Processing integrating and analysing chromatin immunoprecipitation followed by

Processing, integrating and analysing chromatin immunoprecipitation followed by sequencing (Ch. IP-seq) data Bioinformatics methods, models and applications to disease Alex Essebier

Ch. IP-seq experiment • • • To determine protein binding sites in the genome Snapshot of in vivo sites occupied by protein Improve understanding of regulation in genome Improve understanding of epigenetics Transcription factors – TFs Histone modifications – HMs – To tails of histone proteins forming nucleosomes
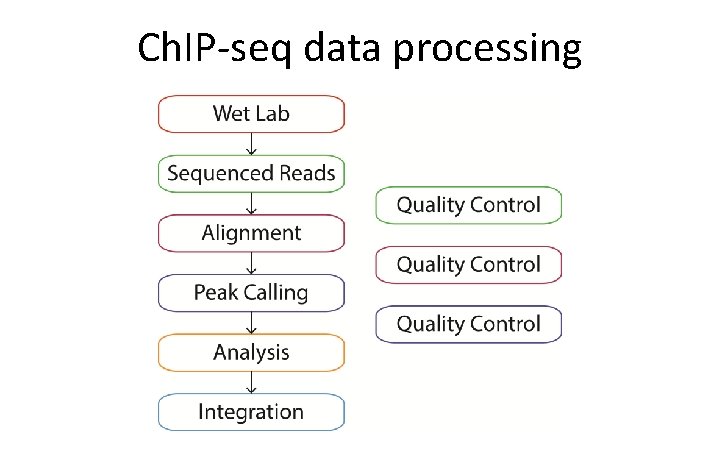
Ch. IP-seq data processing
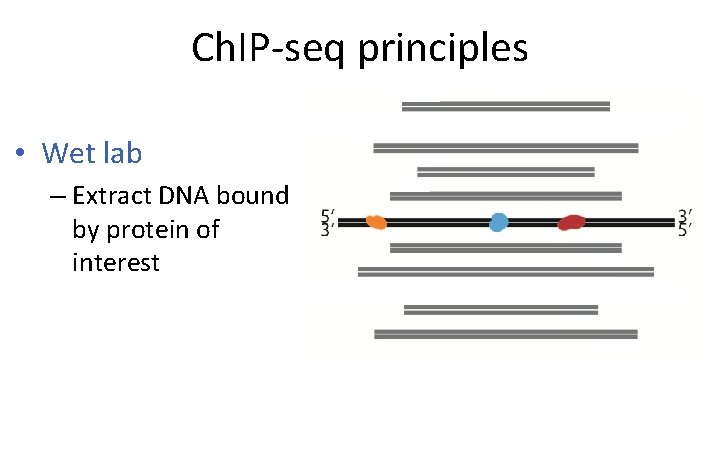
Ch. IP-seq principles • Wet lab – Extract DNA bound by protein of interest
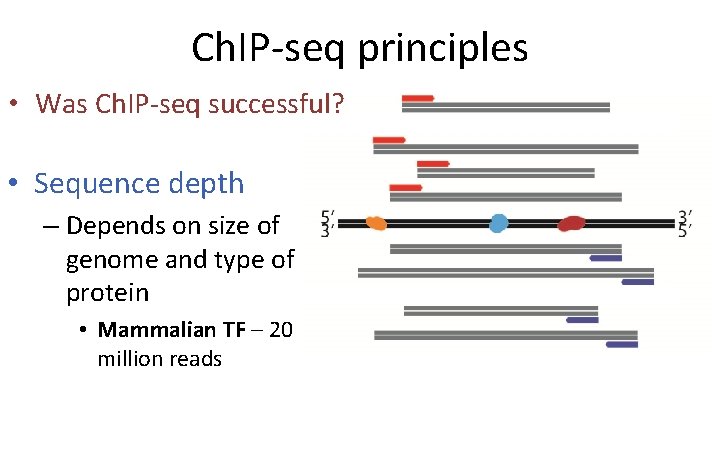
Ch. IP-seq principles • Was Ch. IP-seq successful? • Sequence depth – Depends on size of genome and type of protein • Mammalian TF – 20 million reads
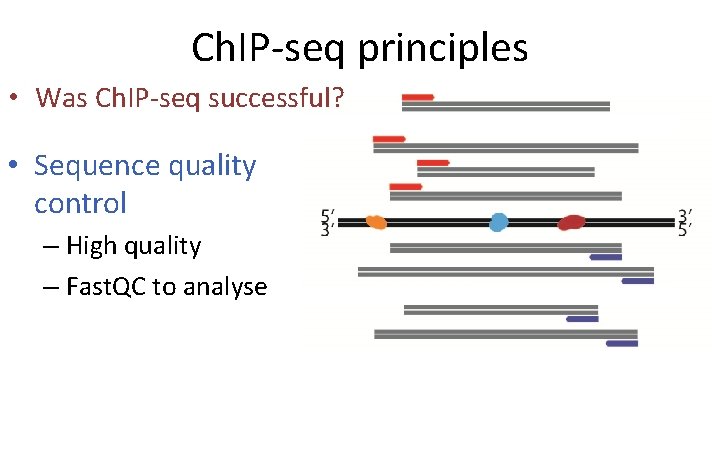
Ch. IP-seq principles • Was Ch. IP-seq successful? • Sequence quality control – High quality – Fast. QC to analyse
