Prof em Klaus Ammann University of Bern Switzerland

Prof. em. Klaus Ammann, University of Bern, Switzerland Gene Editing: Regulation with Scalable Flexibility A solution for the EU, the Cartagena Protocol and elsewhere Brussels, Copa-Cogeca, Rue de Trèves, Room A

Maize Field Trial on Gene Flow: GIS supported results show: 10 -20 m safety distances are enough Maize field I, individual sampling plants Ammann, K. , Kovacovsky, K. , & Bratschi, D. (2009) Maize Geneflow - GPS-Mapping and 3 -dimensional Landscape Modeling, final report WP 2 SIGMEA project, Wider Fields University of Bern Participation pp 25 (Manuscript) http: //www. botanischergarten. ch/SIGMEA-Bern-Final-Report-20090831. pdf Wider Fields Models of Maize Pla

Outcrossing of blue kernel maize surrounding yellow kernel maize, Slightly different growing conditions due to sowing machine turning Change in flowering synchronization: higher outcrossing, SIGMEA Ammann, K. , Kovacovsky, K. , & Bratschi, D. (2009) Maize Geneflow - GPS-Mapping and 3 -dimensional Landscape Modeling, final report WP 2 SIGMEA project, Wider Fields University of Bern Participation pp 25 (Manuscript) http: //www. botanischergarten. ch/SIGMEA-Bern-Final-Report-20090831. pdf Maize Field I and II from gene flow experiment in Switzerland: spots caused through sowing machine turned around, caused change in flowering synchroni-zation: higher outcrossing, but all results well within the legal threshold of the E

The basic monograph, a draft literature review Ammann Klaus. (20160514). Modern Plant Breeding and Future Biosafety Regulation, 750 references with full text links for private use. ASK-FORCE Manuscript, pp. 325. http: //www. ask-force. org/web/Genomic. Misconception/Ammann-Modern-Plant. Breeding-and-Future-Regulation 20160514. pdf
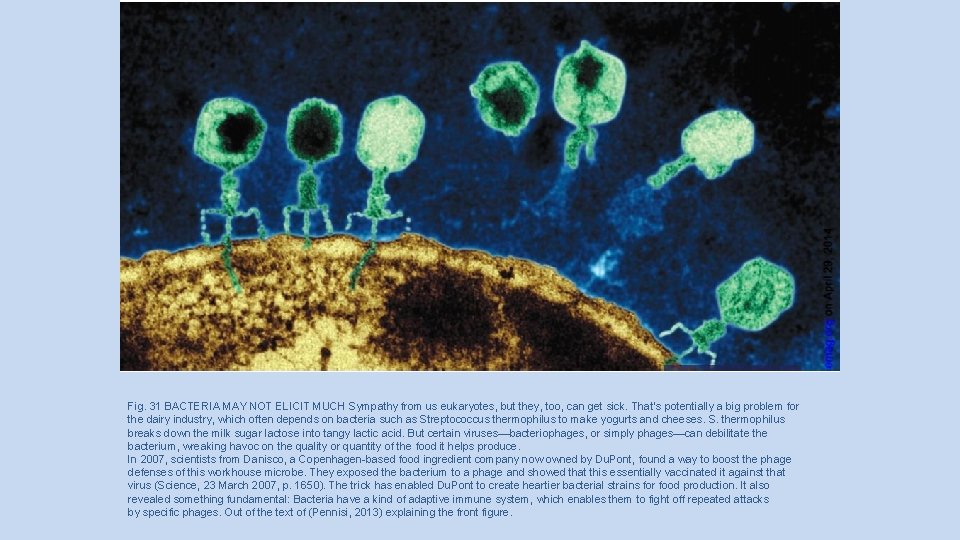
Fig. 31 BACTERIA MAY NOT ELICIT MUCH Sympathy from us eukaryotes, but they, too, can get sick. That’s potentially a big problem for the dairy industry, which often depends on bacteria such as Streptococcus thermophilus to make yogurts and cheeses. S. thermophilus breaks down the milk sugar lactose into tangy lactic acid. But certain viruses—bacteriophages, or simply phages—can debilitate the bacterium, wreaking havoc on the quality or quantity of the food it helps produce. In 2007, scientists from Danisco, a Copenhagen-based food ingredient company now owned by Du. Pont, found a way to boost the phage defenses of this workhouse microbe. They exposed the bacterium to a phage and showed that this essentially vaccinated it against that virus (Science, 23 March 2007, p. 1650). The trick has enabled Du. Pont to create heartier bacterial strains for food production. It also revealed something fundamental: Bacteria have a kind of adaptive immune system, which enables them to fight off repeated attacks by specific phages. Out of the text of (Pennisi, 2013) explaining the front figure.
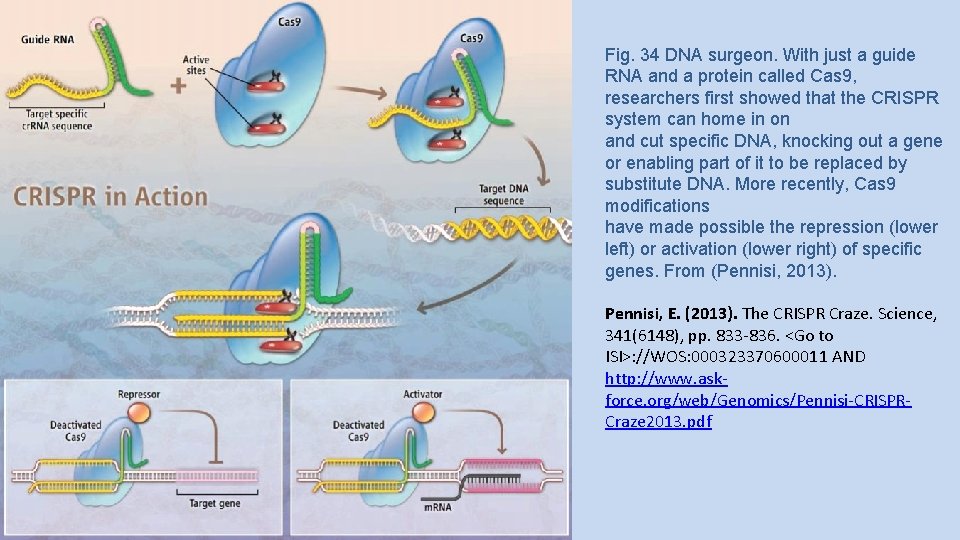
Fig. 34 DNA surgeon. With just a guide RNA and a protein called Cas 9, researchers first showed that the CRISPR system can home in on and cut specific DNA, knocking out a gene or enabling part of it to be replaced by substitute DNA. More recently, Cas 9 modifications have made possible the repression (lower left) or activation (lower right) of specific genes. From (Pennisi, 2013). Pennisi, E. (2013). The CRISPR Craze. Science, 341(6148), pp. 833 -836. <Go to ISI>: //WOS: 000323370600011 AND http: //www. askforce. org/web/Genomics/Pennisi-CRISPRCraze 2013. pdf

