Probabilistic Models of Relational Domains Daphne Koller Stanford

Probabilistic Models of Relational Domains Daphne Koller Stanford University © Daphne Koller, 2003

Overview n n n Relational logic Probabilistic relational models Inference in PRMs Learning PRMs Uncertainty about domain structure Summary © Daphne Koller, 2003

Attribute-Based Models & Relational Models © Daphne Koller, 2003
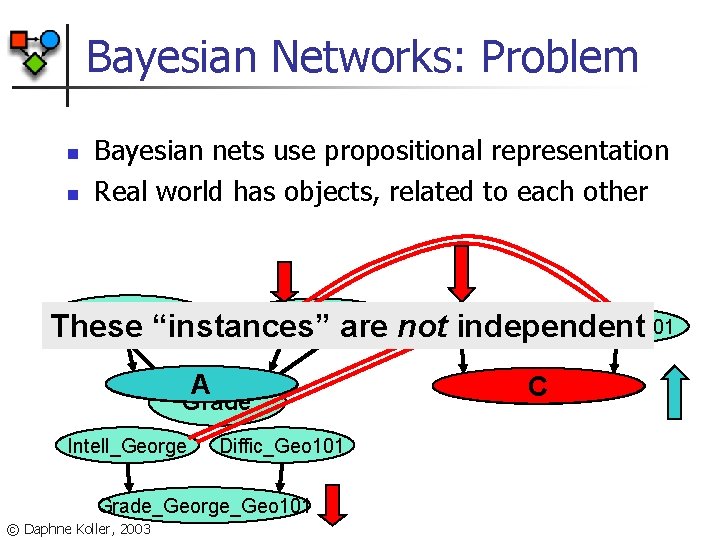
Bayesian Networks: Problem n n Bayesian nets use propositional representation Real world has objects, related to each other Intelligence Diffic_CS 101 Difficulty Intell_Jane Diffic_CS 101 Intell_George These “instances” are not independent Grade_Jane_CS 101 A Grade Intell_George Diffic_Geo 101 Grade_George_Geo 101 © Daphne Koller, 2003 Grade_George_CS 101 C
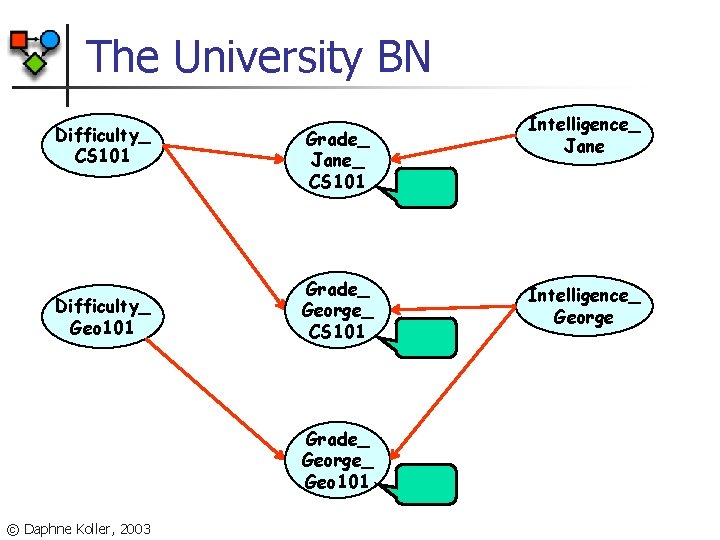
The University BN Difficulty_ CS 101 Difficulty_ Geo 101 Grade_ Jane_ CS 101 Grade_ George_ Geo 101 © Daphne Koller, 2003 Intelligence_ Jane Intelligence_ George
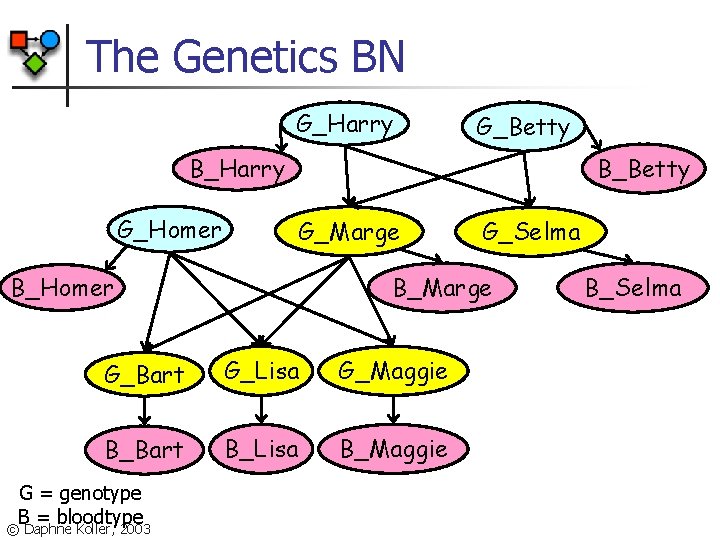
The Genetics BN G_Harry G_Betty B_Harry G_Homer B_Betty G_Marge B_Homer B_Marge G_Bart G_Lisa G_Maggie B_Bart B_Lisa B_Maggie G = genotype B = bloodtype © Daphne Koller, 2003 G_Selma B_Selma

Simple Approach n n n Graphical model with shared parameters … and shared local dependency structure Want to encode this constraint: n n n For human knowledge engineer For network learning algorithm How do we specify n n which nodes share params? shared (but different) structure across nodes? © Daphne Koller, 2003

Simple Approach II n We can write a special-purpose program for each domain: n n n genetic inheritance (family tree imposes constraints) university (course registrations impose constraints) Is there something more general? © Daphne Koller, 2003

Attribute-Based Worlds Smart_Jane & easy_CS 101 Get. A_Jane_CS 101 Smart_Mike & easy_Geo 101 Get. A_Mike_Geo 101 Smart_Jane & easy_Geo 101 Get. A_Jane_Geo 101 Smart_Rick & easy_CS 221 Get. A_Rick_C n World = assignment of values to attributes / truth values to propositional symbols © Daphne Koller, 2003
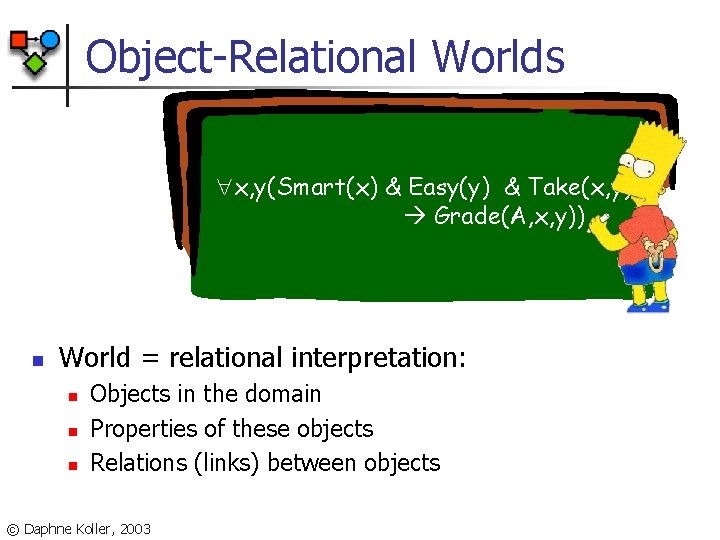
Object-Relational Worlds x, y(Smart(x) & Easy(y) & Take(x, y) Grade(A, x, y)) n World = relational interpretation: n n n Objects in the domain Properties of these objects Relations (links) between objects © Daphne Koller, 2003

Relational Logic n General framework for representing: n n n objects & their properties classes of objects with same model relations between objects Represent a model at the template level, and apply it to an infinite set of domains Given finite domain, each instantiation of the model is propositional, but the template is not © Daphne Koller, 2003
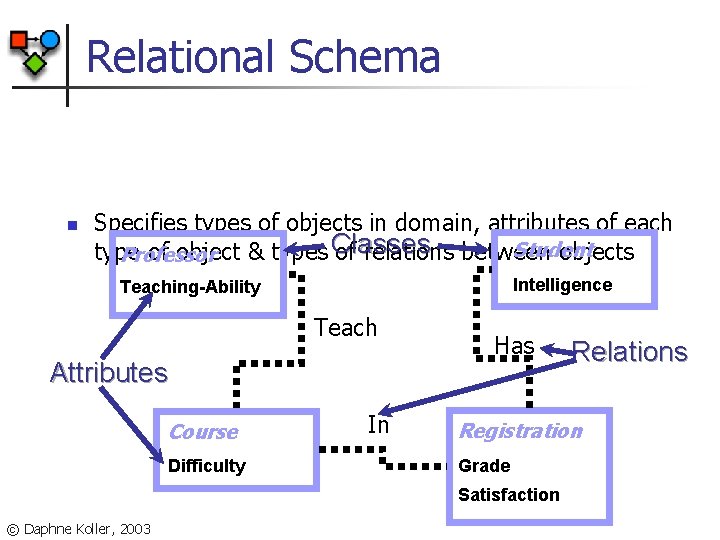
Relational Schema n Specifies types of objects in domain, attributes of each Student type of object & types Classes of relations between objects Professor Intelligence Teaching-Ability Teach Attributes Course Difficulty In Has Registration Grade Satisfaction © Daphne Koller, 2003 Relations
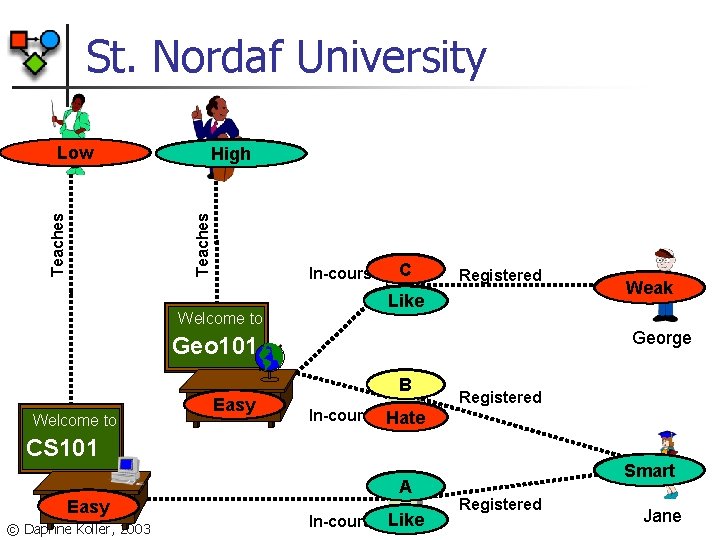
St. Nordaf University Prof. Smith Teaching-ability High Teaches Prof. Jones High Low Teaching-ability Welcome to B C In-course. Grade Registered Satisfac Hate Like George Geo 101 Welcome to Difficulty Easy Grade B C Registered In-course. Satisfac Hate CS 101 Difficulty Easy Hard © Daphne Koller, 2003 Intelligence Weak Grade A Hate Like In-course. Satisfac Intelligence Smart Registered Jane