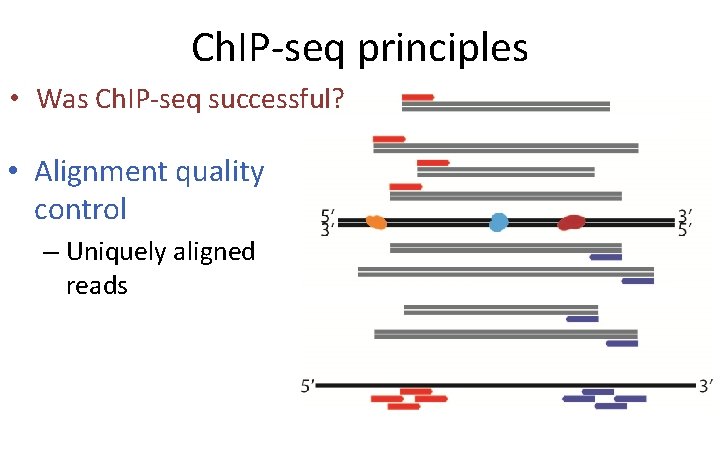
Ch. IP-seq principles • Was Ch. IP-seq successful? • Alignment quality control – Uniquely aligned reads
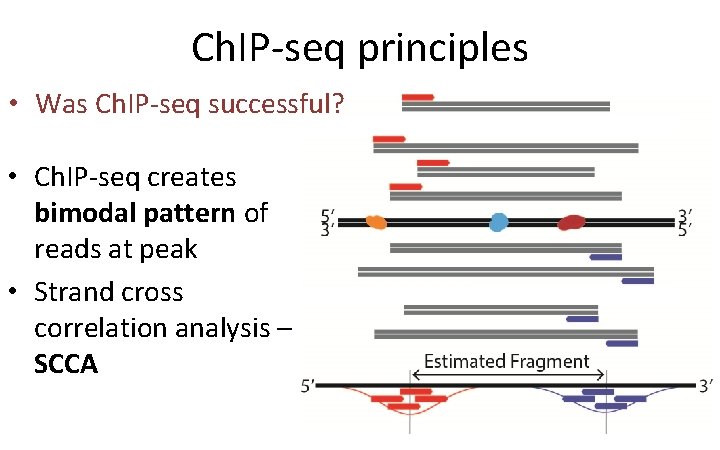
Ch. IP-seq principles • Was Ch. IP-seq successful? • Ch. IP-seq creates bimodal pattern of reads at peak • Strand cross correlation analysis – SCCA
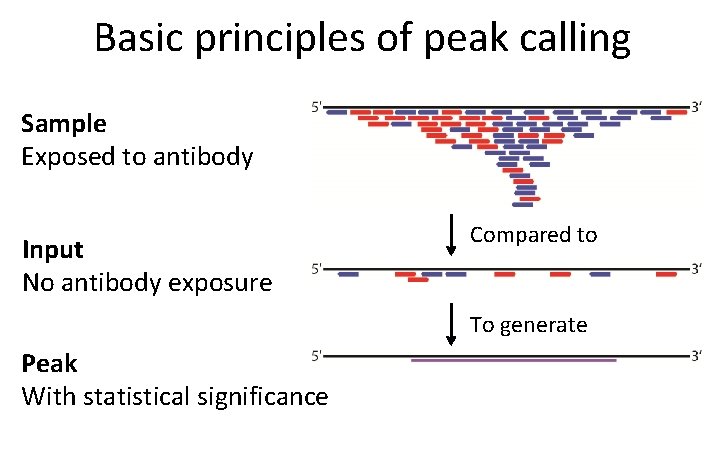
Basic principles of peak calling Sample Exposed to antibody Input No antibody exposure Compared to To generate Peak With statistical significance
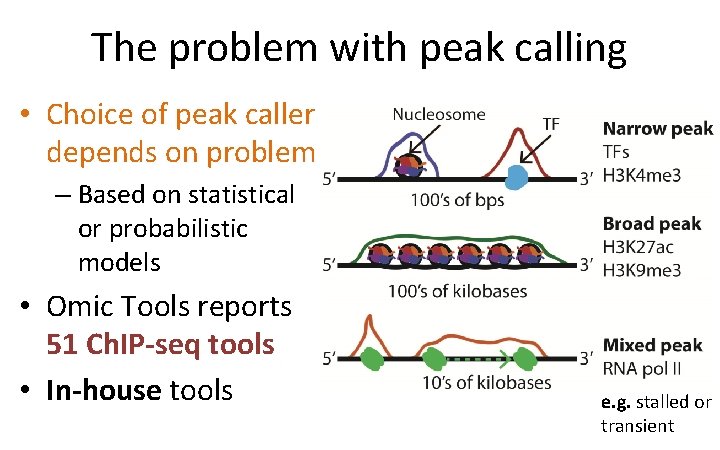
The problem with peak calling • Choice of peak caller depends on problem – Based on statistical or probabilistic models • Omic Tools reports 51 Ch. IP-seq tools • In-house tools e. g. stalled or transient
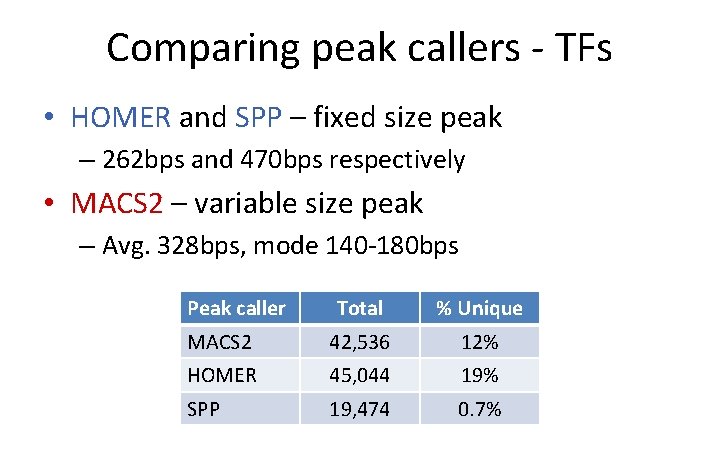
Comparing peak callers - TFs • HOMER and SPP – fixed size peak – 262 bps and 470 bps respectively • MACS 2 – variable size peak – Avg. 328 bps, mode 140 -180 bps Peak caller Total % Unique MACS 2 42, 536 12% HOMER 45, 044 19% SPP 19, 474 0. 7%

