Problems of Subject Mediator Development for Gene Expression

Problems of Subject Mediator Development for Gene Expression Regulation Domain 1 L. A. Kalinichenko, 1 D. O. Briukhov, 1 V. N. Zakharov, 2 O. A. Podkolodnaya, 2, 3 N. L. Podkolodny 1 Institute for Problems of Informatics RAS, Moscow, Russia 2 Institute of Cytology and Genetics SB RAS, Novosibirsk, Russia 3 Institute of Computational Mathematics and Mathematical Geophysics SB RAS, Novosibirsk, Russia

The Mediator Concept • The mediator architecture (Wiederhold, 1992) deals with the problem of integration of heterogeneous information. The sources are "heterogeneous" on many levels: • • • data model and types of data used; the underlying data units; behavior of objects involved; the underlying concepts; the schema that the information may conform cannot be rigid in advance. • Mediator is to provide a uniform query interface to the multiple data sources, thereby freeing the user from having to locate the relevant sources, query each one in isolation, and combine manually the information from the different sources.

Mediation Approaches • integration information from pre-selected sources according to the predefined information needs. A procedural approach is known (TSIMMIS, Squirrel, WHIPS) to integrate information from sources through ad-hoc procedures. When information needs or sources change, a new mediator should be generated. This is known as Global as View (GAV) approach. • integration information from arbitrary sources according to the predefined information needs. A declarative approach is known (Carnot, SIMS, Information Manifold, Infomaster). Mediators contain mechanisms to rewrite queries according to source descriptions. A rewritten query should be contained in the original query. This is known as Local as View (LAV) approach.

Mediator Layers • Federated layer keeps subject mediator specifications, such as ontological definitions of the subject domain, schema description defining structural (types, classes, attributes) and functional (e. g. , facilities for semantic data analysis and predictions, knowledge discovery based on the automatic methods) capabilities of the mediator; • Local layer represents canonical specifications of the heterogeneous sources registered at the mediator; • Intermediate layer defines a mapping of the source specifications into the specifications of the mediator.

Advantages of the Proposed Approach • Semantic integration of heterogeneous information collections can be reached by taking into account structural, value, semantic, quality data heterogeneity; • Users should know only subject definitions that contain concepts, structures and methods as defined by the community; • Querying the subject definitions, users have integrated access to all information registered at the mediators up to the moment of a query; • Personalization providing convenient views for specific groups of users can be formed above the subject definitions. This process is independent of the existing collection and their registration.

The Mediator for Gene Expression Regulation The mediator is oriented on a broad class of problems. The intuition behind them can be provided by an example sequence of interrelated queries to the mediator that are intended for preparation of the training samples of regulatory regions, which may be used by recognition programs: • • to output the set of transcription factor binding sites sequences, which have a definite type of DNA-binding domain, search for transcription factors corresponding to the proteins found, search for transcription factor binding sites; search for the sequences of pre-ordered length including relevant transcription factor binding sites.

Examples of the ontological definitions Name "protein" Definition "A large molecule composed of one or more chains of amino acids in a specific order; the order is determined by the base sequence of nucleotides in the gene coding for the protein. Proteins are required for the structure, function, and regulation of the bodys cells, tissues, and organs, and each protein has unique functions. Examples are hormones, enzymes, and antibodies. “ Name "transcription factor" Definition "A protein that regulates transcription after nuclear translocation by specific binding with DNA or by stoichiometric interaction with a protein that can be assembled into a sequence-specific DNA-protein complex. " Part-of "transcription complex" Subclass-of "protein"
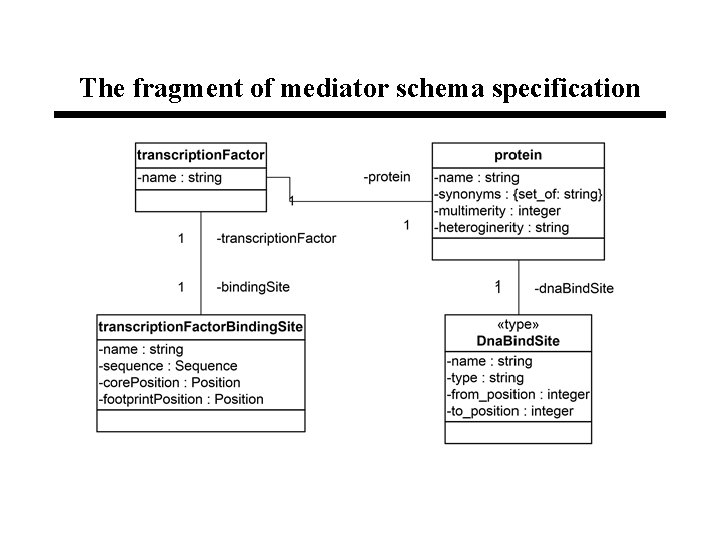
The fragment of mediator schema specification