Rapid Trait Development System (RTDS) in Plants, from: http: //www. cibus. com/technology. php
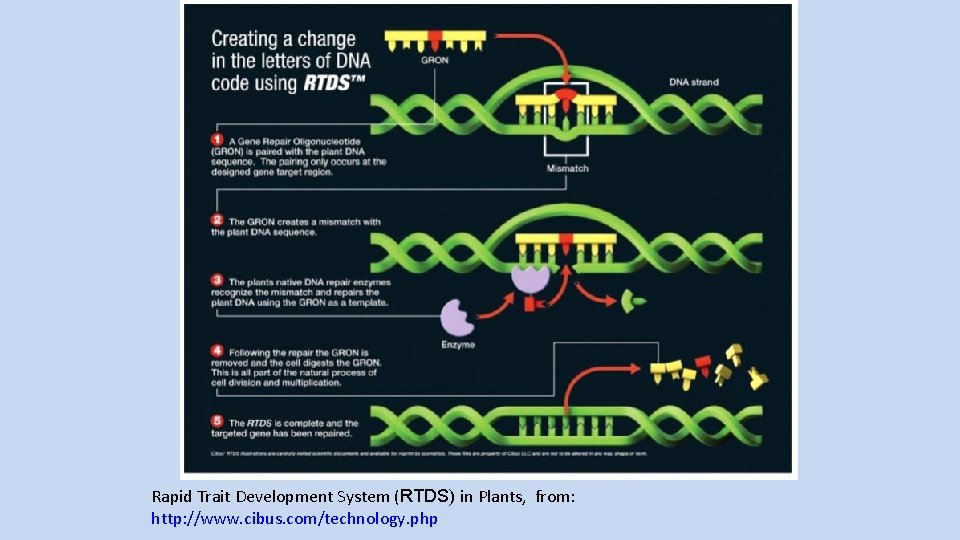
Rapid Trait Development System (RTDS) in Plants, from: http: //www. cibus. com/technology. php
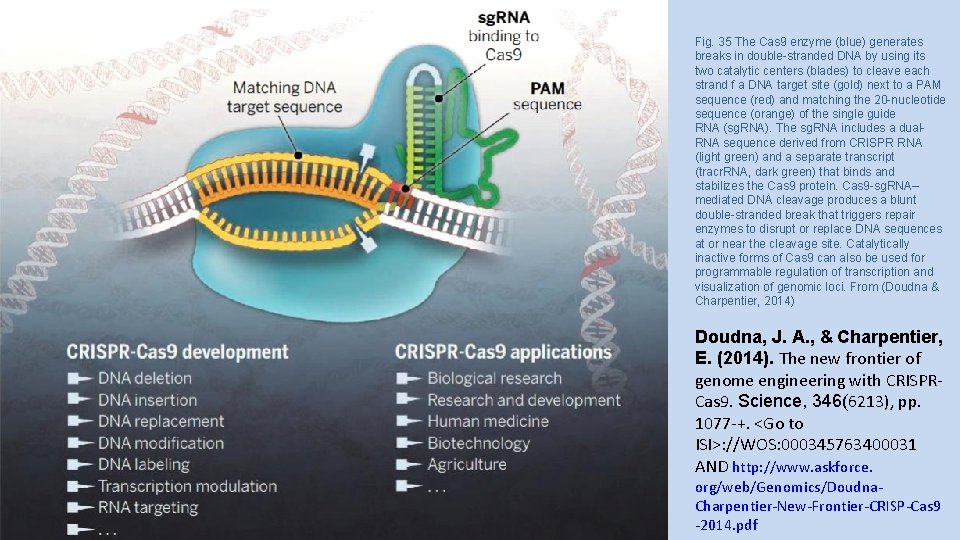
Fig. 35 The Cas 9 enzyme (blue) generates breaks in double-stranded DNA by using its two catalytic centers (blades) to cleave each strand f a DNA target site (gold) next to a PAM sequence (red) and matching the 20 -nucleotide sequence (orange) of the single guide RNA (sg. RNA). The sg. RNA includes a dual. RNA sequence derived from CRISPR RNA (light green) and a separate transcript (tracr. RNA, dark green) that binds and stabilizes the Cas 9 protein. Cas 9 -sg. RNA– mediated DNA cleavage produces a blunt double-stranded break that triggers repair enzymes to disrupt or replace DNA sequences at or near the cleavage site. Catalytically inactive forms of Cas 9 can also be used for programmable regulation of transcription and visualization of genomic loci. From (Doudna & Charpentier, 2014) Doudna, J. A. , & Charpentier, E. (2014). The new frontier of genome engineering with CRISPRCas 9. Science, 346(6213), pp. 1077 -+. <Go to ISI>: //WOS: 000345763400031 AND http: //www. askforce. org/web/Genomics/Doudna. Charpentier-New-Frontier-CRISP-Cas 9 -2014. pdf