Relational Logic: Summary n Vocabulary: n Classes of objects: n n Individual objects in a class: n n George. Intelligence, Reg 1. Grade Relationships between these objects n n George, Jane, … Attributes of these objects: n n Person, Course, Registration, … Of(Reg 1, George), Teaches(CS 101, Smith) A world specifies: n n n A set of objects, each in a class The values of the attributes of all objects The relations that hold between the objects © Daphne Koller, 2003

Binary Relations n Any relation can be converted into an object: n n R(x 1, x 2, …, xk) new “relation” object y, R 1(x 1, y), R 2(x 2, y), …, Rk(xk, y) E. g. , registrations are “relation objects” Can restrict attention to binary relations R(x, y) © Daphne Koller, 2003

Relations & Links n n Binary relations can also be viewed as links: Specify the set of objects related to x via R R(x, y) y x. R 1, x y. R 2 E. g. , Teaches(p, c) n n p. Courses = {courses c : Teaches(p, c)} c. Instructor = {professors p : Teaches(p, c)} © Daphne Koller, 2003

Probabilistic Relational Models: Relational Bayesian Networks © Daphne Koller, 2003

Probabilistic Models n Uncertainty model: n n n In attribute-based models, world specifies n n space of “possible worlds”; probability distribution over this space. assignment of values to fixed set of random variables In relational models, world specifies n n n Set of domain elements Their properties Relations between them © Daphne Koller, 2003

PRM Scope n Entire set of relational worlds is infinite and too broad Assume circumscribed class of sets of worlds consistent with some type of background knowledge PRM is a template defining P ( ) for any such n Simplest class attribute-based PRMs: n n n = relational skeleton: n finite set of entities E and relations between them = all assignments of values to all attributes of entities in E n PRM template defines distribution over for any such n [K. & Pfeffer ’ 98; Friedman, Getoor, K. Pfeffer ’ 99] © Daphne Koller, 2003
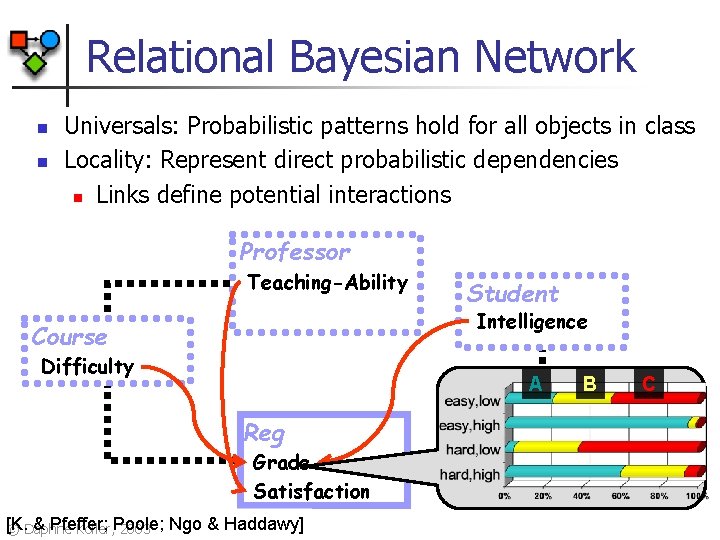
Relational Bayesian Network n n Universals: Probabilistic patterns hold for all objects in class Locality: Represent direct probabilistic dependencies n Links define potential interactions Professor Teaching-Ability Student Intelligence Course Difficulty A Reg Grade Satisfaction [K. & Pfeffer; © Daphne Koller, Poole; 2003 Ngo & Haddawy] B C

RBN: Semantics n n n : set of objects & relations between them : the set of all assignments of values to all attributes of all objects in P ( ) is defined by a ground Bayesian network: n n variables: attributes of all objects dependencies: determined by n n relational links in dependency structure of RBN model © Daphne Koller, 2003
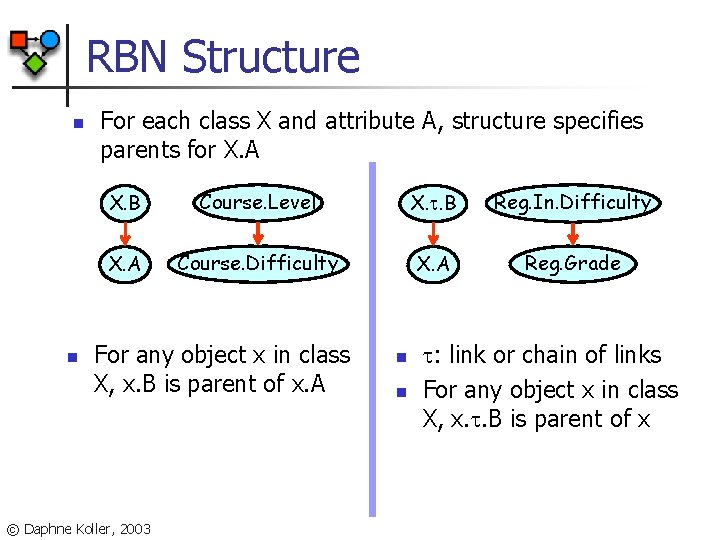
RBN Structure n n For each class X and attribute A, structure specifies parents for X. A X. B Course. Level X. . B Reg. In. Difficulty X. A Course. Difficulty X. A Reg. Grade For any object x in class X, x. B is parent of x. A © Daphne Koller, 2003 n n : link or chain of links For any object x in class X, x. . B is parent of x
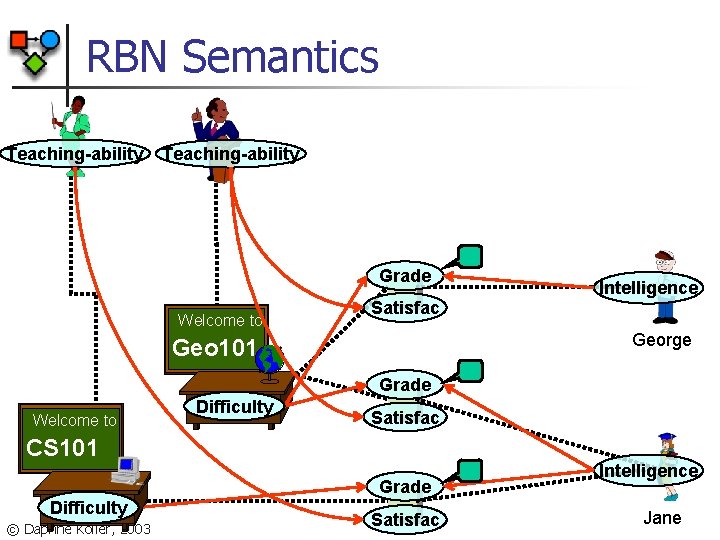
RBN Semantics Prof. Jones Teaching-ability Prof. Smith Teaching-ability Grade Welcome to Satisfac Intelligence George Geo 101 Grade Welcome to to Welcome Difficulty Satisfac CS 101 Grade Difficulty © Daphne Koller, 2003 Satisfac Intelligence Jane
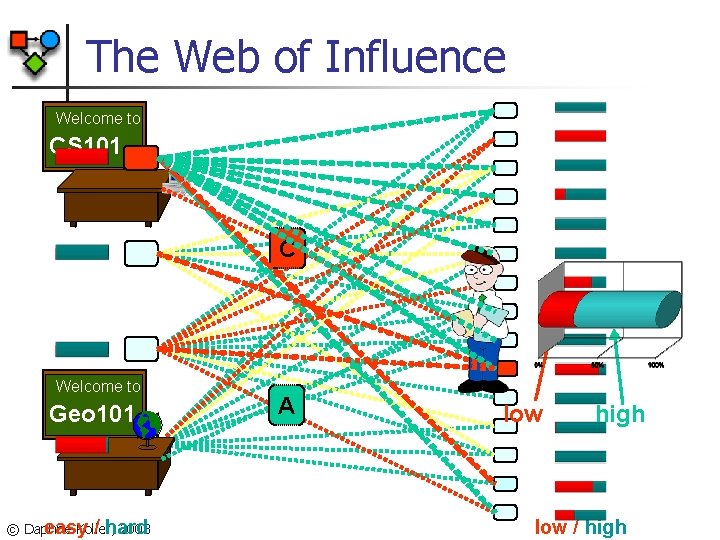
The Web of Influence Welcome to CS 101 C Welcome to Geo 101 © Daphne Koller, 2003 easy / hard A low high low / high
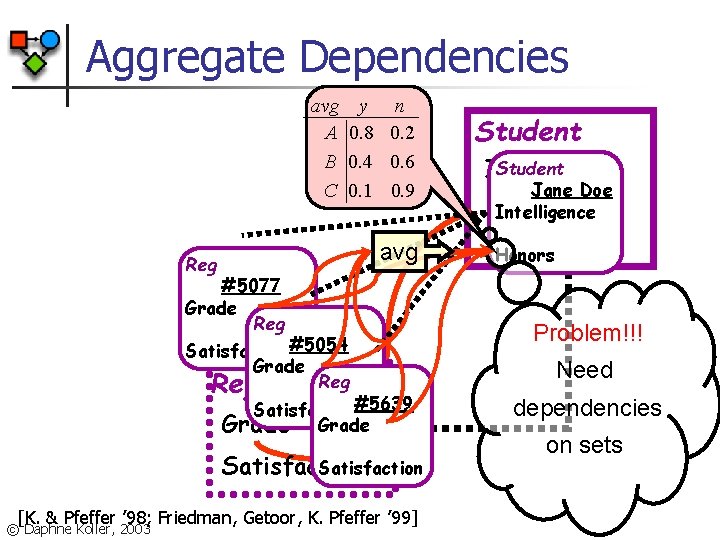
Aggregate Dependencies avg A B C y n 0. 8 0. 2 0. 4 0. 6 0. 1 0. 9 avg Reg #5077 Grade Reg Satisfaction#5054 Grade Reg Satisfaction#5639 Grade Satisfaction [K. & Pfeffer ’ 98; Friedman, Getoor, K. Pfeffer ’ 99] © Daphne Koller, 2003 Student Intelligence Jane Doe Intelligence Honors Problem!!! Need dependencies on sets
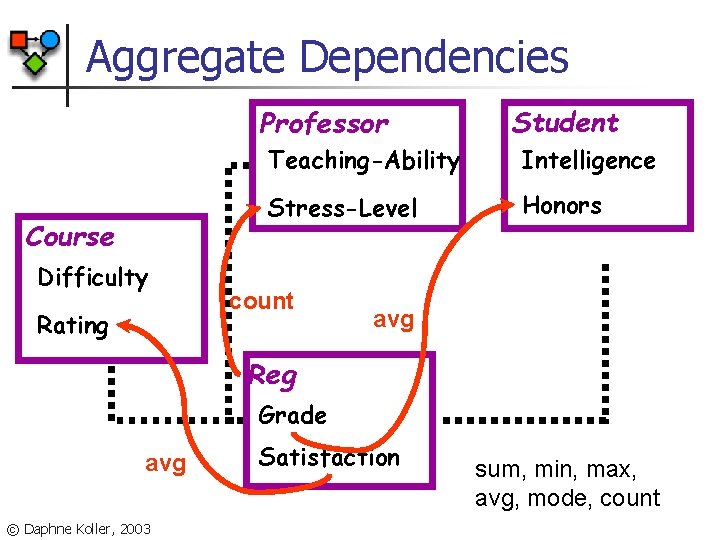
Aggregate Dependencies Professor Course Difficulty Rating Student Teaching-Ability Intelligence Stress-Level Honors count avg Reg Grade avg © Daphne Koller, 2003 Satisfaction sum, min, max, avg, mode, count

