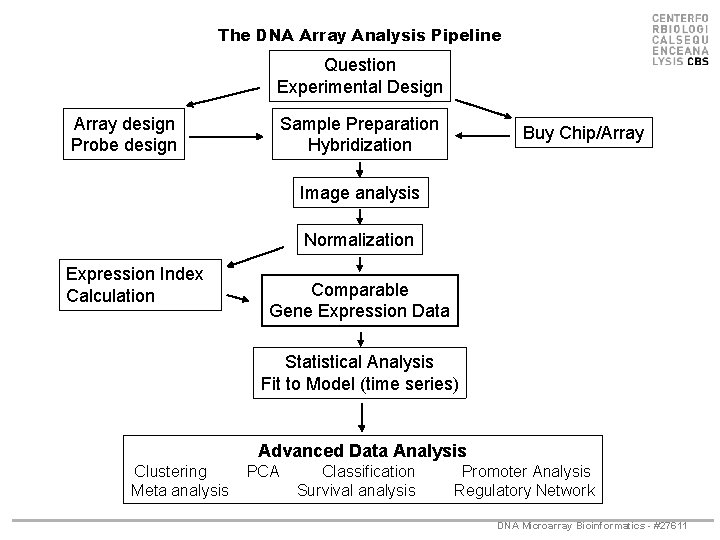
The DNA Array Analysis Pipeline Question Experimental Design Array design Probe design Sample Preparation Hybridization Buy Chip/Array Image analysis Normalization Expression Index Calculation Comparable Gene Expression Data Statistical Analysis Fit to Model (time series) Advanced Data Analysis Clustering Meta analysis PCA Classification Survival analysis Promoter Analysis Regulatory Network DNA Microarray Bioinformatics - #27611

The DNA Array Analysis Pipeline Question Experimental Design Array design Probe design Sample Preparation Hybridization Buy Chip/Array Image analysis Normalization Expression Index Calculation Comparable Gene Expression Data Statistical Analysis Fit to Model (time series) Advanced Data Analysis Clustering Meta analysis PCA Classification Survival analysis Promoter Analysis Regulatory Network DNA Microarray Bioinformatics - #27611

Dimension reduction methods • • • Principal component analysis (PCA) Cluster analysis Multidimensional scaling Correspondance analysis Singular value decomposition Slides stolen more or less from Agnieszka Juncker. DNA Microarray Bioinformatics - #27611

Dimension reduction methods • • • Principal component analysis (PCA) Cluster analysis Multidimensional scaling Correspondance analysis Singular value decomposition DNA Microarray Bioinformatics - #27611

Principal Component Analysis (PCA) • used for visualization of complex data • developed to capture as much of the variation in data as possible DNA Microarray Bioinformatics - #27611

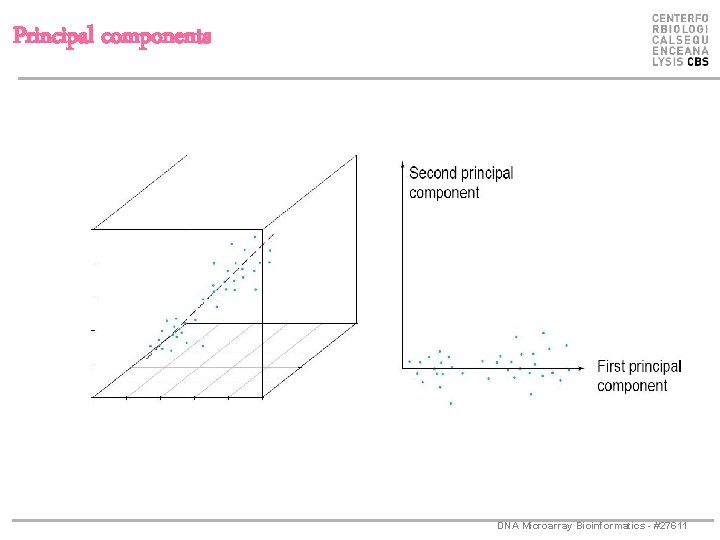
Principal components 1. principal component (PC 1) – the direction along which there is greatest variation 2. principal component (PC 2) – the direction with maximum variation left in data, orthogonal to the 1. PC DNA Microarray Bioinformatics - #27611