Information Sources Initial set of information sources to be registered at the mediator includes: • The database TRRD developed at the Institute of Cytology and Genetics, unique informational resource that has neither world-wide analogs and that contains information about structural and functional organization of extended transcription regulating regions of eukaryotic genes and their expression. • The database SWISSPROT contains an information about the structure and functions of proteins, about their domain structure, sequences, etc. • The databases EMBL/Gen. Bank accumulate information about the sequences DNA, RNA, their exon-intron structure, and other functional layout. • The database Medline/Pub. Med stores bibliography that is necessary for supporting and verifying the data presented.
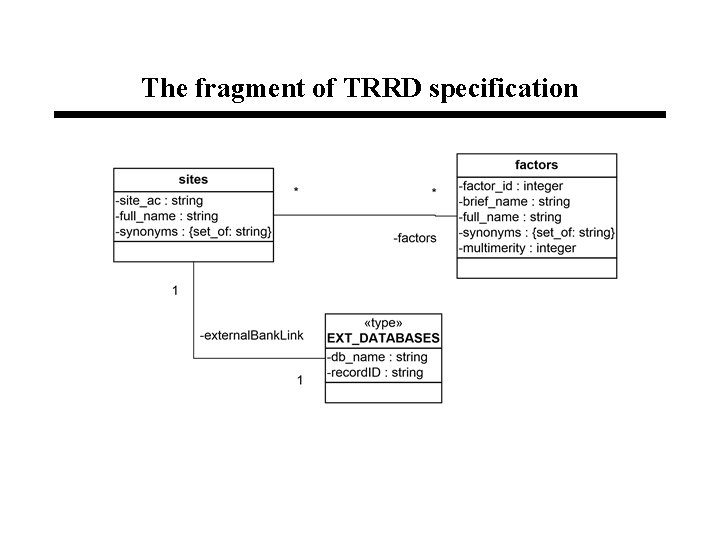
The fragment of TRRD specification
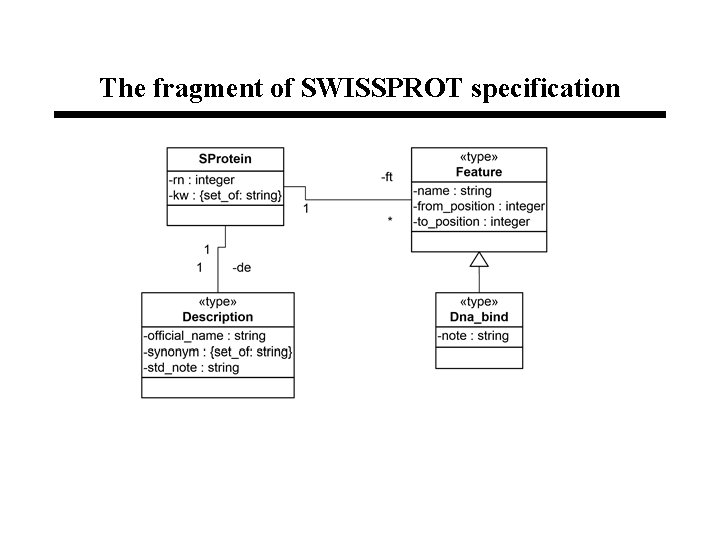
The fragment of SWISSPROT specification

Process of an Information Source Registration For each source class the following steps are required: 1. relevant federated classes identification • 2. Find federated classes that ontologically can be used for defining source class extent in terms of federated classes. To a source class several federated classes may correspond covering with their instance types different reducts of an instance type of the source class. On another hand, several source classes may correspond to one federated class. most common reducts construction For an instance type of each identified federated class do: • Construct most common reducts for instance type of this federated class and source class instance type to concretize (partially) such federated instance type. Most common reduct may include also additional attributes corresponding to those federated type attributes that can be derived from the source type instances to support them. • In this process for each attribute type of the common reduct a concretizing type, concretizing function or their combination should be constructed (this step should be recursively applied).

Process of an Information Source Registration For each source class the following steps are required: 3. partial source view construction • 4. For each relevant federated class construct a partial source view expressing a constraints in terms of the federated class that should be satisfied by values of respective most common reducts of source class instances. Thus partial views over all relevant federated classes will be obtained. partial views composition • Construct compositions of the source type most common reducts obtained for instance types of all federated classes involved. • Construct a source view as a composition of partial views obtained above. This is an expression of a materialized view of an information source in terms of federated classes. An instance type of this view is determined by the most common reducts composition constructed above.