CRISPR-Cas 9 applications in plants and fungi also promise to change the pace and course of agricultural research. Future research directions to improve the technology will include engineering or identifying smaller Cas 9 variants with distinct specificity that may be more amenable to delivery in human cells. Understanding the homology-directed repair mechanisms that follow Cas 9 -mediated DNA cleavage will enhance insertion of new or corrected sequences into genomes. The development of specific methods for efficient and safe delivery of Cas 9 and its guide RNAs to cells and tissues will also be critical for applications of the technology in human gene therapy.
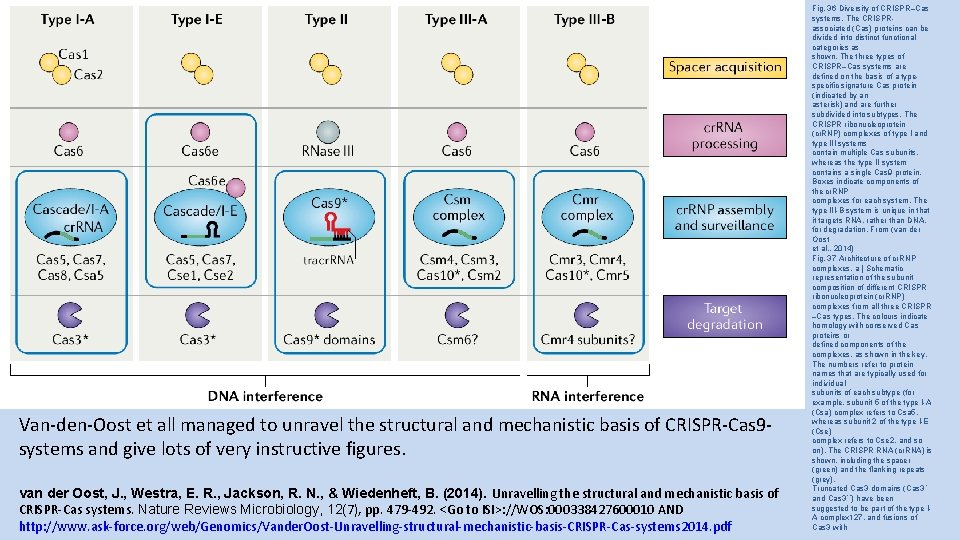
Van-den-Oost et all managed to unravel the structural and mechanistic basis of CRISPR-Cas 9 systems and give lots of very instructive figures. van der Oost, J. , Westra, E. R. , Jackson, R. N. , & Wiedenheft, B. (2014). Unravelling the structural and mechanistic basis of CRISPR-Cas systems. Nature Reviews Microbiology, 12(7), pp. 479 -492. <Go to ISI>: //WOS: 000338427600010 AND http: //www. ask-force. org/web/Genomics/Vander. Oost-Unravelling-structural-mechanistic-basis-CRISPR-Cas-systems 2014. pdf Fig. 36 Diversity of CRISPR–Cas systems. The CRISPRassociated (Cas) proteins can be divided into distinct functional categories as shown. The three types of CRISPR–Cas systems are defined on the basis of a typespecific signature Cas protein (indicated by an asterisk) and are further subdivided into subtypes. The CRISPR ribonucleoprotein (cr. RNP) complexes of type I and type III systems contain multiple Cas subunits, whereas the type II system contains a single Cas 9 protein. Boxes indicate components of the cr. RNP complexes for each system. The type III-B system is unique in that it targets RNA, rather than DNA, for degradation. From (van der Oost et al. , 2014) Fig. 37 Architecture of cr. RNP complexes. a | Schematic representation of the subunit composition of different CRISPR ribonucleoprotein (cr. RNP) complexes from all three CRISPR –Cas types. The colours indicate homology with conserved Cas proteins or defined components of the complexes, as shown in the key. The numbers refer to protein names that are typically used for individual subunits of each subtype (for example, subunit 5 of the type I-A (Csa) complex refers to Csa 5, whereas subunit 2 of the type I-E (Cse) complex refers to Cse 2, and so on). The CRISPR RNA (cr. RNA) is shown, including the spacer (green) and the flanking repeats (grey). Truncated Cas 3 domains (Cas 3ʹ and Cas 3ʹʹ) have been suggested to be part of the type IA complex 127, and fusions of Cas 3 with
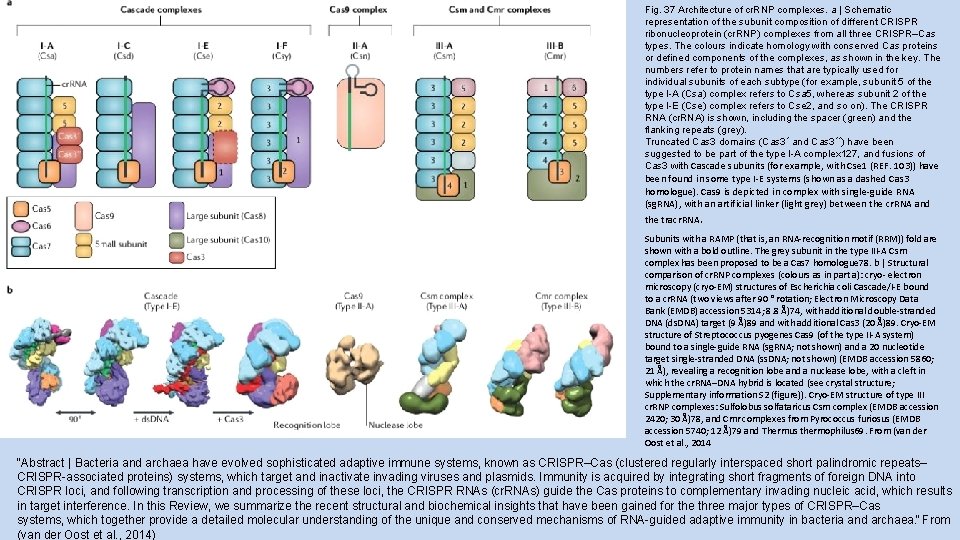
Fig. 37 Architecture of cr. RNP complexes. a | Schematic representation of the subunit composition of different CRISPR ribonucleoprotein (cr. RNP) complexes from all three CRISPR–Cas types. The colours indicate homology with conserved Cas proteins or defined components of the complexes, as shown in the key. The numbers refer to protein names that are typically used for individual subunits of each subtype (for example, subunit 5 of the type I-A (Csa) complex refers to Csa 5, whereas subunit 2 of the type I-E (Cse) complex refers to Cse 2, and so on). The CRISPR RNA (cr. RNA) is shown, including the spacer (green) and the flanking repeats (grey). Truncated Cas 3 domains (Cas 3ʹ and Cas 3ʹʹ) have been suggested to be part of the type I-A complex 127, and fusions of Cas 3 with Cascade subunits (for example, with Cse 1 (REF. 103)) have been found in some type I-E systems (shown as a dashed Cas 3 homologue). Cas 9 is depicted in complex with single-guide RNA (sg. RNA), with an artificial linker (light grey) between the cr. RNA and the tracr. RNA. Subunits with a RAMP (that is, an RNA-recognition motif (RRM)) fold are shown with a bold outline. The grey subunit in the type III-A Csm complex has been proposed to be a Cas 7 homologue 78. b | Structural comparison of cr. RNP complexes (colours as in part a): cryo- electron microscopy (cryo-EM) structures of Escherichia coli Cascade/I-E bound to a cr. RNA (two views after 90 ° rotation; Electron Microscopy Data Bank (EMDB) accession 5314; 8. 8 Å)74, with additional double-stranded DNA (ds. DNA) target (9 Å)89 and with additional Cas 3 (20 Å)89. Cryo-EM structure of Streptococcus pyogenes Cas 9 (of the type II-A system) bound to a single-guide RNA (sg. RNA; not shown) and a 20 nucleotide target single-stranded DNA (ss. DNA; not shown) (EMDB accession 5860; 21 Å), revealing a recognition lobe and a nuclease lobe, with a cleft in which the cr. RNA–DNA hybrid is located (see crystal structure; Supplementary information S 2 (figure)). Cryo-EM structure of type III cr. RNP complexes: Sulfolobus solfataricus Csm complex (EMDB accession 2420; 30 Å)78, and Cmr complexes from Pyrococcus furiosus (EMDB accession 5740; 12 Å)79 and Thermus thermophilus 69. From (van der Oost et al. , 2014 “Abstract | Bacteria and archaea have evolved sophisticated adaptive immune systems, known as CRISPR–Cas (clustered regularly interspaced short palindromic repeats– CRISPR-associated proteins) systems, which target and inactivate invading viruses and plasmids. Immunity is acquired by integrating short fragments of foreign DNA into CRISPR loci, and following transcription and processing of these loci, the CRISPR RNAs (cr. RNAs) guide the Cas proteins to complementary invading nucleic acid, which results in target interference. In this Review, we summarize the recent structural and biochemical insights that have been gained for the three major types of CRISPR–Cas systems, which together provide a detailed molecular understanding of the unique and conserved mechanisms of RNA-guided adaptive immunity in bacteria and archaea. ” From (van der Oost et al. , 2014)
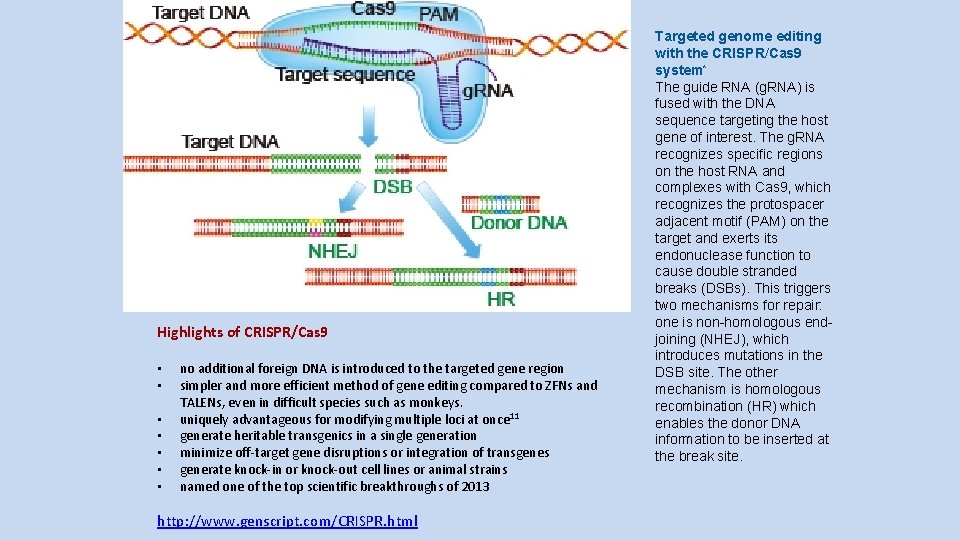
Highlights of CRISPR/Cas 9 • • no additional foreign DNA is introduced to the targeted gene region simpler and more efficient method of gene editing compared to ZFNs and TALENs, even in difficult species such as monkeys. uniquely advantageous for modifying multiple loci at once 11 generate heritable transgenics in a single generation minimize off-target gene disruptions or integration of transgenes generate knock-in or knock-out cell lines or animal strains named one of the top scientific breakthroughs of 2013 http: //www. genscript. com/CRISPR. html Targeted genome editing with the CRISPR/Cas 9 system* The guide RNA (g. RNA) is fused with the DNA sequence targeting the host gene of interest. The g. RNA recognizes specific regions on the host RNA and complexes with Cas 9, which recognizes the protospacer adjacent motif (PAM) on the target and exerts its endonuclease function to cause double stranded breaks (DSBs). This triggers two mechanisms for repair: one is non-homologous endjoining (NHEJ), which introduces mutations in the DSB site. The other mechanism is homologous recombination (HR) which enables the donor DNA information to be inserted at the break site.

DOZENS OF NEW GENE-EDITING METHODS WILL CAUSE A REGULATORY REVOLUTION Fig. 5 Engineered nuclease-mediated genome editing. Engineered nucleases such as the CRISPR-Cas 9 or TAL effectors can be designed to target specific sites in the genome, creating double-strand breaks (DSBs) at desired locations. The natural repair mechanisms of the cell repair the break by either homologous recombination (HR) or non -homologous end joining (NHEJ). HR is more precise, since it requires a template, allowing the introduction of foreign DNA into the target gene. Homologous DNA “donor sequences” can be used with homologydirected repair (HDR) to introduce a defined new DNA sequence. DSB repair by NHEJ is likely to introduce errors such as insertions or deletions (indels), leading to a nonfunctional gene. From (Life Technologies, 2015) Ammann Klaus, & Kuntz Marcel. (12. January 2016). Decades-old GMO regulation unfit for 21 st century, En, Fr, It, Dt. Eur. Activ, 3 p, Brussels, Eur. Activ, http: //www. ask-force. org/web/Genomic-Misconception/Ammann-Kuntz-Euractiv-Gene-Editing-Debate. English-original-20160112. pdf

The origins of CRISPR goes back to a range of selected papers (which might still be incomplete): The CRISPR dynamics has been discovered before it got its name: The first mention of early discovery goes to Ishino Y. et al. (Ishino Y. et al. ): In 1987, Yoshizumi Ishino and his colleagues at Osaka University in Japan published the sequence of a peculiar short repeat, called iap, in the DNA of E. col: Ishino, Y. , Shinagawa, H. , Makino, K. , Amemura, M. , & Nakata, A. (1987). Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. Journal of Bacteriology, 169(12), pp. 5429 -5433. http: //jb. asm. org/content/169/12/5429. abstract AND http: //www. ask-force. org/web/Genomics/Ishino-Nucleotide-Sequence-iap-Gene-Conversion-Ec-identification-Gene-Product-1987. pdf “The iap gene in Escherichia coli is responsible for the isozyme conversion of alkaline phosphatase. We analyzed the 1, 664 -nucleotide sequence of a chromosomal DNA segment that contained the iap gene and its flanking regions. The predicted iap product contained 345 amino acids with an estimated molecular weight of 37, 919. The 24 -aminoacidsequence at the amino terminus showed features characteristic of a signal peptide. Two proteins of different sizes were identified by the maxicell method, one corresponding to the lap protein and the other corresponding to the processed product without the signal peptide. Neither the isozyme-converting activity nor labeled Iap proteins were detected in the osmotic-shock fluid of cells carrying a multicopy iap plasmid. The Iap protein seems to be associated with the membrane. ” From Ishino et al. (1987) The source of a lot of interesting and precise data and events on the birth of CRISPR can be found in an interview of Bob Grant with George Church: Grant Bob. (20151229). Credit for CRISPR: A Conversation with George Church. The media frenzy over the gene-editing technique highlights shortcomings in how journalists and award committees portray contributions to scientific discoveries. The Scientist, 29. December 2015, pp. 3. http: //www. the-scientist. com/? articles. view/article. No/44919/title/Credit-for-CRISPR--A-Conversation-with-George-Church/ AND http: //www. askforce. org/web/Genomics/Grant-Church-Credit-for-CRISPR-2015. pdf