• Number Peak quality control – How active is the protein? • Read coverage – Are peak locations enriched for reads? – Fraction of reads in peaks (FRi. P) > 1% • Generally observe > 10% • E. g. below 6/50 reads in peak -> 12%
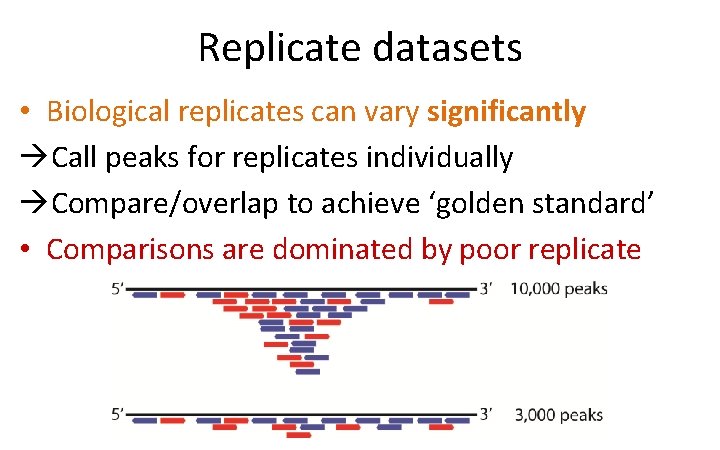
Replicate datasets • Biological replicates can vary significantly Call peaks for replicates individually Compare/overlap to achieve ‘golden standard’ • Comparisons are dominated by poor replicate