Principal components DNA Microarray Bioinformatics - #27611

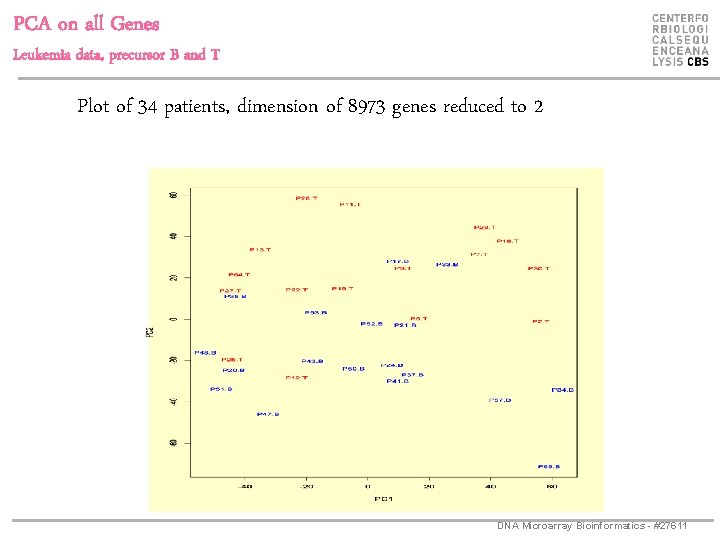
PCA on all Genes Leukemia data, precursor B and T Plot of 34 patients, dimension of 8973 genes reduced to 2 DNA Microarray Bioinformatics - #27611

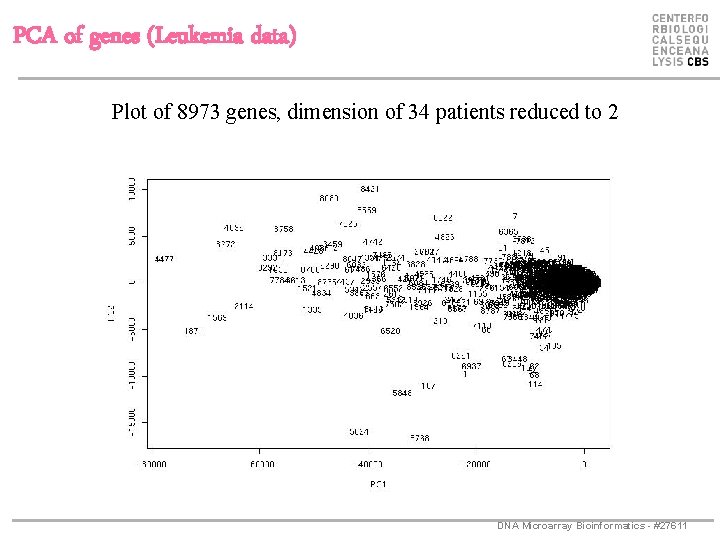
PCA of genes (Leukemia data) Plot of 8973 genes, dimension of 34 patients reduced to 2 DNA Microarray Bioinformatics - #27611

Principal components General about principal components – – summary variables linear combinations of the original variables uncorrelated with each other capture as much of the original variance as possible DNA Microarray Bioinformatics - #27611

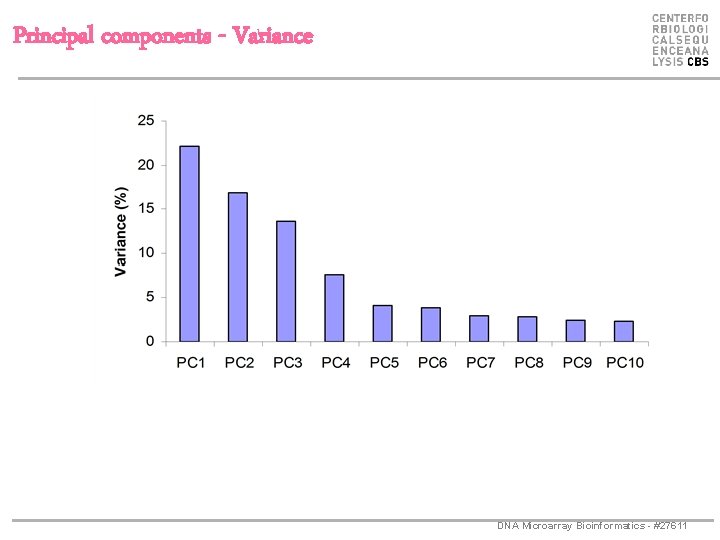
Principal components - Variance DNA Microarray Bioinformatics - #27611