Paul Anderson, best video explaining the new Gene Editing with CRISPR-Cas 9 https: //www. youtube. com/watch? v=Mn. Yppmstx. Is&app=desktop Published on 18 Feb 2016 In this video Paul Andersen explains how the CRISPR/Cas immune system was identified in bacteria and how the CRISPR/Cas 9 system was developed to edit genomes. FIRST MENTION OF CRISPR IN A PUBLICATIOIN Mojica, F. J. M. , Díez-Villaseñor, C. , García-Martínez, J. , & Soria, E. (2005). Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol. , 60, pp. http: //dx. doi. org/10. 1007/s 00239 -0046 -3 AND http: //www. askforce. org/web/Genomics/Mojina-Intervening. Sequences-Regularly-Spaced-Prokaryotic-Repeats-2005. pdf “Prokaryotes contain short DNA repeats known as CRISPR, recognizable by the regular spacing existing between the recurring units. They represent the most widely distributed family of repeats among prokaryotic genomes, suggesting a biological function.

FIRST MENTION OF CRISPR IN A PUBLICATIOIN Mojica, F. J. M. , Díez-Villaseñor, C. , García-Martínez, J. , & Soria, E. (2005). Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol. , 60, pp. http: //dx. doi. org/10. 1007/s 00239 -0046 -3 AND http: //www. askforce. org/web/Genomics/Mojina-Intervening-Sequences. Regularly-Spaced-Prokaryotic-Repeats-2005. pdf “Prokaryotes contain short DNA repeats known as CRISPR, recognizable by the regular spacing existing between the recurring units. They represent the most widely distributed family of repeats among prokaryotic genomes, suggesting a biological function

Lundgren Magnus, Charpentier Emmanuelle, & Fineran Peter C. (Eds. ). (2015). CRISPR, Methods and Protocols, Springer Science+Business Media LLC New York is part of Springer Science+Business Media www. springer. com Humana Press is a brand of Springer. DOI 10. 1007/978 -1 -4939 -2687 -9 AND http: //rd. springer. com/book/10. 1007/978 -1 -4939 -2687 -9 AND http: //www. ask-force. org/web/Genomics/Lundgren-CRISPR-Methods-Protocols-2015. pdf Quetier, F. (2016). The CRISPR-Cas 9 technology: Closer to the ultimate toolkit for targeted genome editing. Plant Science, 242, pp. 65 -76. <Go to ISI>: //WOS: 000367106100007 AND http: //www. ask-force. org/web/Genomics/Quetier-CRISP-Cas 9 technology-closer-to-ultimate-toolkit-2016. pdf “The first period of plant genome editing was based on Agrobacterium; chemical mutagenesis by EMS (ethyl methanesulfonate) and ionizing radiations; each of these technologies led to randomly distributed genome modifications. The second period is associated with the discoveries of homing and meganuclease enzymes during the 80 s and 90 s, which were then engineered to provide efficient tools for targeted editing. From 2006 to 2012, a few crop plants were successfully and precisely modified using zincfinger nucleases. A third wave of improvement in genome editing, which led to a dramatic decrease in off-target events, was achieved in 2009– 2011 with the TALEN technology. The latest revolution surfaced in 2013 with the CRISPR-Cas 9 system, whose high efficiency and technical ease of use is really impressive; scientists can use in-house kits or commercially available kits; the only two requirements are to carefully choose the location of the DNA double strand breaks to be induced and then to order an oligonucleotide. While this close-to- ultimate toolkit for targeted editing of genomes represents dramatic scientific progress which allows the development of more complex useful agronomic traits through synthetic biology, the social acceptance of genome editing remains regularly questioned by anti-GMO citizens and organizations. ” From (Quetier, 2016)

Ammann Klaus, & Kuntz Marcel. (20160112, 12. January 2016). Decades-old GMO regulation unfit for 21 st century, En, Fr, It, Dt. Eur. Activ, 3 p, Brussels, Eur. Activ, original link: http: //www. ask-force. org/web/Genomic-Misconception/Ammann-Kuntz-Euractiv-Gene-Editing-Debate-English-original-20160112. pdf



Natural GMO? Sweet Potato Genetically Modified 8, 000 Years Ago Tyna Kynth Gent University Belgium Main author Of PNAS paper http: //www. npr. org/sections/goatsandsoda/2015/05/05/404198552 natural-gmo-sweet-potato-genetically-modified-8 -000 -years-ago Kyndt, T. , Quispe, D. , Zhai, H. , Jarret, R. , Ghislain, M. , Liu, Q. , Gheysen, G. , & Kreuze, J. F. (2015). The genome of cultivated sweet potato contains Agrobacterium T-DNAs with expressed genes: An example of a naturally transgenic food crop. Proceedings of the National Academy of Sciences, http: //www. ask-force. org/web/Genomics/Kynth-Genome-Cultivated-Sweet-Potato-Naturally-Transgenic-2015. pdf For those who cannot get enough science, watch http: //www. scoop. it/t/articles-published-by-cip-staff/? tag=Sweet+potatoes See also the collection of natural transgenic plants of David Tribe from Melbourne http: //gmopundit. blogspot. ch/search/label/Natural%20 GMOs
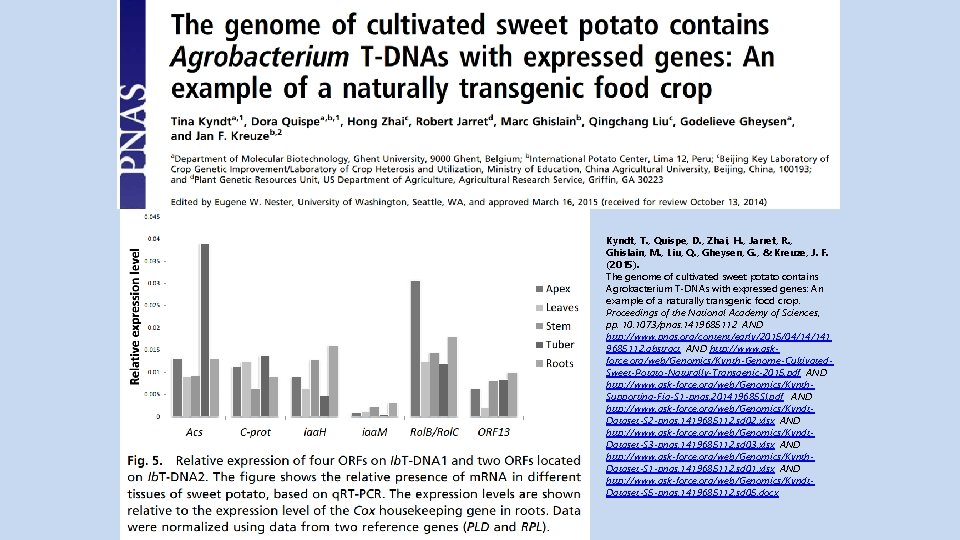
Kyndt, T. , Quispe, D. , Zhai, H. , Jarret, R. , Ghislain, M. , Liu, Q. , Gheysen, G. , & Kreuze, J. F. (2015). The genome of cultivated sweet potato contains Agrobacterium T-DNAs with expressed genes: An example of a naturally transgenic food crop. Proceedings of the National Academy of Sciences, pp. 1073/pnas. 1419685112 AND http: //www. pnas. org/content/early/2015/04/14/141 9685112. abstract AND http: //www. askforce. org/web/Genomics/Kynth-Genome-Cultivated. Sweet-Potato-Naturally-Transgenic-2015. pdf AND http: //www. ask-force. org/web/Genomics/Kynth. Supporting-Fig-S 1 -pnas. 201419685 SI. pdf AND http: //www. ask-force. org/web/Genomics/Kyndt. Dataset-S 2 -pnas. 1419685112. sd 02. xlsx AND http: //www. ask-force. org/web/Genomics/Kyndt. Dataset-S 3 -pnas. 1419685112. sd 03. xlsx AND http: //www. ask-force. org/web/Genomics/Kynth. Dataset-S 1 -pnas. 1419685112. sd 01. xlsx AND http: //www. ask-force. org/web/Genomics/Kyndt. Dataset-S 5 -pnas. 1419685112. sd 05. docx