PEAK ANALYSIS Exploring the peaks generated from Ch. IP-seq

Transcription Factors • Confirm in vitro and in silico results – Overlapping peaks with motifs • Identify consensus motif – For TFs which do not have an existing/known motif – To identify variations in motif • Differential peak binding – To identify differences in binding patterns – Compare cell types or time points

Histone Modifications • Epigenetic analysis – Generate epigenetic profiles – Identify chromatin states genome wide • E. g. Chrom. HMM – Identify regulatory modules • E. g. promoters or enhancers • Differential peak binding – Identify differences in epigenetic patterns

INTEGRATING DATA Combining data sets to improve outcomes

Data integration • Experiments capture dependent regulatory events – Ch. IP-seq – regulatory elements – DNase I hypersensitivity (DHS) – chromatin accessibility – RNA-seq – expression patterns • Consider multiple datasets to: – Improve confidence and understanding – Support hypotheses
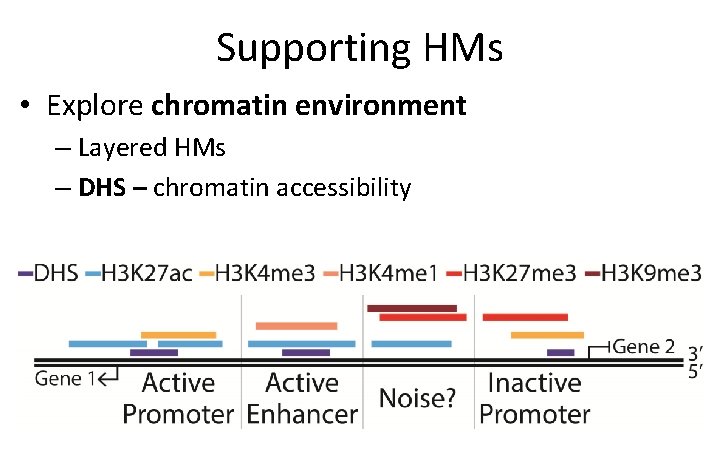
Supporting HMs • Explore chromatin environment – Layered HMs – DHS – chromatin accessibility