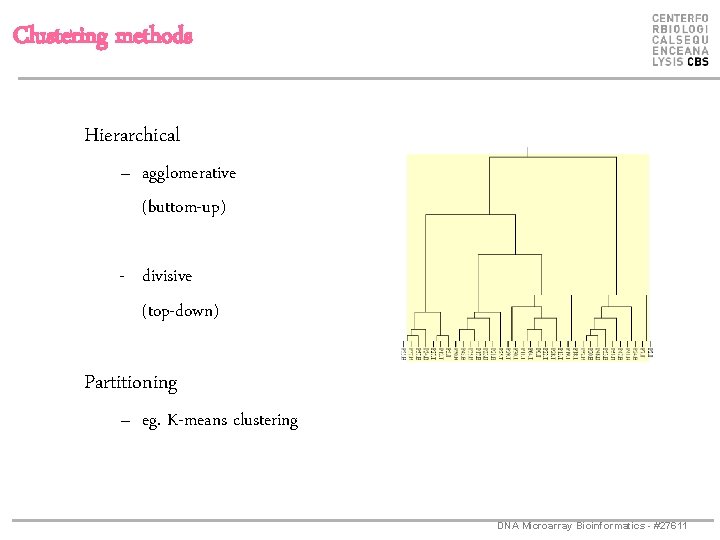
Clustering methods Hierarchical – agglomerative (buttom-up) - divisive (top-down) Partitioning – eg. K-means clustering DNA Microarray Bioinformatics - #27611

Hierarchical clustering Representation of all pairwise distances Parameters: none (distance measure) Results: – in one large cluster – hierarchical tree (dendrogram) Deterministic DNA Microarray Bioinformatics - #27611

Hierarchical clustering – UPGMA Algorithm Assign each item to its own cluster Join the nearest clusters Reestimate the distance between clusters Repeat for 1 to n DNA Microarray Bioinformatics - #27611

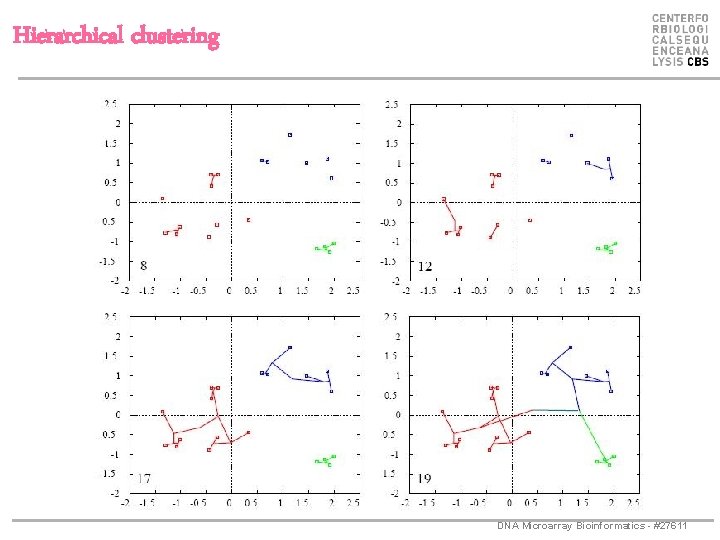
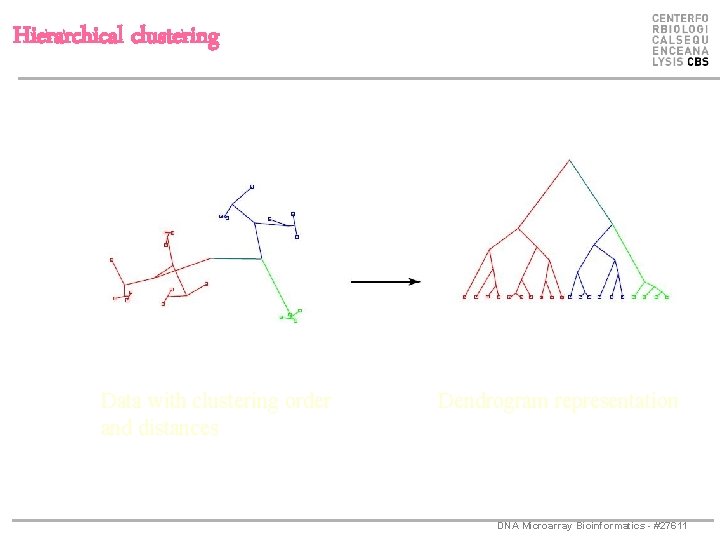
Hierarchical clustering DNA Microarray Bioinformatics - #27611