Urban Myth Transgenic and non-transgenic crops are basically different Wrong: the molecular processes are with both the same Ammann, K. (20120706) Genomic Misconception: A fresh look at the biosafety of transgenic and conventional crops, a plea for a process agnostic regulation New Biotechnology, in press, pp 32 http: //www. ask-force. org/web/New. Biotech/Genomic-Misconception-20120706 -names-def. pdf

Interestingly, naturally occurring molecular evolution, i. e. the spontaneous generation of genetic variants has been seen to follow exactly the same three strategies as those used in genetic engineering 14. These three strategies are (after W. Arber, Nobel Laureate 1978) (a) small local changes in the nucleotide sequences, (b) internal reshuffling of genomic DNA segments, and (c) acquisition of usually rather small segments of DNA from another type of organism by horizontal gene transfer. Arber, W. (2002) Roots, strategies and prospects of functional genomics. Current Science, 83, 7, pp 826 -828 http: //www. botanischergarten. ch/Mutations/Arber-Comparison-2002. pdf Arber, W. (2010) Genetic engineering compared to natural genetic variations. New Biotechnology, 27, 5, pp 517 -521 http: //www. ask-force. org/web/Vatican-PAS-Studyweek-Elsevier-publ-20101130/Arber-Werner-PAS-Genetic-Engineering. Compared-20101130 -publ. pdf

However, there is a principal difference between the procedures of genetic engineering and those serving in nature for biological evolution. While the genetic engineer pre-reflects his alteration and verifies its results, nature places its genetic variations more randomly and largely independent of an identified goal. After ca. 15 years of testing the GM crops are brought to the field by millions in a few years Arber, W. (2002) Roots, strategies and prospects of functional genomics. Current Science, 83, 7, pp 826 -828 http: //www. botanischergarten. ch/Mutations/Arber-Comparison-2002. pdf Ammann Klaus. (2014). Genomic Misconception: a fresh look at the biosafety of transgenic and conventional crops. A plea for a process agnostic regulation. New Biotechnology, 31(1), pp. 1 -17. http: //dx. doi. org/10. 1016/j. nbt. 2013. 04. 008 AND open source: http: //www. askforce. org/web/New. Biotech/Genomic-Misconception-new-20140821 -names-links. pdf AND http: //www. ask -force. org/web/New. Biotech/Ammann-Genomic-Misconception-printed-2014. pdf

Pontifical Academy of Science Vatican Conference on Transgenic Plants for Food Security in the Context of Development Bishop Marcelo Sanchez – Sorondo, Secretary, Prof. Dr. Werner Arber, President, Nobel Laureate Potrykus, I. and K. Ammann (2010), Transgenic Plants for Food Security in the Context of Development, Conference Proceedings and Statements of the Pontifical Academy of Sciences, M. Taussig, New Biotechnology, 27, open source, pp. 445 -717, Elsevier, Amsterdam, http: //www. sciencedirect. com/science/issue/43660 -2010 -999729994 -2699796 AND on Vatican Website http: //www. vatican. va/roman_curia/pontifical_academies/acdscien/2010/newbiotechnologynov 2010. pdf AND on Host ASK-FORCE: http: //www. ask-force. org/web/Vatican-PAS-Studyweek-Elsevier-publ-20101130/Potrykus-Ammann-Conference-Volume-Newbiotechnology 2010. pdf Press release http: //www. ask-force. org/web/Vatican-PAS-Studyweek-Elsevier-publ-20101130/Press-Release-PAS-Studyweek 20101127. pdf

Full bibliography of the open source volume of NEW BIOTECHNOLOGY, Elsevier 27/5, p. 445 -718, November 30, 2010 All published papers, statements and conference presentations in: http: //www. sciencedirect. com/science/issue/43 660 -2010 -999729994 -2699796 It must be understood, that statements by the participants regarding the event do not constitute the opinion of the Vatican or the Pontifical Academy of Sciences. The official information, beyond any interview, is laid out in the English version of the ‚Statement’ agreed upon unanimously by all participants http: //www. ask-force. org/web/PAS-Statement. English. pdf and in additional 15 world languages, see link above For interviews contact Prof. em. Ingo Potrykus ingo@potrykus. ch or Prof. em. Klaus Ammann, klaus. ammann@ips. unibe. ch or anybody else from the participants list: http: //www. ask-force. org/web/Participants-List 2010. pdf
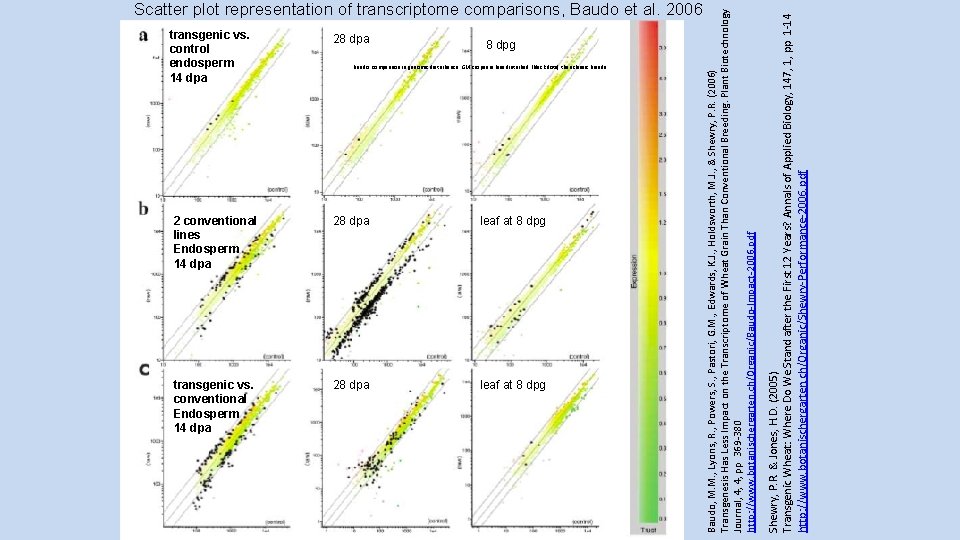
transgenic vs. control endosperm 14 dpa 28 dpa 8 dpg Baudo: comparison in genomic disturbance: GM crops are less disturbed (black dots) than classic breeds 2 conventional lines Endosperm 14 dpa 28 dpa leaf at 8 dpg transgenic vs. conventional Endosperm 14 dpa 28 dpa leaf at 8 dpg Shewry, P. R. & Jones, H. D. (2005) Transgenic Wheat: Where Do We Stand after the First 12 Years? Annals of Applied Biology, 147, 1, pp 1 -14 http: //www. botanischergarten. ch/Organic/Shewry-Performance-2006. pdf Baudo, M. M. , Lyons, R. , Powers, S. , Pastori, G. M. , Edwards, K. J. , Holdsworth, M. J. , & Shewry, P. R. (2006) Transgenesis Has Less Impact on the Transcriptome of Wheat Grain Than Conventional Breeding. Plant Biotechnology Journal, 4, 4, pp 369 -380 http: //www. botanischergarten. ch/Organic/Baudo-Impact-2006. pdf Scatter plot representation of transcriptome comparisons, Baudo et al. 2006

Gamma Field for radiation breeding 100 m radius Radiation breeding as field experiments 60 min 89 TBq Co-60 source at the center Shielding dike 8 m high 4 min Better Institute of Radiation Breeding Ibaraki-ken, JAPAN http: //www. irb. affrc. go. jp/ spaghettis, whisky 1800 new plants

Reuters, May 10, 2010 UN's International Atomic Energy Agency since 1963, 3217 new plant varieties, including Italian durum wheat, have been created using radioactive substances such as cobalt and X-rays. 70% of the crops under cultivation worldwide are radiation mutation varieties Charles Margulis of Greenpeace USA: "But now they tell us that scientists have been artificially hybridizing plants since the 1960 s. That's, like, really uncool. "

Activists, supported by Jane Rissler, called for a ban, since those irradiated varieties have never been tested for food safety, which would have wiped out 70% of the food products on shelfs. Rissler: “Compared to these plants, genetically modified food is about as dangerous as a one-legged man in an ass-kicking contest. ” But excellent repair mechanisms working like zippers are reducing radiation damage considerably And worldwide there has been no correlation established between radiation mutation and negative food safety facts. (Reuters 2001 continued)

FRANKENSTEIN Durum Wheat, Triticum durum: all major breeds have gone though massive and inprecise radiation breeding, but with Important success unnecessary fearmongering

http: //cen. acs. org/articles/93/i 40/Tomas-Lindahl-Paul-Modrich-Aziz. html Lindahl, T. (2013). My journey to DNA repair. Genomics, proteomics & bioinformatics, 11(1), pp. 2 -7. <Go to ISI>: //MEDLINE: 23453014 AND http: //www. ask-force. org/web/Genomics/Lindahl-My-Journey-to-DNA-Repair 2013. pdf Friedberg, E. C. (2003). DNA damage and repair. Nature, 421(6921), pp. 436 -440. http: //dx. doi. org/10. 1038/nature 01408 AND http: //www. askforce. org/web/Genomics/Friedberg-DNA-damage-repair-Nature-2003. pdf AND http: //www. nature. com/collections/wqswtxswtg