Most Common Reduct Between Mediator Type Protein and SWISSPROT Type SProtein {R_Protein_SProtein; in: reduct; metaslot of: Protein; taking: {name, synonyms, keywords, dna. Bind. Site}; c_reduct: CR_Protein_SProtein end }

Most Common Reduct Between Mediator Type Protein and SWISSPROT Type SProtein {CR_Protein_SProtein; in: c_reduct; . . . simulating: { R_Protein. name ~ get_name, R_Protein. synonyms ~ get_synonyms, R_Protein. key. Words ~ R_Protein. kw, R_Protein. dna. Bind. Site ~ get_dna. Bind. Site} get_name: {in: function; params: {+ext/CR_Protein_SProtein, -returns/string}; predicative: {ex p/SProtein ((p/CR_Protein_SProtein = ext) & returns = p. de. official_name)}}. . . get_dna. Bind. Site: {in: function; params: {+ext/CR_Protein_SProtein, -returns/DNABind. Site}; predicative: {ex p/SProtein ((p/CR_Protein_SProtein = ext) & ex d/Dna_bind (in(p. ft, d) & returns = d/CR_Dna. Bind. Site_Dna_bind))}} }

Partial Source View Construction (Example) The formula expressing the SWISSPROT class sprotein is terms of the mediator class protein is defined as: sprotein(p/CR_Protein_SProtein) protein(p/R_Protein_SProtein) Specification of a class (actually, this is local as view class) containing this formula is: {v_sprotein_protein; in: class; class_section: { lav: invariant, {subseteq (v_sprotein_protein(p), protein(p/R_Protein_SProtein))} }; instance_section: CR_Protein_SProtein }

Example of formulas expressing the source classes is terms of the mediator classes sprotein(p/CR_Protein_SProtein) protein(p/R_Protein_SProtein) factors(p/CR_Transcription. Factor_FACTORS) transcription. Factor(p/R_Transcription. Factor_FACTORS) sites(p/CR_Transcription. Factor. Binding. Site_SITES) transcription. Factor. Binding. Site (p/R_Transcription. Factor. Binding. Site_SITES)

Example of inverse rules protein(p/Protein_SProtein) : - protein(p/Protein_SProtein) transcription. Factor(t/Transcription. Factor_FACTORS) : FACTORS(t/Transcription. Factor_FACTORS) transcription. Factor. Binding. Site(s/Transcription. Factor. Binding. Site_SITES) : SITES(s/Transcription. Factor. Binding. Site_SITES)

Query Rewriting in Terms of the Sources • We consider an example of a query to the mediator: Display the transcription factor binding sites with the definite types of DNA binding domain • In the mediator’s canonical model this query is expressed as: Q: transcription. Factor. Binding. Site(s) & protein(p) & s. transcription. Factor. protein = p & p. dna. Bind. Site. type = “HOMEBOX” • Rewrite query by adding classes that participates in associations (e. g. s. transcription. Factor. protein = p is replaced by transcription. Factor(t) & s. transcription. Factor = t & t. protein = p ): Q’: transcription. Factor. Binding. Site(s) & transcription. Factor(t) & protein(p) & s. transcription. Factor = t & t. protein = p & p. structure. type = “HOMEBOX”

Query Rewriting in Terms of the Sources (cont. ) • After query rewriting applying the inverse rules above, we get the query: RQ 1: FACTORS(t/Transcription. Factor_FACTORS) & SITES(s/Transcription. Factor. Binding. Site_SITES) & sprotein(p/Protein_SProtein) & s. transcription. Factor = t & t. protein = p & p. structure. type = “HOMEBOX” • This query is implemented by a subquery SQ 1 to TRRD and a subquery SQ 2 to SWISSPROT with the remaining postprocessing in the mediator SQ 3: SQ 1(s, t): - FACTORS(t/Transcription. Factor_FACTORS) & SITES(s/Transcription. Factor. Binding. Site_SITES) & s. transcription. Factor = t SQ 2(p): - sprotein(p/Protein_SProtein) & p. structure. type = “HOMEBOX” SQ 3(s, t, p) : - SQ 1(s, t) & SQ 2(p) & t. protein = p

Conclusions • subject mediator for gene expression regulation domain was introduced; • issues of heterogeneous sources registration at the mediator and query rewriting in terms of registered sources was shown; • an approach developed is based on information and software sources in the gene expression regulation domain, which is being developed at the Institute of Cytology and Genetics of SB RAS.
- Slides: 21