Basic RBN: Summary n RBN specifies n A probabilistic dependency structure S: n n n A set of parents X. . B for each class attribute X. A A set of local probability models: n Aggregator to use for each multi-valued dependency n Set of CPD parameters X. A Given relational skeleton structure , RBN induces a probability distribution over worlds n Distribution defined via ground BN over attributes x. A © Daphne Koller, 2003 Attributes Objects

Extension: Class Hierarchy n n Subclasses inherit all attributes of parents, but may have additional ones For inherited attribute X. A, subclass can: n n n inherit parent’s probabilistic model overwrite with local probabilistic model Example: n n n Professor has subclasses assistant, associate, full Inherit distribution over Stress-Level Modify distribution over Salary [K. & Pfeffer ’ 98] © Daphne Koller, 2003

Extension: Class Hierarchies n Hierarchies allow reuse in knowledge engineering and in learning n n Parameters and dependency models shared across more objects If class assignments specified in , class hierarchy does not introduce complications [K. & Pfeffer ’ 98] © Daphne Koller, 2003
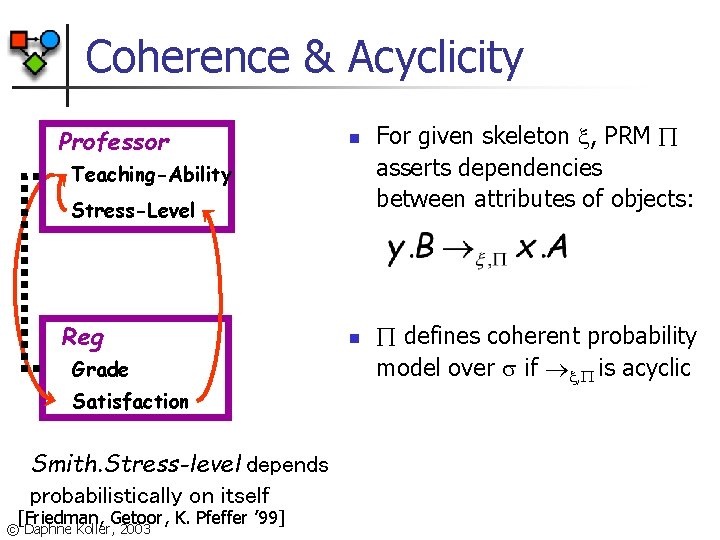
Coherence & Acyclicity Professor n Teaching-Ability Stress-Level Reg Grade Satisfaction Smith. Stress-level depends probabilistically on itself [Friedman, Getoor, K. Pfeffer ’ 99] © Daphne Koller, 2003 n For given skeleton , PRM asserts dependencies between attributes of objects: defines coherent probability model over if , is acyclic

Guaranteeing Acyclicity How do we guarantee that a PRM is acyclic for every object skeleton ? PRM dependency structure S template-level dependency graph Y. B X. A Attribute stratification: if X. . B Parents(X. A), and class of X. is Y class dependency graph acyclic , acyclic for all © Daphne Koller, 2003
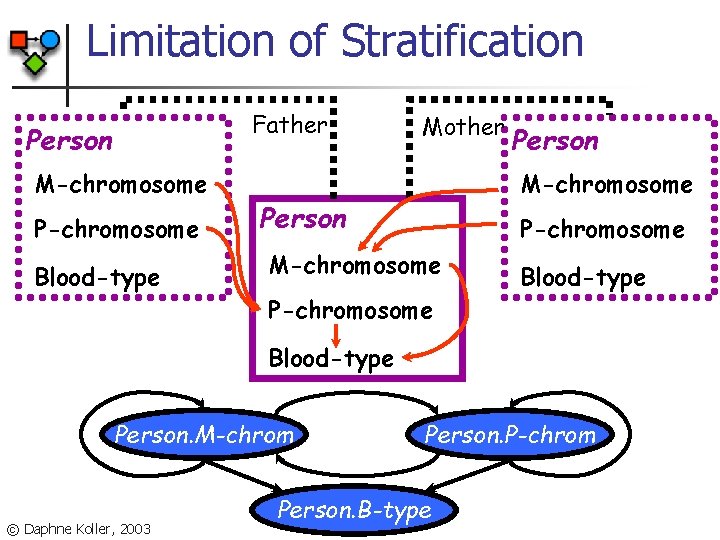
Limitation of Stratification Father Person M-chromosome P-chromosome Blood-type Mother Person M-chromosome Person P-chromosome M-chromosome P-chromosome Blood-type Person. M-chrom © Daphne Koller, 2003 Person. P-chrom Person. B-type
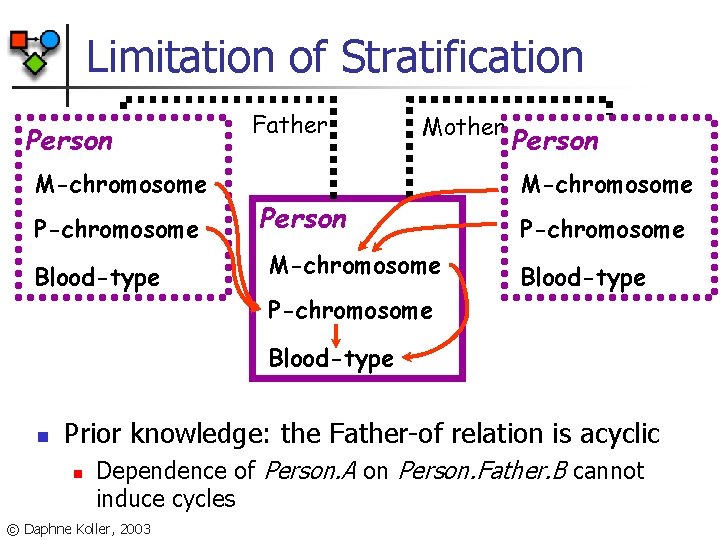
Limitation of Stratification Person M-chromosome P-chromosome Blood-type Father Mother Person M-chromosome P-chromosome Blood-type n Prior knowledge: the Father-of relation is acyclic n Dependence of Person. A on Person. Father. B cannot induce cycles © Daphne Koller, 2003
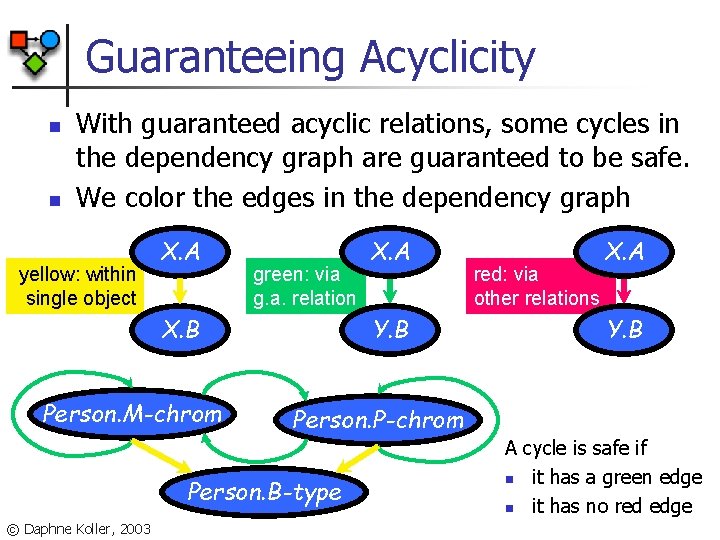
Guaranteeing Acyclicity n n With guaranteed acyclic relations, some cycles in the dependency graph are guaranteed to be safe. We color the edges in the dependency graph yellow: within single object X. A green: via g. a. relation X. B Person. M-chrom Y. B red: via other relations X. A Y. B Person. P-chrom Person. B-type © Daphne Koller, 2003 X. A A cycle is safe if n it has a green edge n it has no red edge