Following DNA damage, the cell must first recognize the damage and then alert the system that there’s a problem. The recognition machinery then activates various factors to halt cell growth until the damage has been repaired. And if things are too far gone, additional factors are poised and ready to induce cell death. That’s the most basic way to think about the DNA damage response pathway, as a simple chain of events. Of course it’s a lot more complicated, a complex network of checks and balances to ensure that the DNA damage is not only recognized but clearly identified to ensure that the correct factors are recruited to repair the lesion. https: //theconversation. com/chemistry-nobel-dna-research-laysfoundation-for-new-ways-to-fight-cancer-48800 And https: //theconversation. com/nobel-prize-in-chemistry-highlightshow-our-bodies-can-repair-our-fragile-dna-48816

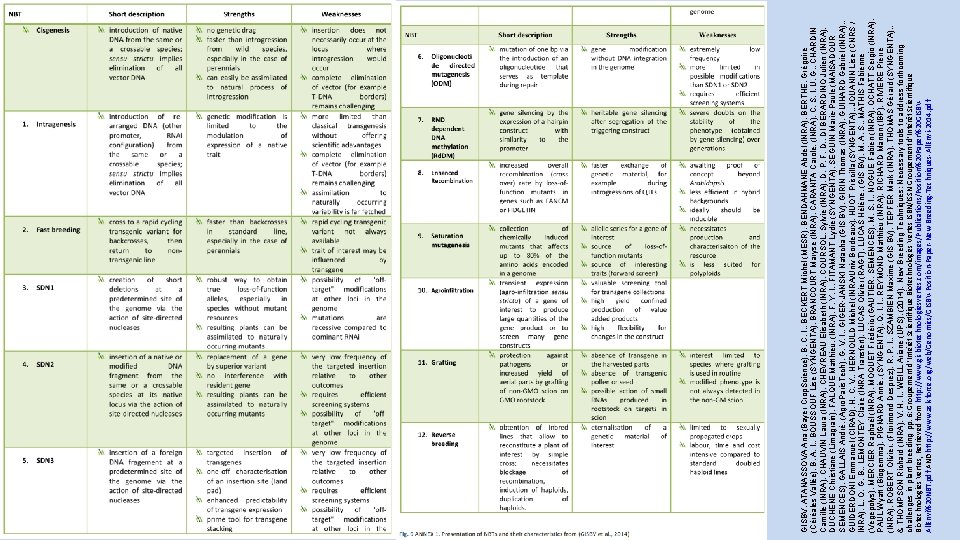
GISBV, ATANASSOVA Ana (Bayer Crop. Science), B. C. I. , BECKERT Michel (MESR), BENDAHMANE Abdel (INRA), BERTHE, Grégoire (Céréales Vallée), B. A. I. , BOUSSOUF Lise (SYNGENTA), BRANCOURT Maryse (INRA), CARANTA Carole, (INRA), C. S. I. U. G. , CHARDIN Camille (INRA), CHAUVIN Laura (INRA), CHEVREAU Elisabeth (INRA), COURSOL, Sylvie (INRA), D. P. F. D. , DI BERARDINO Julien (INRA), DUCHENE Christiane (Limagrain), FALQUE Matthieu, (INRA), F. Y. I. , FITAMANT Lydie (SYNGENTA), SEGUIN Marie-Paule (MAISADOUR SEMENCES), GALLAIS André, (Agro. Paris. Tech), G. V. I. , GIGER-JANSKI Natacha (GIS BV), GIRIN Thomas (INRA), GUIHARD Gabriel (INRA), , GUIDERDONI Emmanuel (CIRAD), H. C. V. , HERNOULD Michel (INRA/Univ Bordeaux), HUOT Priscilla (SYNGENTA), , JOUANIN Lise (CNRS / INRA), L. O. G. B. , LEMONTEY Claire (INRA Transfert), LUCAS Olivier (RAGT), LUCAS Hélène, (GIS BV), M. A. S. , MATHIS Fabienne (Vegepolys), MERCIER Raphaël (INRA), MOQUET Frédéric (GAUTIER, SEMENCES), M. S. I. , NOGUE Fabien (INRA), OCHATT Sergio (INRA), PAUL Wyatt (Biogemma), PIGNARD Annie, (SYNGENTA), Q. I. I. , REYMOND Matthieu (INRA), RICHARD Manon (IBP), RIVIERE Pierre (INRA), ROBERT Olivier, (Florimond Desprez), R. P. I. , SZAMBIEN Maxime (GIS BV), TEPFER Mark (INRA), THOMAS Gérard (SYNGENTA), , & THOMPSON Richard (INRA), V. H. I. , WEILL Ariane (UPS). (2014). New Breeding Techniques: Necessary tools to address forthcoming challenges in plant breeding pp. 6: Groupement d’Intérêt Scientifique Biotechnologies Vertes ISBN/ISSN Groupement d’Intérêt Scientifique Biotechnologies Vertes, Retrieved from http: //www. gisbiotechnologiesvertes. com/images/Publications/Position%20 Paper%20 GISBVAll. Envi%20 NBT. pdf AND http: //www. ask-force. org/web/Genomics/GISBV-Position-Paper-New-Breeding-Techniques-All. Envi-2014. pdf
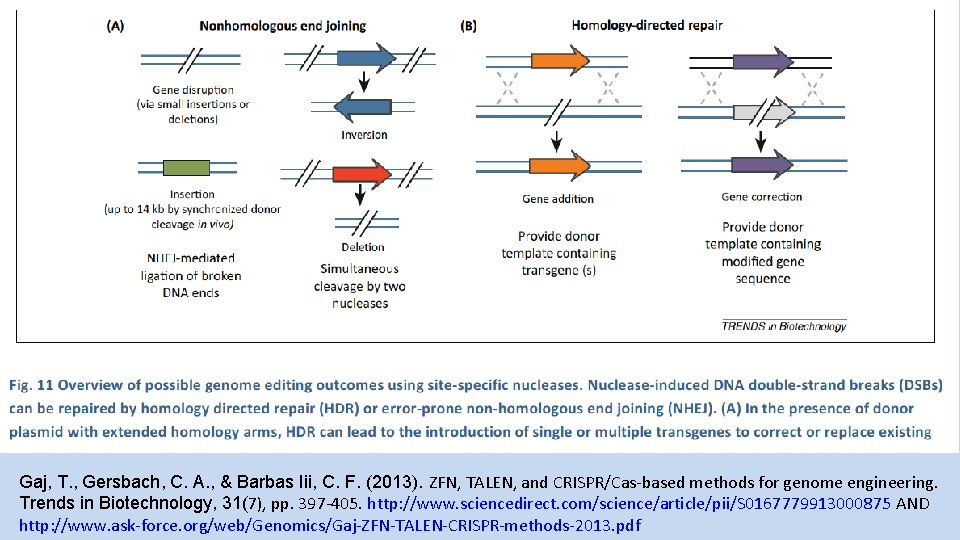
Gaj, T. , Gersbach, C. A. , & Barbas Iii, C. F. (2013). ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends in Biotechnology, 31(7), pp. 397 -405. http: //www. sciencedirect. com/science/article/pii/S 0167779913000875 AND http: //www. ask-force. org/web/Genomics/Gaj-ZFN-TALEN-CRISPR-methods-2013. pdf