Ch. IP-seq complications • Possible to observe multiple states at one location • False negatives – Can’t detect small sub-populations • False positives – General non-specific chromatin being pulled down – Bias not removed by input
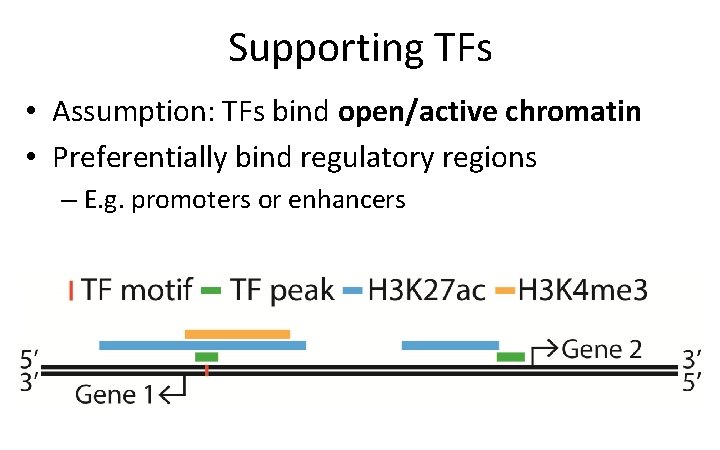
Supporting TFs • Assumption: TFs bind open/active chromatin • Preferentially bind regulatory regions – E. g. promoters or enhancers
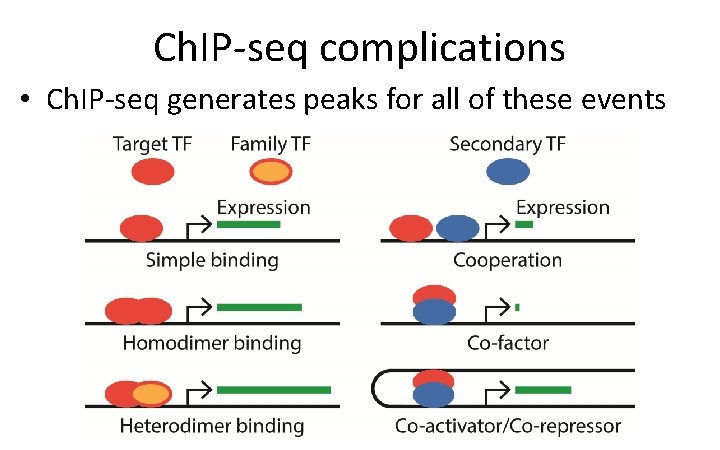
Ch. IP-seq complications • Ch. IP-seq generates peaks for all of these events
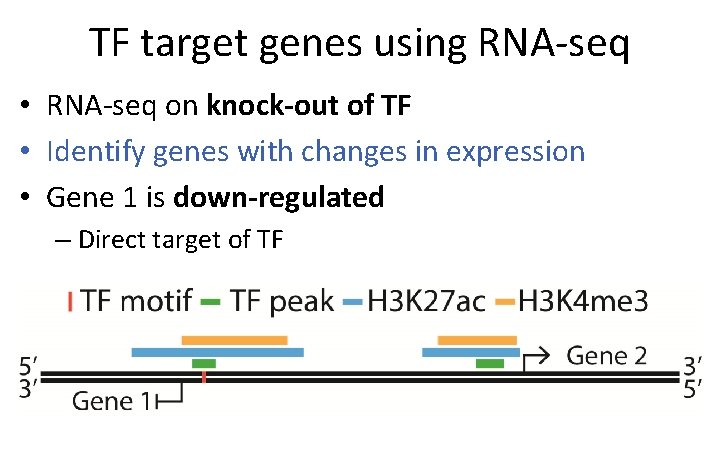
TF target genes using RNA-seq • RNA-seq on knock-out of TF • Identify genes with changes in expression • Gene 1 is down-regulated – Direct target of TF

PRACTICAL EXAMPLE The role of Math 1 in differentiation of cerebellum
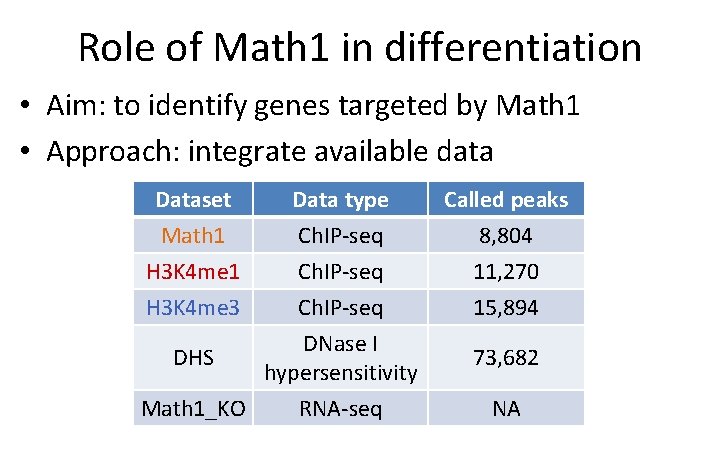
Role of Math 1 in differentiation • Aim: to identify genes targeted by Math 1 • Approach: integrate available data Dataset Math 1 H 3 K 4 me 3 Data type Ch. IP-seq DNase I DHS hypersensitivity Math 1_KO RNA-seq Called peaks 8, 804 11, 270 15, 894 73, 682 NA

Combining replicates • Two replicates for H 3 K 4 me 1 • Two peak callers: – MACS 2 – HOMER Data set MACS 2_rep 1 MACS 2_rep 2 HOMER_rep 1 HOMER_rep 2 Peaks Overlap 8, 183 5, 269 9, 789 71, 534 48, 661 70, 469 H 3 K 4 me 1 rep 1 H 3 K 4 me 1 rep 2 Ig. G Control MACS 2_rep 1 HOMER_rep 1 MACS 2_rep 2 HOMER_rep 2