Object-Oriented Bayesian Nets* n OOBNs are RBN with only one type of relation n n One object can be a “part-of” another Objects can only interact with component parts Other types of relationships must be embedded into the “part-of” framework Defines “neat” hierarchy of related objects Provides clearly defined object interface between object x and its enclosing object y * Simplified view © Daphne Koller, 2003 [K. & Pfeffer ’ 97]
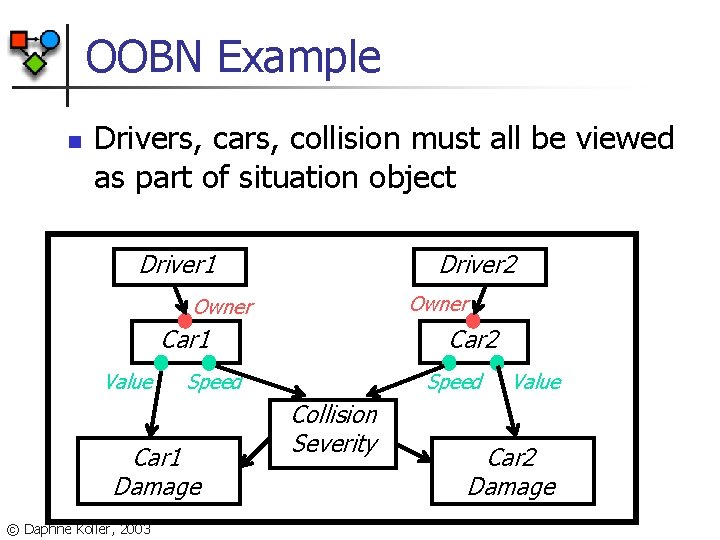
OOBN Example n Drivers, cars, collision must all be viewed as part of situation object Driver 1 Driver 2 Owner Car 1 Value Speed Car 1 Damage © Daphne Koller, 2003 Car 2 Speed Collision Severity Value Car 2 Damage
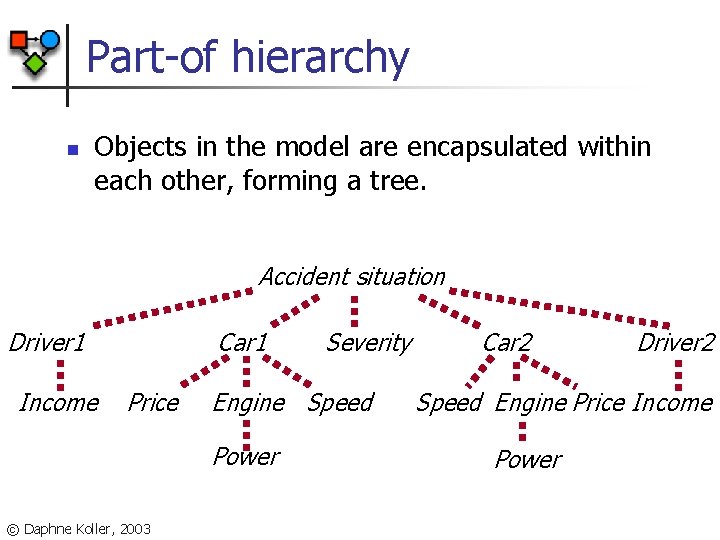
Part-of hierarchy n Objects in the model are encapsulated within each other, forming a tree. Accident situation Driver 1 Income Car 1 Price Engine Speed Power © Daphne Koller, 2003 Severity Car 2 Driver 2 Speed Engine Price Income Power
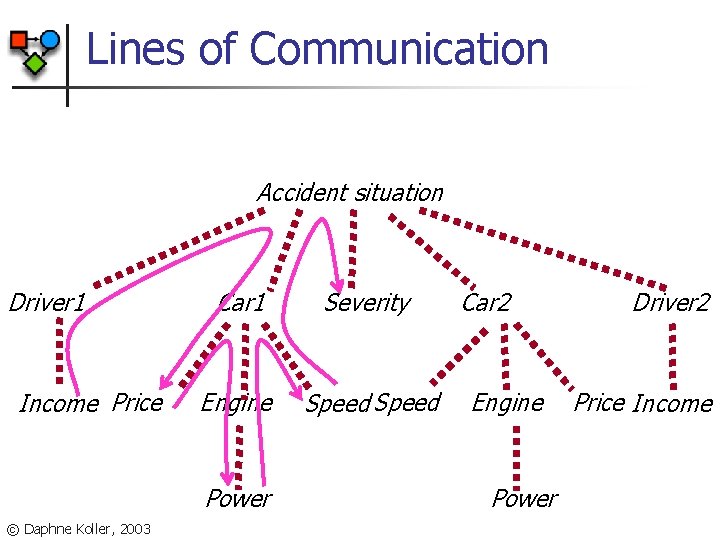
Lines of Communication Accident situation Driver 1 Income Price Car 1 Severity Engine Speed Power © Daphne Koller, 2003 Car 2 Engine Power Driver 2 Price Income

Probabilistic Relational Models: Relational Markov Networks © Daphne Koller, 2003
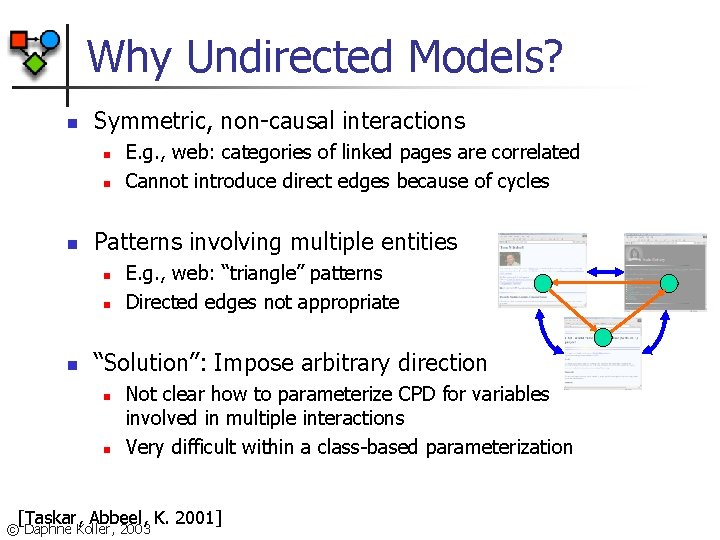
Why Undirected Models? n Symmetric, non-causal interactions n n n Patterns involving multiple entities n n n E. g. , web: categories of linked pages are correlated Cannot introduce direct edges because of cycles E. g. , web: “triangle” patterns Directed edges not appropriate “Solution”: Impose arbitrary direction n n Not clear how to parameterize CPD for variables involved in multiple interactions Very difficult within a class-based parameterization [Taskar, Abbeel, K. 2001] © Daphne Koller, 2003
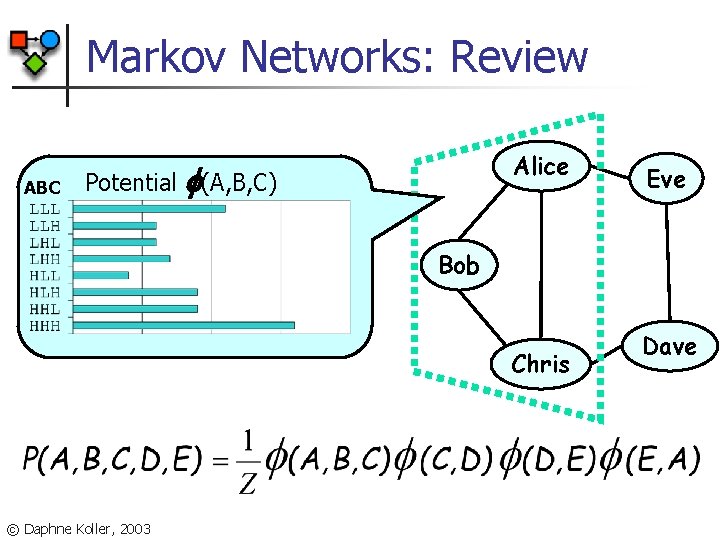
Markov Networks: Review ABC Potential Alice (A, B, C) Eve Bob Chris © Daphne Koller, 2003 Dave

Markov Networks: Review n n A Markov network is an undirected graph over some set of variables V Graph associated with a set of potentials i n n Each potential is factor over subset Vi Variables in Vi must be a (sub)clique in network © Daphne Koller, 2003
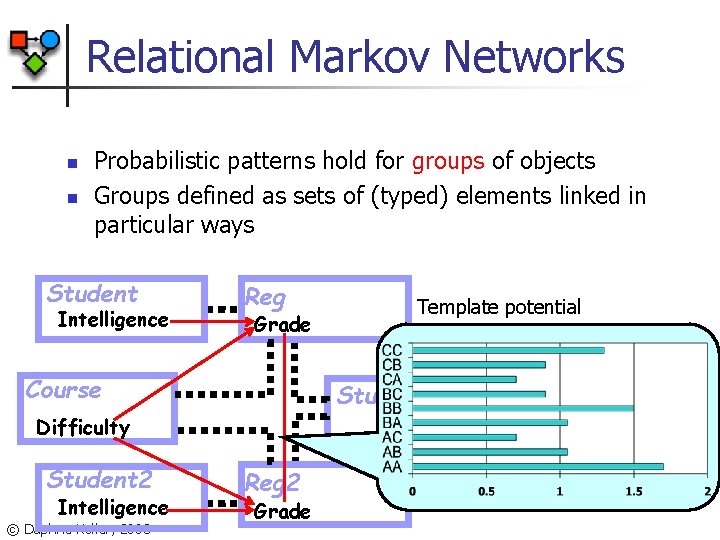
Relational Markov Networks n n Probabilistic patterns hold for groups of objects Groups defined as sets of (typed) elements linked in particular ways Student Intelligence Reg Grade Course Study Group Difficulty Student 2 Intelligence © Daphne Koller, 2003 Template potential Reg 2 Grade

RMN Language n Define clique templates n n n All tuples {reg R 1, reg R 2, group G} s. t. In(G, R 1), In(G, R 2) Compatibility potential (R 1. Grade, R 2. Grade) Ground Markov network contains potential (r 1. Grade, r 2. Grade) for all appropriate r 1, r 2 © Daphne Koller, 2003
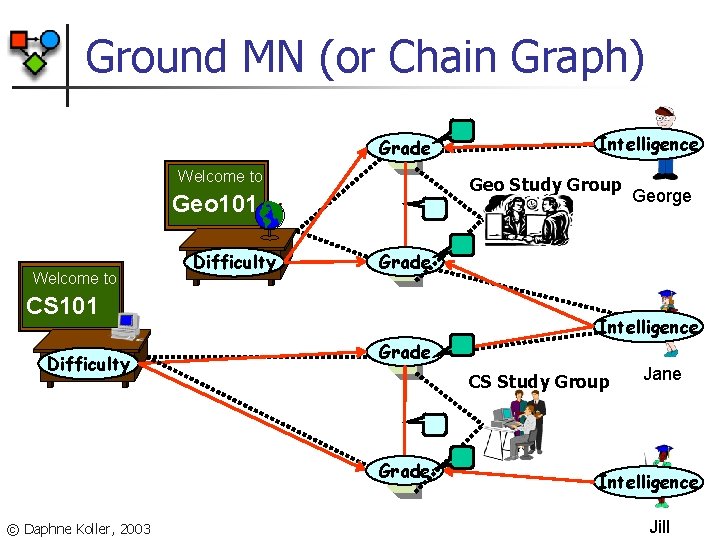
Ground MN (or Chain Graph) Grade Welcome to Geo Study Group Geo 101 Welcome to Difficulty Intelligence Grade CS Study Group Grade © Daphne Koller, 2003 George Grade CS 101 Difficulty Intelligence Jane Intelligence Jill

PRM Inference © Daphne Koller, 2003

Inference: Simple Method n Define ground network as in semantics Apply standard inference methods n Problem: n n n Very large models can be specified very easily Resulting ground network often highly connected Exact inference is typically intractable In practice, often must resort to approximate methods such as belief propagation © Daphne Koller, 2003
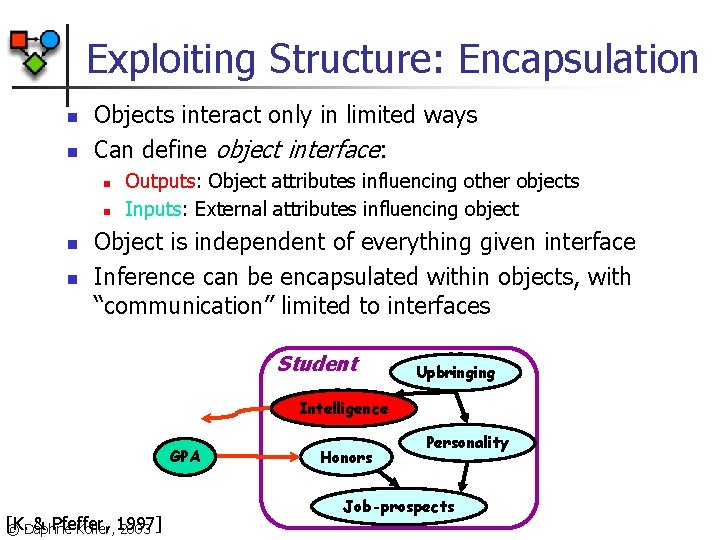
Exploiting Structure: Encapsulation n n Objects interact only in limited ways Can define object interface: n n Outputs: Object attributes influencing other objects Inputs: External attributes influencing object Object is independent of everything given interface Inference can be encapsulated within objects, with “communication” limited to interfaces Student Upbringing Intelligence GPA [K. & Pfeffer, © Daphne Koller, 1997] 2003 Honors Personality Job-prospects