Qasim Waseem, M. P. , Persis Jal Amrolia 2*, Sujith Samarasinghe, MD, Ph. D 3*, Sara Ghorashian, MD, Ph. D, FRCPath 1*, Hong Zhan, Ph. D 4*, Sian Stafford, PHD 1*, Katie Butler, PHD 1*, Gul Ahsan 5*, Kimberly Gilmour 5*, Stuart Adams, PHD 5*, Danielle Pinner 5*, Robert Chiesa 5*, Steve Chatters, PHD 5*, Sue Swift, PHD 1*, Nicholas Goulden, MD, Ph. D 3, Karl Peggs, MBBChir, MRCP, FRCPath 6*, Adrian J Thrasher, MD, Ph. D 1*, Paul Veys 2* and Martin Pule, Ph. D 7. (20151206). 2046 First Clinical Application of Talen Engineered Universal CAR 19 T Cells in B-ALL Poster. Paper presented at the ASH 57 th Annual Meeting and Exposition, Orlando, Florida USA. http: //www. askforce. org/web/Genomics/Qasim-First-Clinical-Application-TALEN-Leucaemia-20151205. pdf Gene editing saves girl dying from leukaemia
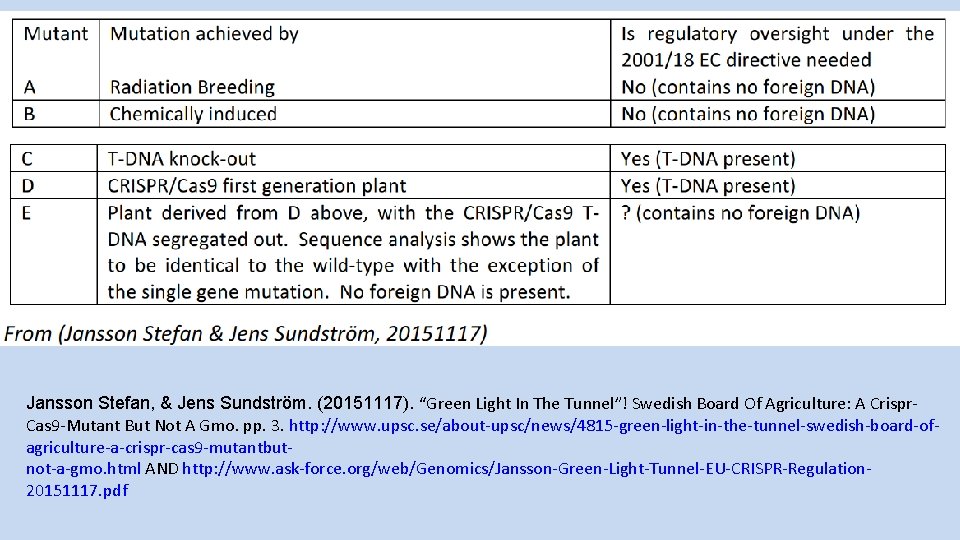
Jansson Stefan, & Jens Sundström. (20151117). “Green Light In The Tunnel”! Swedish Board Of Agriculture: A Crispr. Cas 9 -Mutant But Not A Gmo. pp. 3. http: //www. upsc. se/about-upsc/news/4815 -green-light-in-the-tunnel-swedish-board-ofagriculture-a-crispr-cas 9 -mutantbutnot-a-gmo. html AND http: //www. ask-force. org/web/Genomics/Jansson-Green-Light-Tunnel-EU-CRISPR-Regulation 20151117. pdf
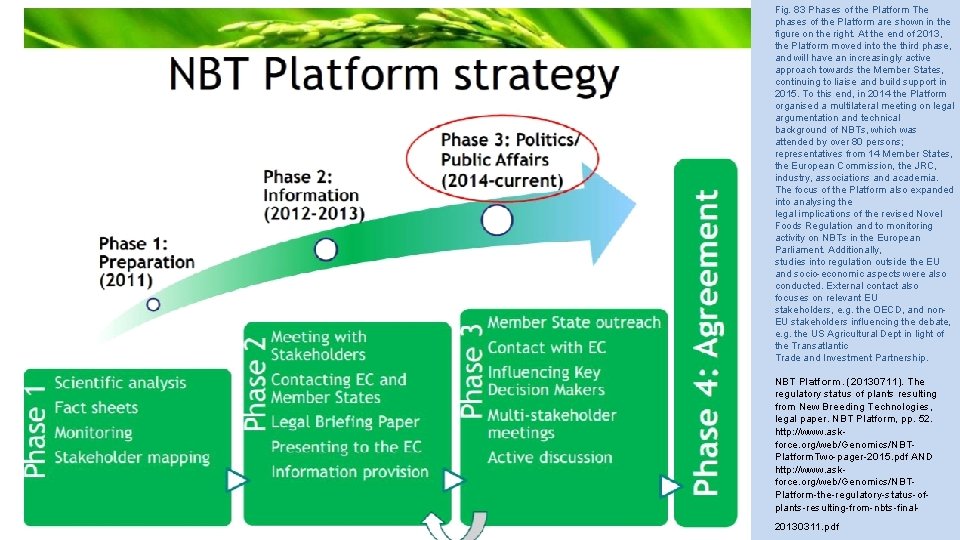
Fig. 83 Phases of the Platform The phases of the Platform are shown in the figure on the right. At the end of 2013, the Platform moved into the third phase, and will have an increasingly active approach towards the Member States, continuing to liaise and build support in 2015. To this end, in 2014 the Platform organised a multilateral meeting on legal argumentation and technical background of NBTs, which was attended by over 80 persons; representatives from 14 Member States, the European Commission, the JRC, industry, associations and academia. The focus of the Platform also expanded into analysing the legal implications of the revised Novel Foods Regulation and to monitoring activity on NBTs in the European Parliament. Additionally, studies into regulation outside the EU and socio-economic aspects were also conducted. External contact also focuses on relevant EU stakeholders, e. g. the OECD, and non. EU stakeholders influencing the debate, e. g. the US Agricultural Dept in light of the Transatlantic Trade and Investment Partnership. NBT Platform. (20130711). The regulatory status of plants resulting from New Breeding Technologies, legal paper. NBT Platform, pp. 52. http: //www. askforce. org/web/Genomics/NBTPlatform. Two-pager-2015. pdf AND http: //www. askforce. org/web/Genomics/NBTPlatform-the-regulatory-status-ofplants-resulting-from-nbts-final 20130311. pdf


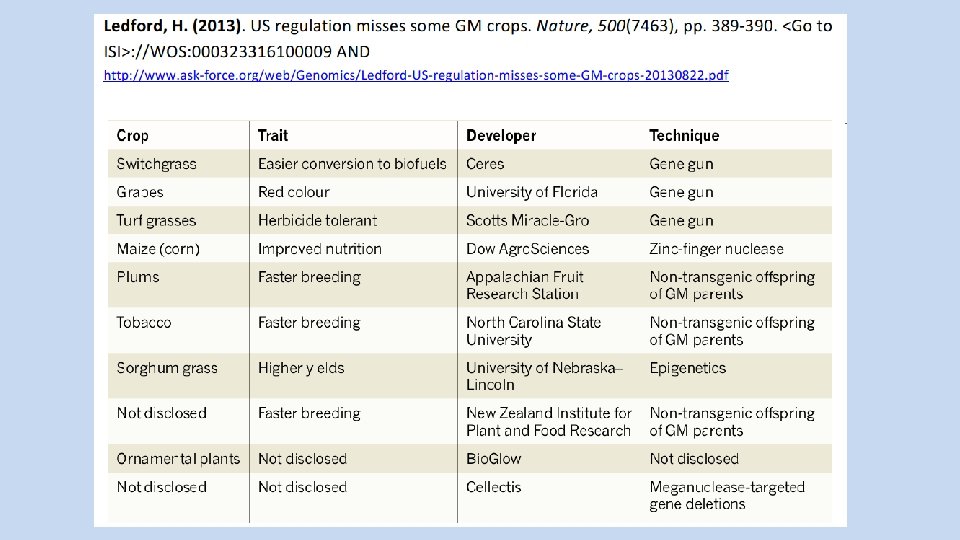
Ledford Heidi. (20160414). US rethinks crop regulation. Committee begins study to guide oversight of gene-edited organisms. Nature, 532, pp. 159 -159. http: //www. nature. com/polopoly_fs/1. 19724!/menu/main/top. Columns/top. Left. Column/pdf/532158 a. pdf AND http: //www. askforc/www/ask-force. org/web/Genomics/Ledford-US-rethinks-crop-regulation-Nature-2016. pd
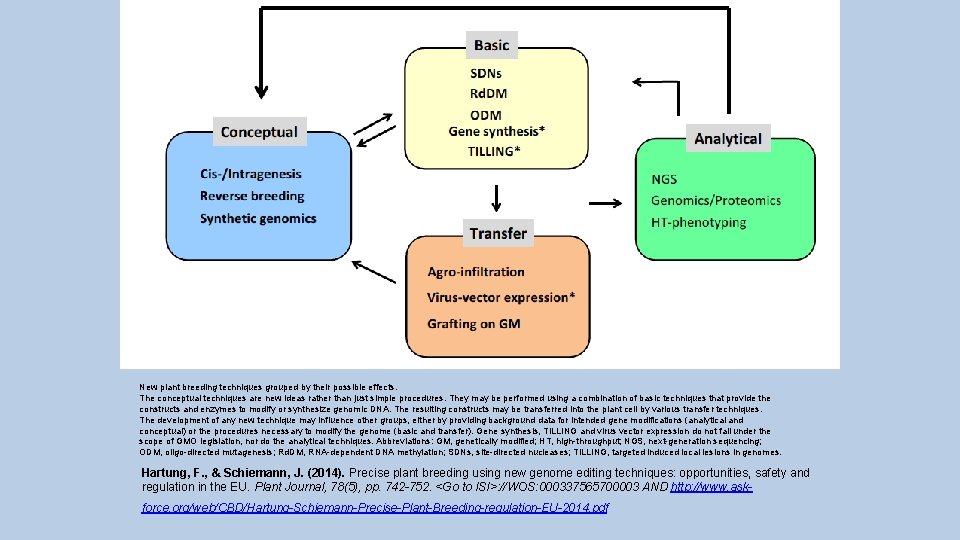
New plant breeding techniques grouped by their possible effects. The conceptual techniques are new ideas rather than just simple procedures. They may be performed using a combination of basic techniques that provide the constructs and enzymes to modify or synthesize genomic DNA. The resulting constructs may be transferred into the plant cell by various transfer techniques. The development of any new technique may influence other groups, either by providing background data for intended gene modifications (analytical and conceptual) or the procedures necessary to modify the genome (basic and transfer). Gene synthesis, TILLING and virus vector expression do not fall under the scope of GMO legislation, nor do the analytical techniques. Abbreviations: GM, genetically modified; HT, high-throughput; NGS, next-generation sequencing; ODM, oligo-directed mutagenesis; Rd. DM, RNA-dependent DNA methylation; SDNs, site-directed nucleases; TILLING, targeted induced local lesions in genomes. Hartung, F. , & Schiemann, J. (2014). Precise plant breeding using new genome editing techniques: opportunities, safety and regulation in the EU. Plant Journal, 78(5), pp. 742 -752. <Go to ISI>: //WOS: 000337565700003 AND http: //www. askforce. org/web/CBD/Hartung-Schiemann-Precise-Plant-Breeding-regulation-EU-2014. pdf