Combining replicates • Two peak callers: – MACS 2 – HOMER • Generate high quality merged output – Requires called peak in 3 of 4 data sets – 11, 270 peaks in total H 3 K 4 me 1 rep 1 H 3 K 4 me 1 rep 2 Ig. G Control MACS 2_rep 1 HOMER_rep 1 MACS 2_rep 2 HOMER_rep 2 Merged_out

Identify regulatory regions • Three outputs from epigenetic data: – H 3 K 4 me 1_DHS sites putative enhancers – H 3 K 4 me 3_DHS sites putative promoters – H 3 K 4 me 1_H 3 K 4 me 3_DHS sites other Comparison H 3 K 4 me 1_DHS H 3 K 4 me 3_DHS H 3 K 4 me 1_H 3 K 4 me 3_DHS Sites Overlap 9, 011 80% of H 3 K 4 me 1 15, 098 95% of H 3 K 4 me 3 919

Bound Math 1 • Identify regulatory regions bound by Math 1 • Math 1 binds preferentially to putative enhancer • >50% Math 1 binding sites do not overlap a defined regulatory region Putative Enhancer Putative Promoter No Overlap
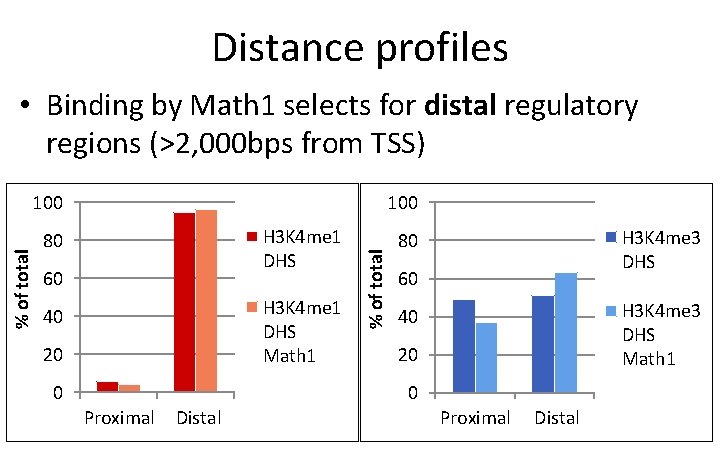
Distance profiles • Binding by Math 1 selects for distal regulatory regions (>2, 000 bps from TSS) 100 H 3 K 4 me 1 DHS 80 60 H 3 K 4 me 1 DHS Math 1 40 20 0 Proximal Distal % of total 100 H 3 K 4 me 3 DHS 80 60 H 3 K 4 me 3 DHS Math 1 40 20 0 Proximal Distal
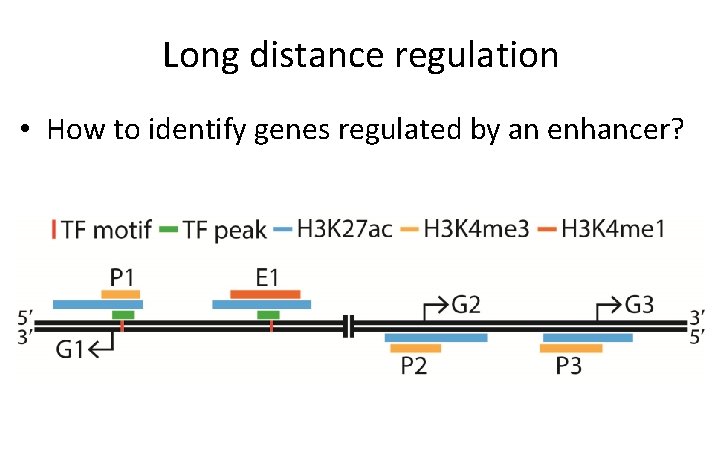
Long distance regulation • How to identify genes regulated by an enhancer?
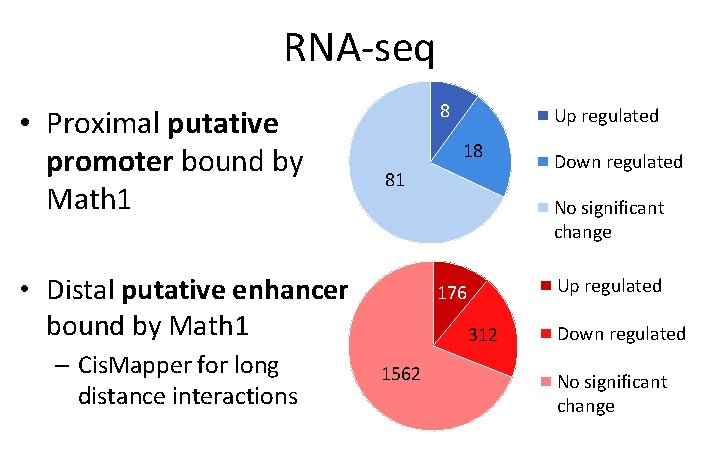
RNA-seq • Proximal putative promoter bound by Math 1 8 18 81 Down regulated No significant change • Distal putative enhancer bound by Math 1 – Cis. Mapper for long distance interactions Up regulated 176 312 1562 Down regulated No significant change
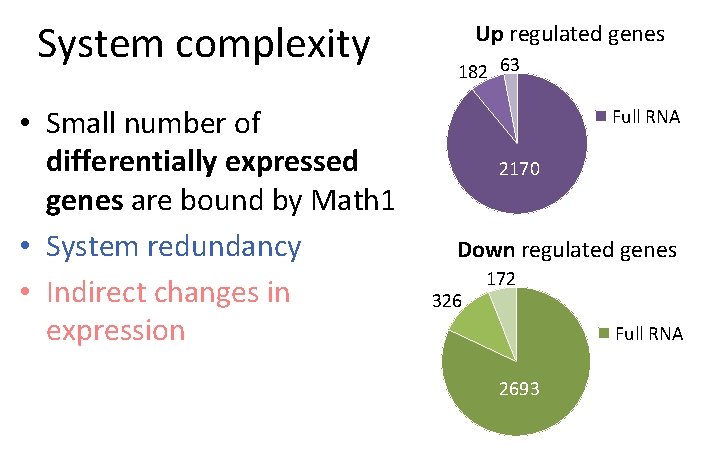
System complexity • Small number of differentially expressed genes are bound by Math 1 • System redundancy • Indirect changes in expression Up regulated genes 182 63 Full RNA 2170 Down regulated genes 326 172 Full RNA 2693