Exploiting Structure: Encapsulation n n Marginalize object distribution onto interface Dependency graph over interfaces induced by n n Perform inference over interfaces If interaction graph has low tree-width, can use exact inference n n Inter-object dependencies And hence by the relational structure E. g. , part-of hierarchy in some OOBNs If relational structure more complex, can use BP n A form of Kikuchi BP, where cluster selection is guided by relational structure © Daphne Koller, 2003
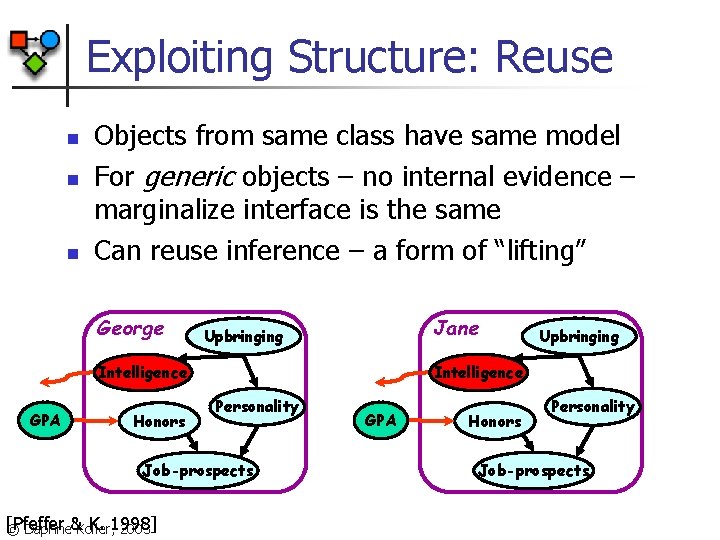
Exploiting Structure: Reuse n n n Objects from same class have same model For generic objects – no internal evidence – marginalize interface is the same Can reuse inference – a form of “lifting” George Jane Upbringing Intelligence GPA Honors Intelligence Personality Job-prospects [Pfeffer K. 1998] © Daphne&Koller, 2003 Upbringing GPA Honors Personality Job-prospects

Exploiting Structure: Reuse n Generic objects often play same role in model n n n Multiple students that all take the same class Reuse: compute interface once Combinatorics: compute total contribution to probability in closed form P(k students like the class | teaching-ability = low) = P(generic student likes the class | teaching ability = low)k [Pfeffer K. 1998] © Daphne&Koller, 2003

Case study Battlefield situation assessment for missile units n n several locations many units each has detailed model Example object classes: n n n Battalion Battery Vehicle Location Weather. © Daphne Koller, 2003 n Example relations: n n At-Location Has-Weather Sub-battery/In-battalion Sub-vehicle/In-battery
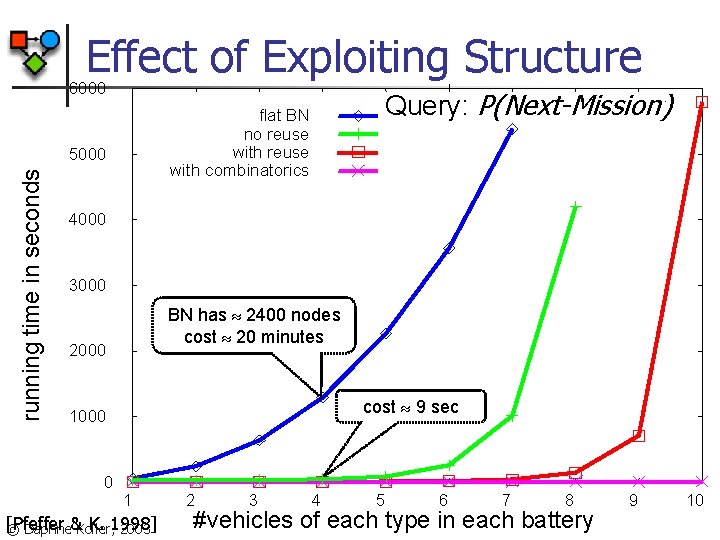
Effect of Exploiting Structure 6000 5000 running time in seconds Query: P(Next-Mission) flat BN no reuse with combinatorics 4000 3000 BN has 2400 nodes cost 20 minutes 2000 cost 9 sec 1000 0 1 [Pfeffer K. 1998] © Daphne&Koller, 2003 2 3 4 5 6 7 8 #vehicles of each type in each battery 9 10

PRM Learning © Daphne Koller, 2003

Outline n Relational Bayesian networks n n n Relational Markov networks n n Likelihood function ML parameter estimation EM Structure learning Parameter estimation Applications: n n Collective classification – web data Relational clustering – biological data © Daphne Koller, 2003
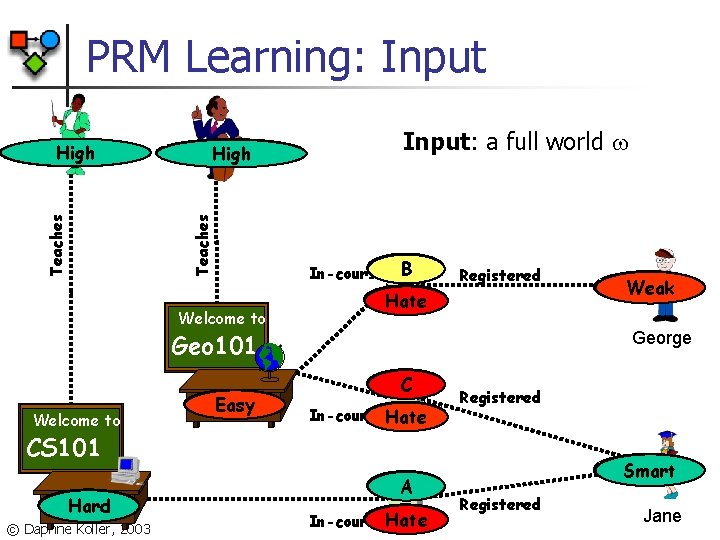
PRM Learning: Input Prof. Smith Teaching-ability High Teaches Prof. Jones High Teaching-ability Welcome to Input: a full world B In-course. Grade Registered Satisfac Hate Geo 101 Welcome to Difficulty Easy George Grade C In-course Satisfac Hate Registered CS 101 Difficulty Hard © Daphne Koller, 2003 Intelligence Weak Grade A Satisfac Hate In-course Intelligence Smart Registered Jane

Likelihood Function Attributes Objects n n Likelihood of a BN with shared parameters Joint likelihood is a product of likelihood terms n n One for each attribute X. A and its family For each X. A, the likelihood function aggregates counts from all occurrences x. A in world [Friedman, Getoor, © Daphne Koller, 2003 K. , Pfeffer, 1999]

Likelihood Function: Multinomials Log-likelihood: Sufficient statistics: © Daphne Koller, 2003

RBN Parameter Estimation n MLE parameters: n Bayesian estimation: n n Prior for each attribute X. A Posterior uses aggregated sufficient statistics © Daphne Koller, 2003

Learning w. Missing Data n EM Algorithm applies essentially unchanged n n n E-step computes expected sufficient statistics, aggregated over all objects in class M-step uses ML (or MAP) parameter estimation Key difference: n n n In general, the hidden variables are not independent Computation of expected sufficient statistics requires inference over entire network [Same reasoning as forward-backward algorithm in temporal models] © Daphne Koller, 2003
![Learning w. Missing Data: EM Students [Dempster et al. 77] Courses A B C Learning w. Missing Data: EM Students [Dempster et al. 77] Courses A B C](http://slidetodoc.com/presentation_image_h2/c6681bd5b58fc444db3e9390e221cff9/image-61.jpg)
Learning w. Missing Data: EM Students [Dempster et al. 77] Courses A B C P(Registration. Grade | Course. Difficulty, Student. Intelligence) easy / hard © Daphne Koller, 2003 low / high

Learning RBN Structure n Define set of legal RBN structures n n Ones with legal class dependency graphs Define scoring function Bayesian score n Product of family scores: One for each X. A n Uses aggregated sufficient statistics n n Search for high-scoring legal structure [Friedman, Getoor, © Daphne Koller, 2003 K. , Pfeffer, 1999]