Hierarchical clustering Data with clustering order and distances Dendrogram representation DNA Microarray Bioinformatics - #27611

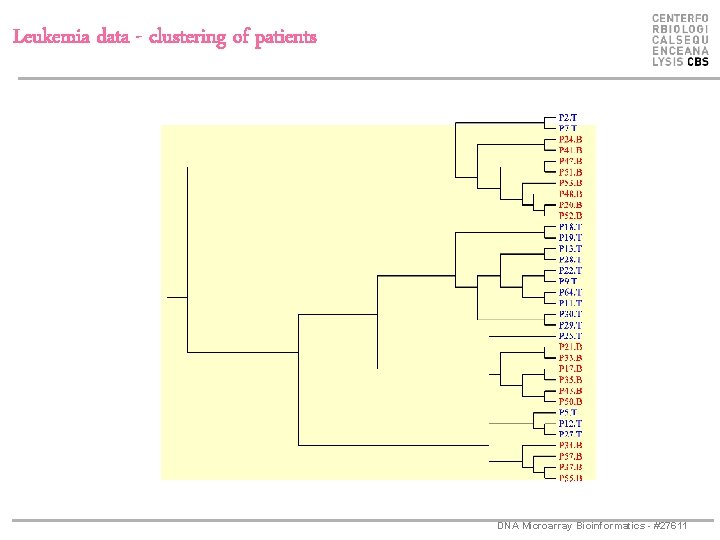
Leukemia data - clustering of patients DNA Microarray Bioinformatics - #27611

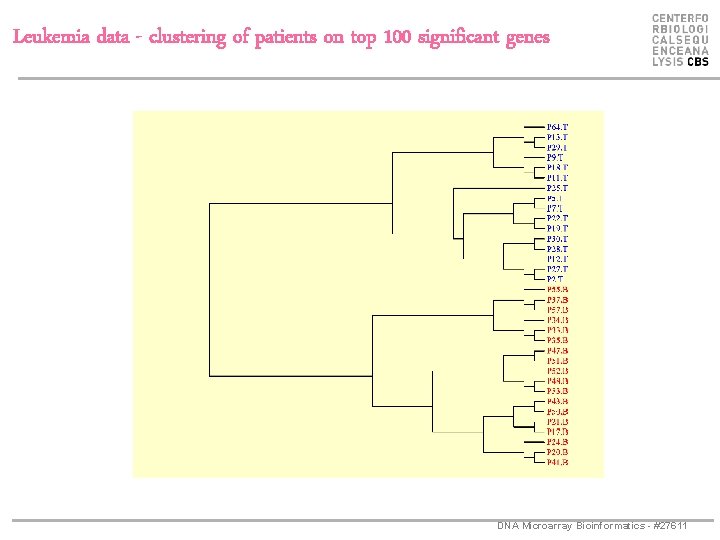
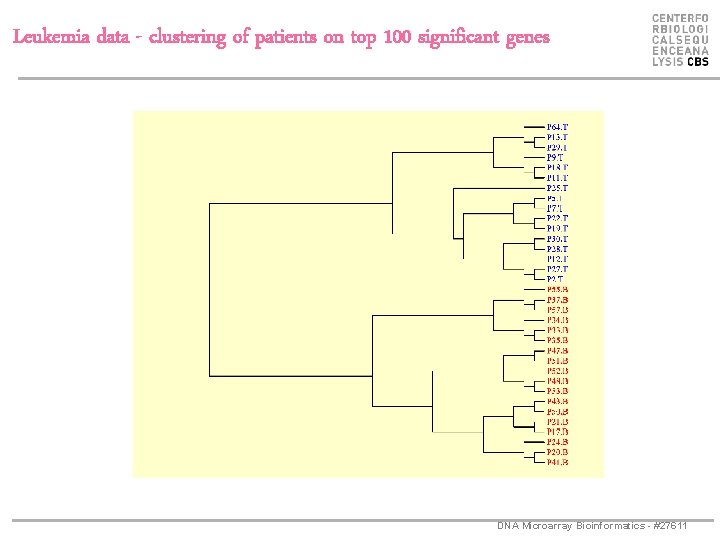
Leukemia data - clustering of patients on top 100 significant genes DNA Microarray Bioinformatics - #27611