Take home messages • Understand your data and how best to use it • Quality control • Peak calling – Use multiple where possible – Keep up to date with advances • Data integration – Use all available data to gain a more complete picture

Resources • Data – – Klisch, T. J. , Xi, Y. , Flora, A. , Wang, L. , Li, W. , & Zoghbi, H. Y. (2011). In vivo Atoh 1 targetome reveals how a proneural transcription factor regulates cerebellar development. Proceedings of the National Academy of Sciences, 108(8), 3288 -3293. Frank, C. L. , Liu, F. , Wijayatunge, R. , Song, L. , Biegler, M. T. , Yang, M. G. , . . . & West, A. E. (2015). Regulation of chromatin accessibility and Zic binding at enhancers in the developing cerebellum. Nature neuroscience, 18(5), 647 -656. • Useful papers – – – Bailey, T. , Krajewski, P. , Ladunga, I. , Lefebvre, C. , Li, Q. , Liu, T. , . . . & Zhang, J. (2013). Practical guidelines for the comprehensive analysis of Ch. IP-seq data. PLo. S Comput Biol, 9(11), e 1003326. Landt, S. G. , Marinov, G. K. , Kundaje, A. , Kheradpour, P. , Pauli, F. , Batzoglou, S. , . . . & Chen, Y. (2012). Ch. IP-seq guidelines and practices of the ENCODE and mod. ENCODE consortia. Genome research, 22(9), 1813 -1831. Farnham, P. J. (2009). Insights from genomic profiling of transcription factors. Nature Reviews Genetics, 10(9), 605 -616. Zhang, Y. , Liu, T. , Meyer, C. A. , Eeckhoute, J. , Johnson, D. S. , Bernstein, B. E. , . . . & Liu, X. S. (2008). Model-based analysis of Ch. IPSeq (MACS). Genome biology, 9(9), 1. Heinz, S. , Benner, C. , Spann, N. , Bertolino, E. , Lin, Y. C. , Laslo, P. , . . . & Glass, C. K. (2010). Simple combinations of lineagedetermining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Molecular cell, 38(4), 576 -589.
- Slides: 35