Learning RBN Structure n All operations done at class level n n Dependency structure = parents for X. A Acyclicity checked using class dependency graph Score computed at class level Individual objects only contribute to sufficient statistics n Can be obtained efficiently using standard DB queries © Daphne Koller, 2003
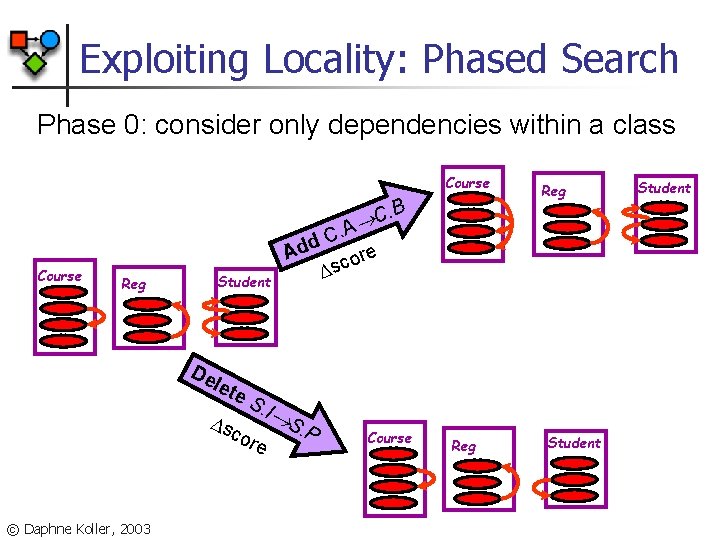
Exploiting Locality: Phased Search Phase 0: consider only dependencies within a class Course . B Add Course Reg Student De let s e. S cor . I © Daphne Koller, 2003 e S. P Reg C . A C e or c s Course Reg Student
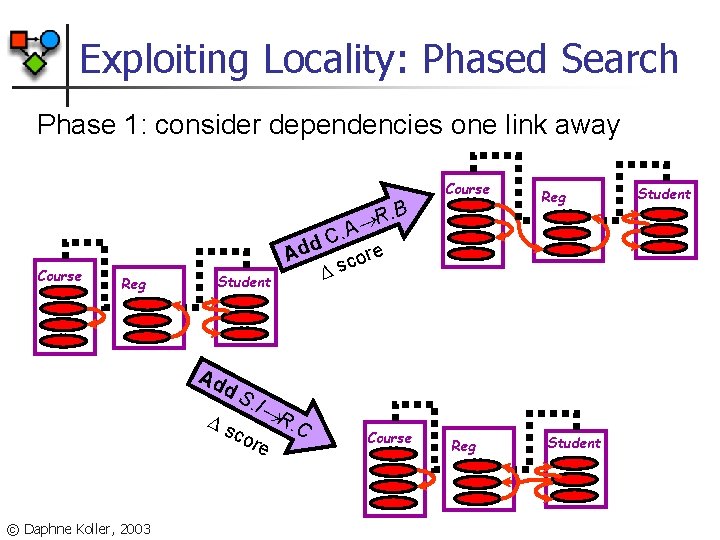
Exploiting Locality: Phased Search Phase 1: consider dependencies one link away Course . B R Course Reg Student Ad d s S. I cor © Daphne Koller, 2003 Reg C. A d re Ad o c s R e . C Course Reg Student
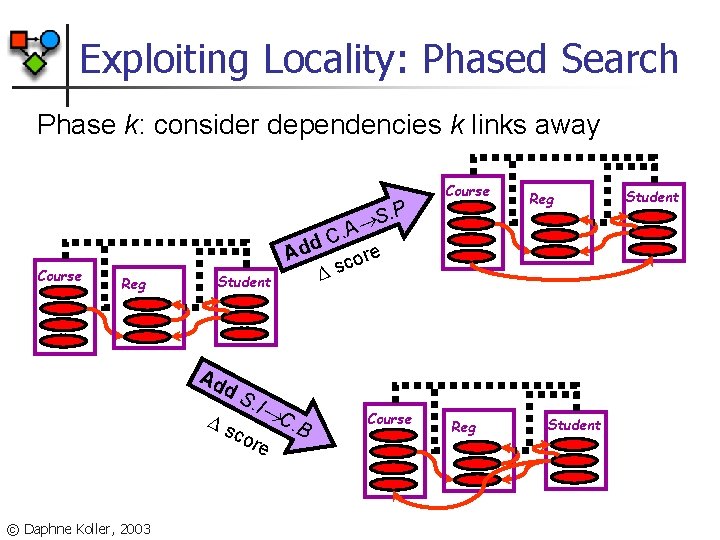
Exploiting Locality: Phased Search Phase k: consider dependencies k links away. P S Course Reg Student Ad d s S. I cor © Daphne Koller, 2003 Reg C. A d Ad re o c s C e Course . B Course Reg Student
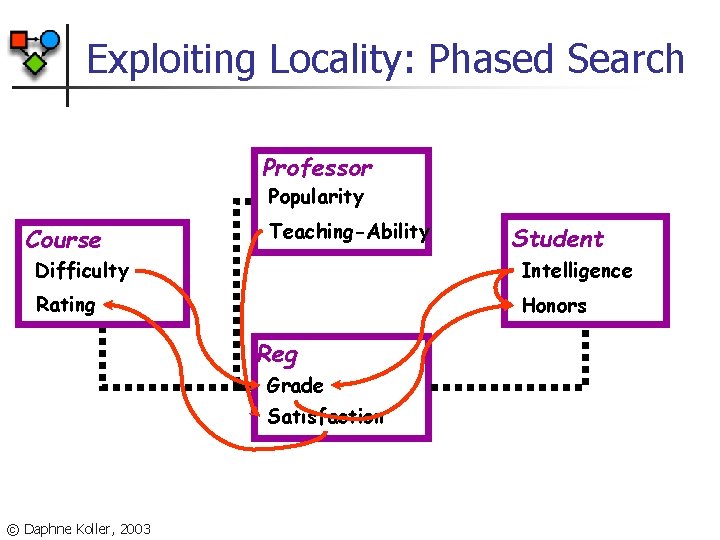
Exploiting Locality: Phased Search Professor Popularity Course Teaching-Ability Student Difficulty Intelligence Rating Honors Reg Grade Satisfaction © Daphne Koller, 2003
![TB Patients in San Francisco Strains Patients [Getoor, Rhee, K. , Small, 2001] © TB Patients in San Francisco Strains Patients [Getoor, Rhee, K. , Small, 2001] ©](http://slidetodoc.com/presentation_image_h2/c6681bd5b58fc444db3e9390e221cff9/image-68.jpg)
TB Patients in San Francisco Strains Patients [Getoor, Rhee, K. , Small, 2001] © Daphne Koller, 2003 Contacts
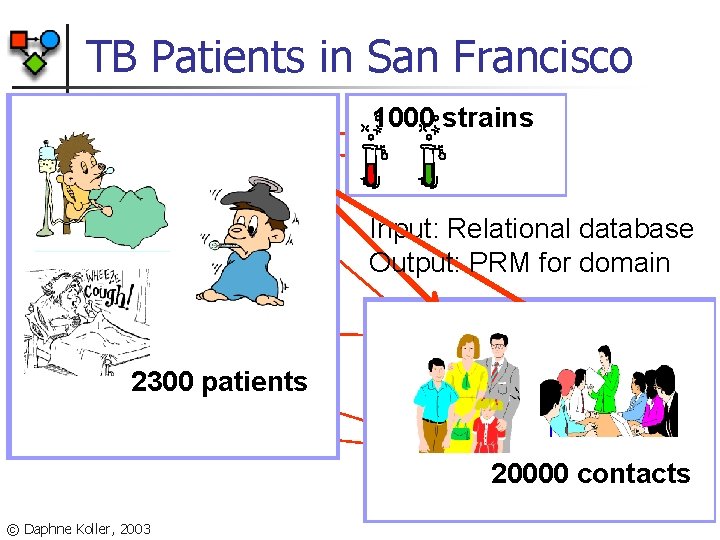
TB Patients in San Francisco 1000 strains Suscept Gender POB Race HIV Age TB-type XRay Outbreak Strain Input: Relational database Output: PRM for domain Infected 2300 patients Patient Smear Subcase Care. Loc Contact © Daphne Koller, 2003 Treatment Close. Cont 20000 contacts Relation

Outline n Relational Bayesian networks n n n Relational Markov networks n n Likelihood function ML parameter estimation EM Structure learning Parameter estimation Applications: n n Collective classification – web data Relational clustering – biological data © Daphne Koller, 2003
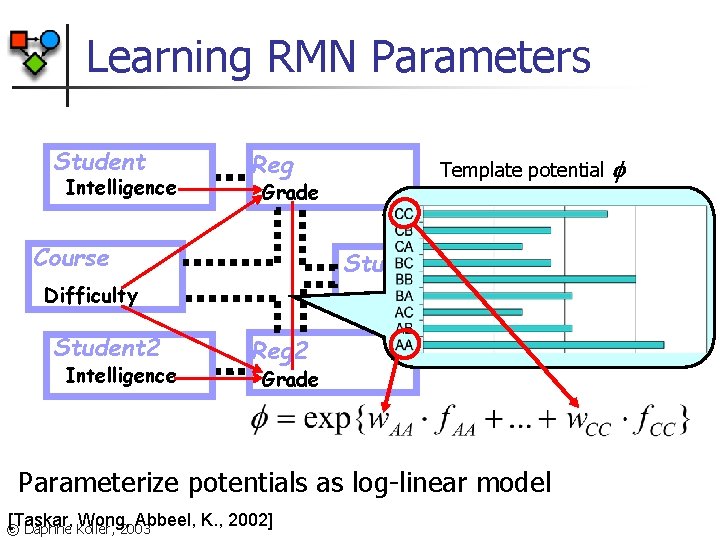
Learning RMN Parameters Student Intelligence Reg Grade Course Template potential Study Group Difficulty Student 2 Intelligence Reg 2 Grade Parameterize potentials as log-linear model [Taskar, Wong, Abbeel, K. , 2002] © Daphne Koller, 2003

Learning RMN Parameters Counts in Partition function For example: f. AA ( ) = # of tuples {reg r 1, reg r 2, group g} s. t. In(g, r 1), In(g, r 2); r 1. Grade=A, r 2. Grade=A © Daphne Koller, 2003

Learning RMN Parameters n n n Parameter estimation is not closed form Convex problem unique global maximum Can use methods such as conjugate gradient actual count - expected count n n Gradient process tries to find parameters s. t. expected counts = actual counts Computing expected counts requires inference over ground Markov network © Daphne Koller, 2003
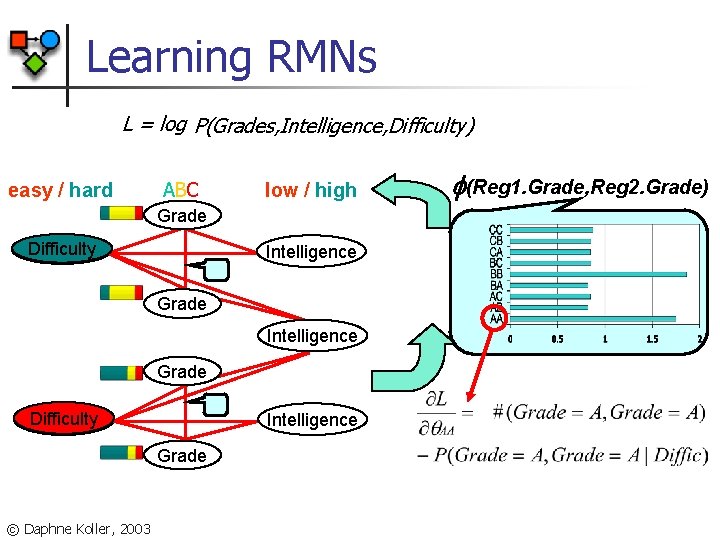
Learning RMNs L = log P(Grades, Intelligence, Difficulty) easy / hard ABC low / high Grade Difficulty Intelligence Grade © Daphne Koller, 2003 (Reg 1. Grade, Reg 2. Grade)