The DNA Array Analysis Pipeline Question Experimental Design Array design Probe design Sample Preparation Hybridization Buy Chip/Array Image analysis Normalization Expression Index Calculation Comparable Gene Expression Data Statistical Analysis Fit to Model (time series) Advanced Data Analysis Clustering Meta analysis PCA Classification Survival analysis Promoter Analysis Regulatory Network DNA Microarray Bioinformatics - #27611

Leukemia data - clustering of patients on top 100 significant genes DNA Microarray Bioinformatics - #27611

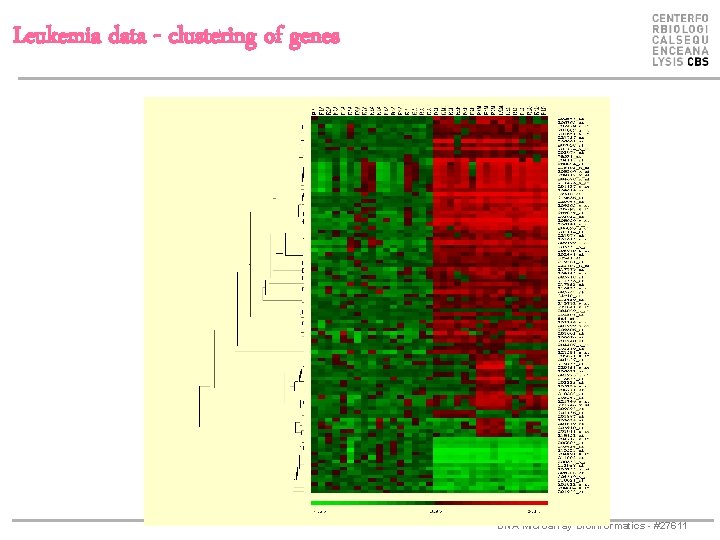
Leukemia data - clustering of genes DNA Microarray Bioinformatics - #27611

DNA Microarray Bioinformatics - #27611

K-means clustering Partition data into K clusters Parameter: Number of clusters (K) must be chosen Randomilized initialization: – different clusters each time DNA Microarray Bioinformatics - #27611

K-means - Algorithm Assign each item a class in 1 to K (randomly) For each class 1 to K – Calculate the centroid (one of the K-means) – Calculate distance from centroid to each item Assign each item to the nearest centroid Repeat until no items are re-assigned (convergence) DNA Microarray Bioinformatics - #27611