Based on the history of GMO legislation in the EU, it is not expected that such a paradigm shift will occur short-term or even medium-term. Therefore, in the near future, we urgently require more pragmatic handling of the NPBTs, such that modified plants that do not contain recombinant DNA are exempt from regulation (e. g. inclusion in Annex IB of Directive 2001/18/EC), and those containing recombinant DNA (which is not a hazard per se) are de-regulated in some way as unintended side-effects are expected to be lower than in first-generation transgenic plants (European Food Safety Authority Panel on Genetically Modified Organisms, 2012 b; Podevin et al. , 2012; Pauwels et al. , 2013).

Conclusions for a new regulatory system It should be science based and build on a truly interdisciplinary discourse including various kinds of knowledge It should be product-oriented with a detailed view of the technologies used, from Conventional over transgenic to gene edited crops, pre-evaluated according to the Canadian system with some adaption: Important, the regulation should work as a dynamically scalable system, adapted to the technology used, embracing the whole continuum in breeding technologies, from conventional breeding including artificial mutation, or any other kind of forced Mutation, and including also special cases of conventional breeding. Transgenic and widely distributed crops with many years of safe use should be Exempted from regulation. Gene editing as a new method should be tested along the genomic impact depth of the method, but not explicitly excluded. See for a suggestion of dynamically scalable regulation:
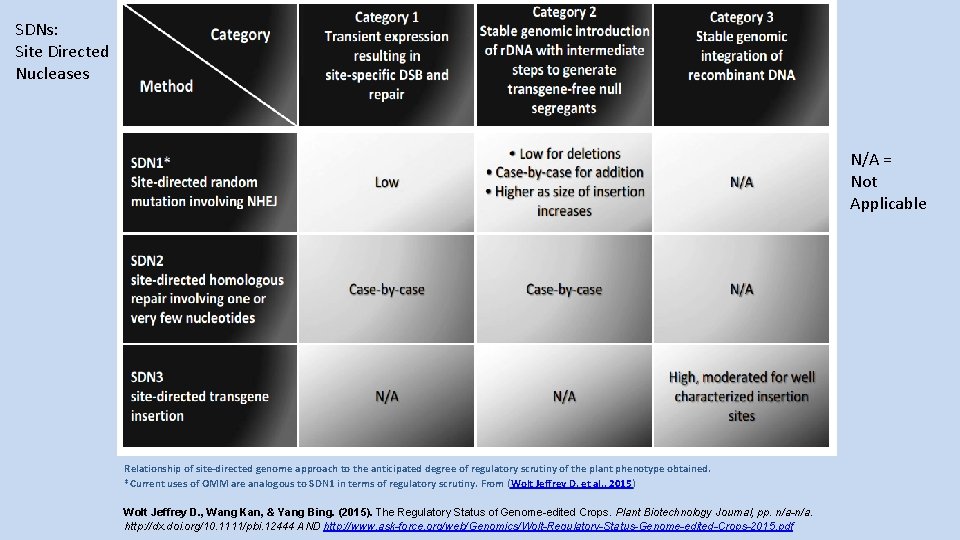
SDNs: Site Directed Nucleases N/A = Not Applicable Relationship of site-directed genome approach to the anticipated degree of regulatory scrutiny of the plant phenotype obtained. *Current uses of OMM are analogous to SDN 1 in terms of regulatory scrutiny. From (Wolt Jeffrey D. et al. , 2015) Wolt Jeffrey D. , Wang Kan, & Yang Bing. (2015). The Regulatory Status of Genome-edited Crops. Plant Biotechnology Journal, pp. n/a-n/a. http: //dx. doi. org/10. 1111/pbi. 12444 AND http: //www. ask-force. org/web/Genomics/Wolt-Regulatory-Status-Genome-edited-Crops-2015. pdf

It is an unacceptable paradox: We cannot exclude oligo-mutated, finally DNA-free NBTs (ODMs) from regulation True, they are not mentioned in the present day EU and Cartagena Regulation And they are free of “foreign” DNA, but still: These are new breeding products done with a still little known new methodology This is why we need a dynamically scalable regulation, the same is true for conventional NBTs, pre-selected similar to a Canadian method But we might exclude those ODMs and conventional NBTs after a very few years of successful testing from future regulation

Process or product oriented ? ? The useless fight in regulatory Paragraphs: EU regulation and Cartagena-Protocol politically seen as “process-oriented”, but it is possible to interpret some paragraphs as product-oriented in order to construct an excuse to exclude Gene Editing methods not using DNA BUT: ACTUALLY: EVERY PRODUCT IS MADE BY A PROCESS, EVERY PROCESS ENDS IN A PRODUCT. This is why regulatory evaluation has to begin with the product, but process details need to be included

Remarks from the discussion of the meeting: about Monocultures The speaker made some critical remarks about the monoculture of Soybeans in South America and emphasized at the same time that the topic of Monocultures is combined with lots of misunderstandings and wrong concepts. First a caveat: it is not correct to criticize monocultures in an undifferentiated way for two reasons: 1. All globally grown crops with their un-comparable economic success have been selected by our early farmers from monodominant fields of crops such as rice and wheat etc. Read the excellent account of Wood, D. , & Lenne, J. (2001). Nature's fields: a neglected model for increasing food production. Outlook on Agriculture, 30(3), pp. 161 -170. <Go to ISI>: //000171396200003 AND http: //www. ask-force. org/web/Organic/Wood-Natures-Fields-2001. pdf 2. A basic book about the origins of organic farming: part of it goes back to the Nazi regime De. Gregori, T. R. (2004). The Origins of the Organic Agriculture Debate. pp. 232: Iowa State University Press, Blackwell. 0 -8138 -0513 -9 http: //www. amazon. co. uk/gp/product/0813805139/202 -6942345 -6387021? v=glance&n=266239 AND Chapter 7 on Nazis http: //www. ask-force. org/web/Organic/de. Gregori-Organic-Ag-Chapter-7 -Nazis-2004. pdf 3. Lies about double cancer rate in soybean fields in Argentina: Dr. Angelika Hilbeck, a well-known Swiss opponent of GMO-agriculture: Recently, a report of the Argentine Government revealed a double cancer rate in soybean fields of Cordoba. This statement was published in a widely distributed brochure about Nature Conservation. The contrary is the case: the official report confirms no correlation between soybean monocultures and cancer rates: Levitus Gabriela. (2014). Agrochemicals and deaths from cancer in Cordoba Province, Argentina, a rebuttal, draft. http: //www. ask-force. org/web/Herbizide. Tol/Levitus-Agrochemicals-and-cancer-deaths-Cordoba-Argentina-Rebuttal-2014. pdf 4. Scientifically proven loss of biodiversity in region-wide monocultures: many examples like Candolfi (see samples of non-target Insects in conventional maize monocultures). One of the many possible solutions: mixed cropping: Lithourgidis, A. , Dordas, C. , Damalas, C. , & Vlachostergios, D. (2001). Annual intercrops: an alternative pathway for sustainable agriculture. Australian journal of crop science, 5(4), pp. 396. http: //www. ask-force. org/web/Organotransgenic/Lithourgidis-Annual-intercrops-alternative-pathway-sustainable-ag-2001. pdf
- Slides: 54