Outline n Relational Bayesian networks n n n Relational Markov networks n n Likelihood function ML parameter estimation EM Structure learning Parameter estimation Applications: n n Collective classification – web data Relational clustering – biological data © Daphne Koller, 2003
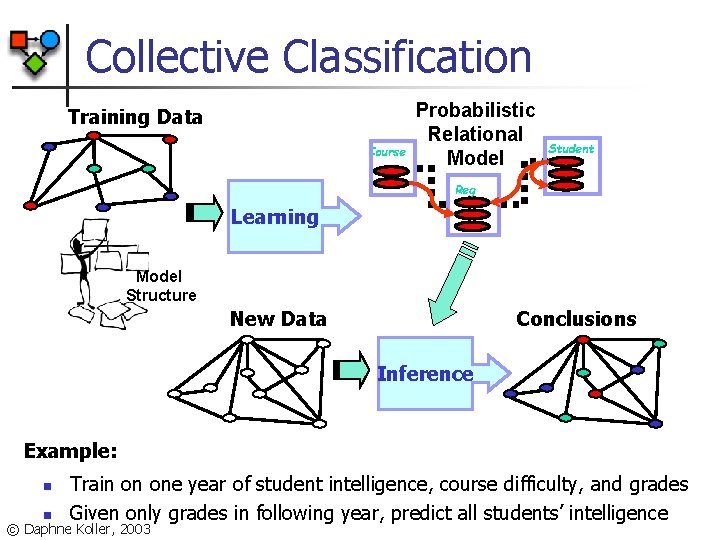
Collective Classification Training Data Course Probabilistic Relational Model Student Reg Learning Model Structure New Data Conclusions Inference Example: n n Train on one year of student intelligence, course difficulty, and grades Given only grades in following year, predict all students’ intelligence © Daphne Koller, 2003

Discriminative Training n n n Goal: Given values of observed variables . O=o, predict desired target values . T=t* Do not necessarily want the model to fit the joint distribution P(. O=o, . T=t*) To maximize classification accuracy, we can consider other optimization criteria n Maximize conditional log likelihood n Maximize margin [Taskar, Abbeel, K. , 2002; Guestrin, ©Taskar, Daphne Koller, 2003 K. 2003] P(second highest probability label)
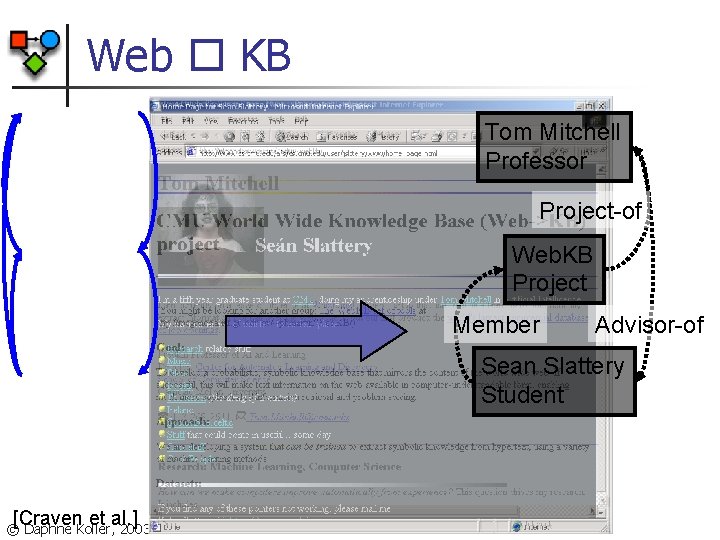
Web KB Tom Mitchell Professor Project-of Web. KB Project Member Advisor-of Sean Slattery Student [Craven et al. ] © Daphne Koller, 2003

Web Classification Experiments n Web. KB dataset n n n Four CS department websites Bag of words on each page Links between pages Anchor text for links Experimental setup n n n Trained on three universities Tested on fourth Repeated for all four combinations © Daphne Koller, 2003
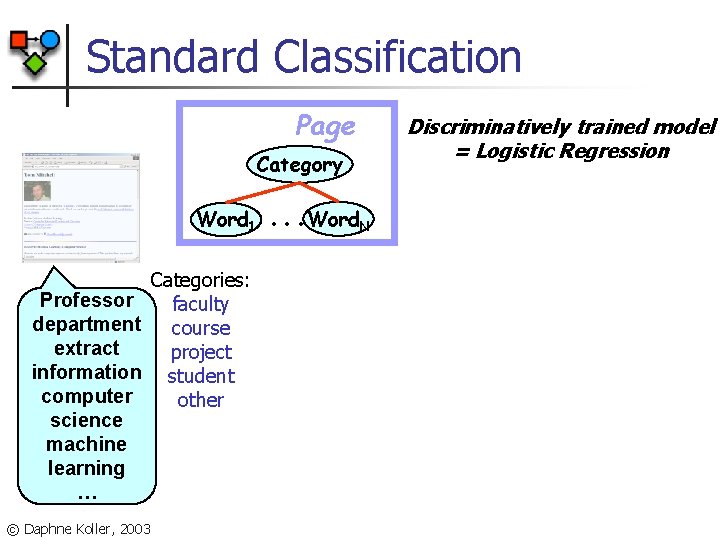
Standard Classification Page Category Word 1 Professor department extract information computer science machine learning … © Daphne Koller, 2003 Categories: faculty course project student other . . . Word. N Discriminatively trained model = Logistic Regression

Standard Classification Page Category . . . Word. N. . . Link. Word. N Word 1 0. 18 working 0. 16 with 0. 14 Tom 0. 12 0. 1 Mitchell 0. 08 … 0. 06 0. 04 0. 02 0 © Daphne Koller, 2003 Logistic
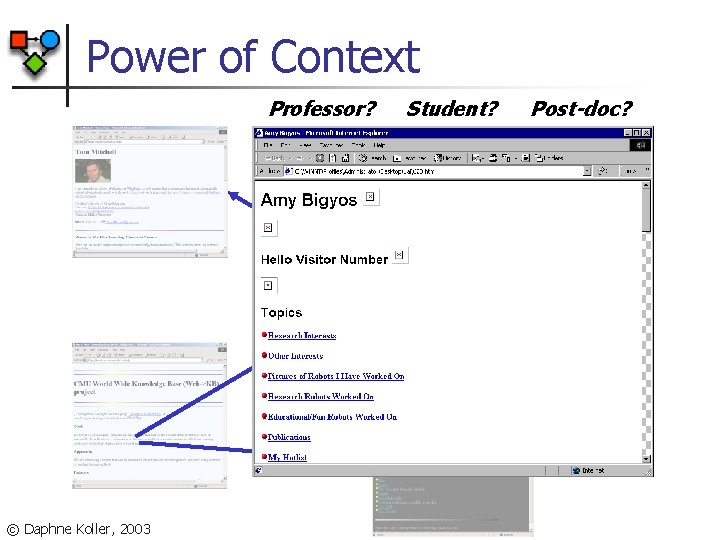
Power of Context Professor? © Daphne Koller, 2003 Student? Post-doc?
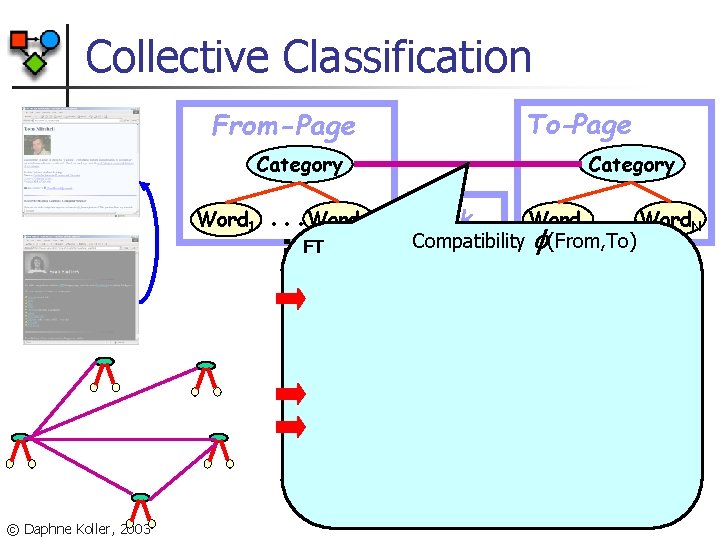
Collective Classification To-Page From-Page Category Word 1 © Daphne Koller, 2003 . . . Word. N FT Link . . . Word. N Word 1 Compatibility (From, To)
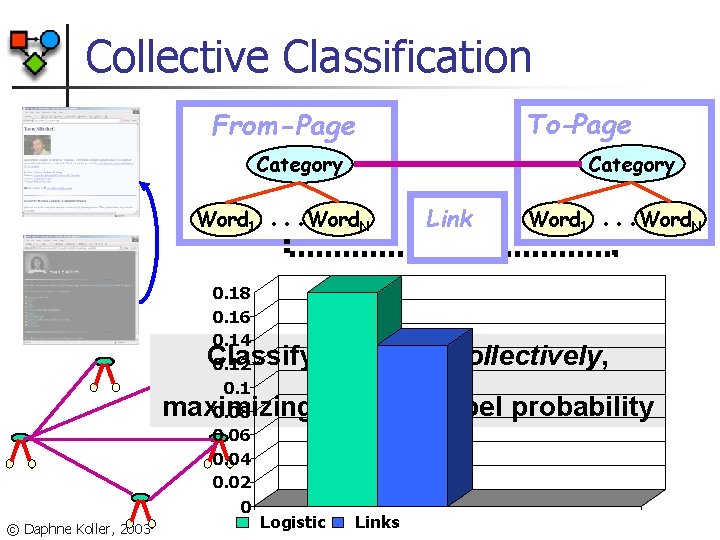
Collective Classification To-Page From-Page Category Word 1 0. 18 0. 16 0. 14 0. 12 0. 1 0. 08 0. 06 0. 04 0. 02 0 . . . Word. N Link Word 1 . . . Word. N Classify all pages collectively, maximizing the joint label probability © Daphne Koller, 2003 Logistic Links
More Complex Structure © Daphne Koller, 2003
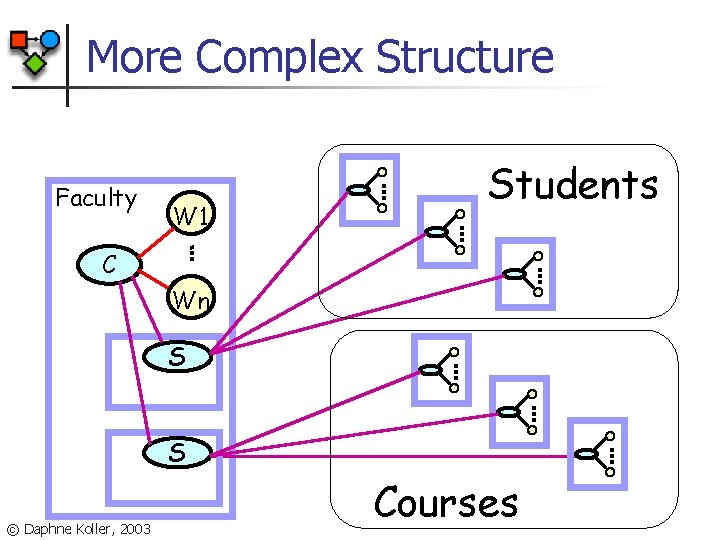
More Complex Structure Faculty W 1 Students C Wn S S © Daphne Koller, 2003 Courses
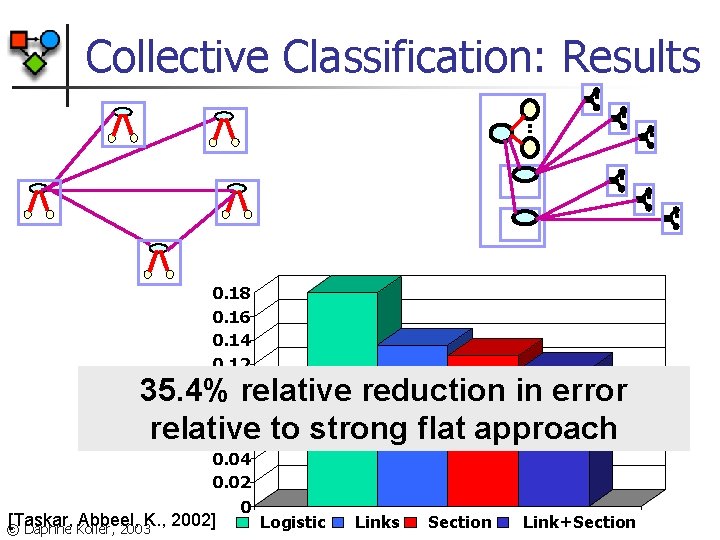
Collective Classification: Results 0. 18 0. 16 0. 14 0. 12 0. 1 0. 08 0. 06 0. 04 0. 02 0 35. 4% relative reduction in error relative to strong flat approach [Taskar, Abbeel, K. , 2002] © Daphne Koller, 2003 Logistic Links Section Link+Section
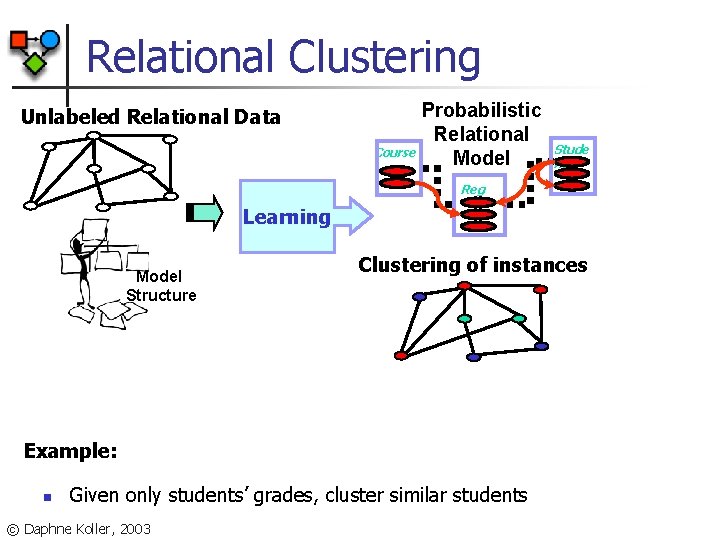
Relational Clustering Unlabeled Relational Data Course Probabilistic Relational Model Stude nt Reg Learning Model Structure Clustering of instances Example: n Given only students’ grades, cluster similar students © Daphne Koller, 2003
Movie Data © Daphne Koller, 2003 Internet Movie Database http: //www. imdb. com
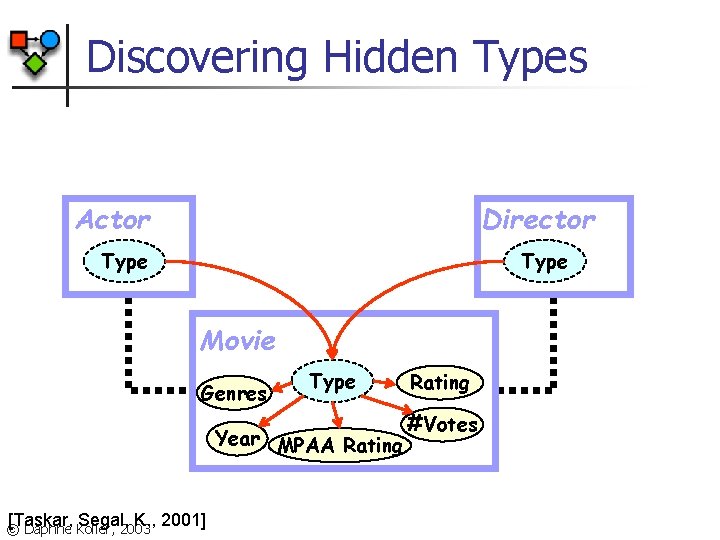
Discovering Hidden Types Actor Director Type Movie Genres Type Year MPAA Rating [Taskar, Segal, K. , 2001] © Daphne Koller, 2003 Rating #Votes