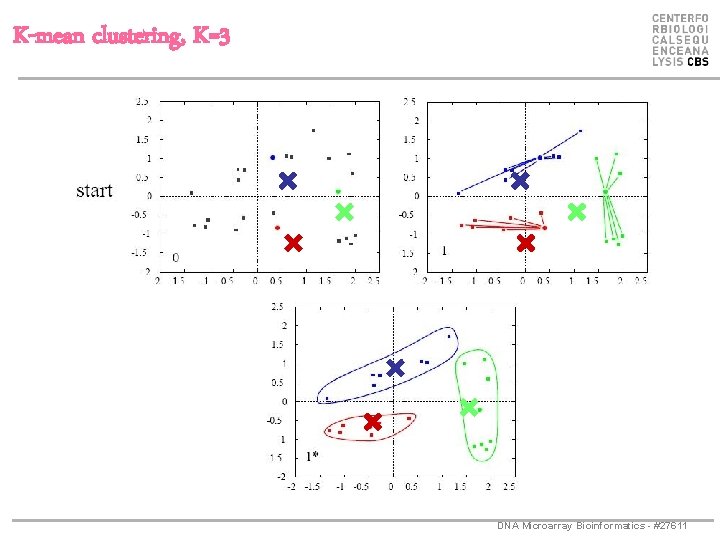
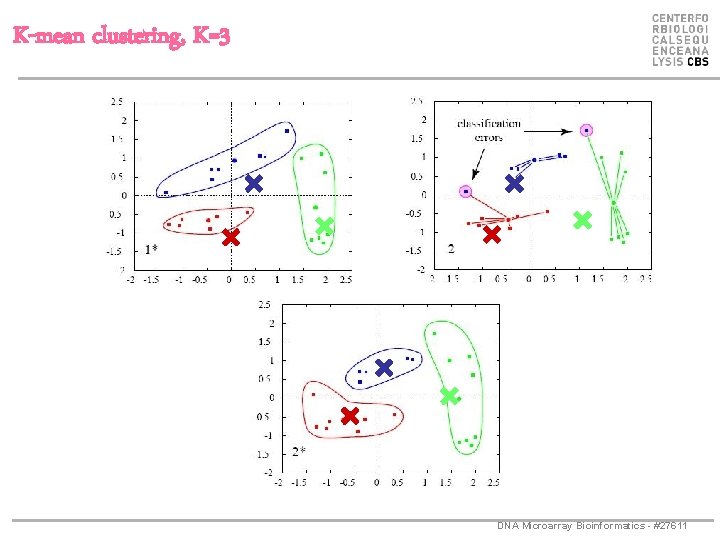
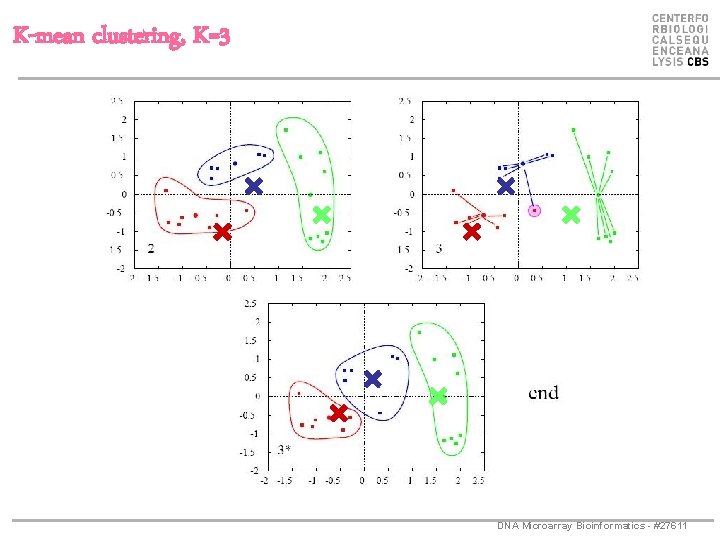
K-mean clustering, K=3 DNA Microarray Bioinformatics - #27611

K-mean clustering, K=3 DNA Microarray Bioinformatics - #27611

K-mean clustering, K=3 DNA Microarray Bioinformatics - #27611

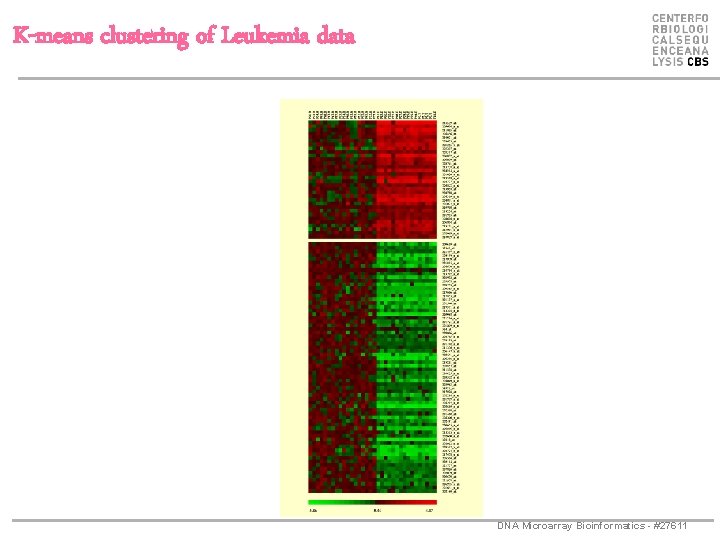
K-means clustering of Leukemia data DNA Microarray Bioinformatics - #27611

Steen Knudsen: A Biologist’s guide to Analysis of microarray data. • Chapter 4: Visualization by Reduction of Dimensionality (PCA) • Chapter 5: Cluster Analysis DNA Microarray Bioinformatics - #27611

Bioinformatics – Real science or fortunetelling? 1. Changing paradigm 1. From final answer to qualified guessing 2. From student to “real” scientist 3. YOU are evolving with this course 2. Always cheating 1. NOT real biology - Approximation to the truth 2. No final answer (in our life time) DNA Microarray Bioinformatics - #27611