Discovering Hidden Types Movies Actors Directors Wizard of Oz Cinderella Sound of Music The Love Bug Pollyanna The Parent Trap Mary Poppins Swiss Family Robinson Sylvester Stallone Bruce Willis Harrison Ford Steven Seagal Kurt Russell Kevin Costner Jean-Claude Van Damme Arnold Schwarzenegger Alfred Hitchcock Stanley Kubrick David Lean Milos Forman Terry Gilliam Francis Coppola Terminator 2 Batman Forever Golden. Eye Starship Troopers Mission: Impossible Hunt for Red October Anthony Hopkins Robert De Niro Tommy Lee Jones Harvey Keitel Morgan Freeman Gary Oldman Steven Spielberg Tim Burton Tony Scott James Cameron John Mc. Tiernan Joel Schumacher … © Daphne Koller, 2003 …
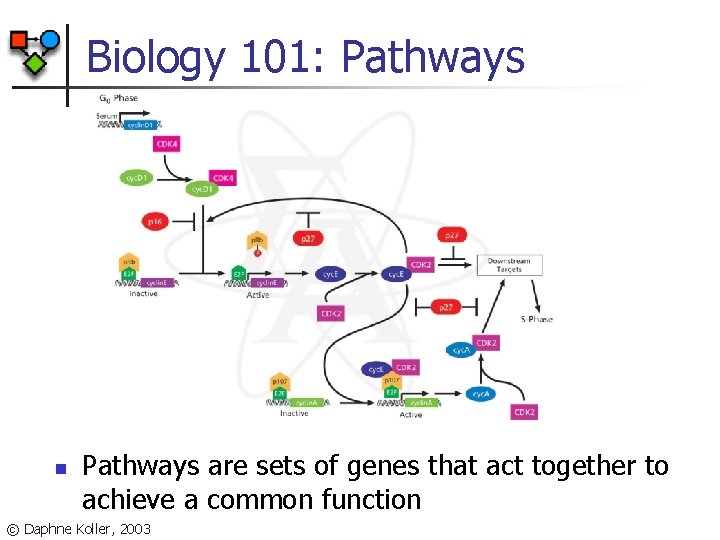
Biology 101: Pathways n Pathways are sets of genes that act together to achieve a common function © Daphne Koller, 2003
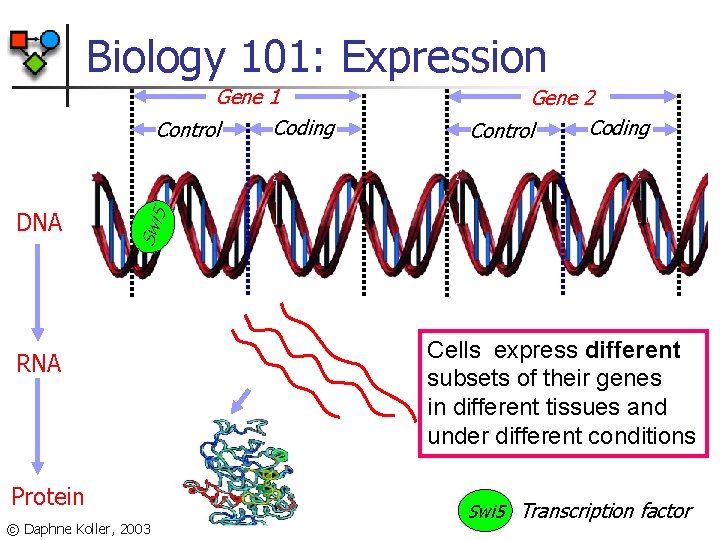
Biology 101: Expression DNA Gene 2 Coding Control Swi 5 Gene 1 Coding Control RNA Protein © Daphne Koller, 2003 Cells express different subsets of their genes in different tissues and under different conditions Swi 5 Transcription factor
Finding Pathways: Attempt I n Use protein-protein interaction data Pathway III © Daphne Koller, 2003

Finding Pathways: Attempt I n n Use protein-protein interaction data Problems: n n Data is very noisy Structure is lost: n Large connected component (3527/3589 genes) in interaction graph © Daphne Koller, 2003

Finding Pathways: Attempt II n Use gene expression data n Thousands of arrays available under different conditions Pathway I Clustering Pathway II © Daphne Koller, 2003

Finding Pathways: Attempt II n Use gene expression data n n Thousands of arrays available under different conditions Problems: n n n Expression is only ‘weak’ indicator of interaction Data is noisy Interacting pathways are not separable © Daphne Koller, 2003 Pathway II
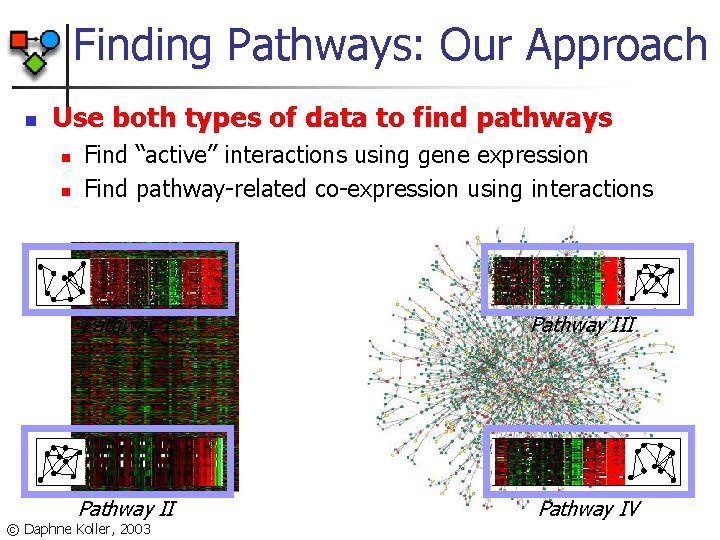
Finding Pathways: Our Approach n Use both types of data to find pathways n n Find “active” interactions using gene expression Find pathway-related co-expression using interactions Pathway III Pathway IV © Daphne Koller, 2003

Probabilistic Model n Genes are partitioned into “pathways”: n n n Expression component: n n Every gene is assigned to one of ‘k’ pathways Random variable for each gene with domain {1, …, k} Model likelihood is higher when genes in the same pathway have similar expression profiles Interaction component: n Model likelihood is higher when genes in the same pathway interact [Segal, Wang, K. , 2003] © Daphne Koller, 2003
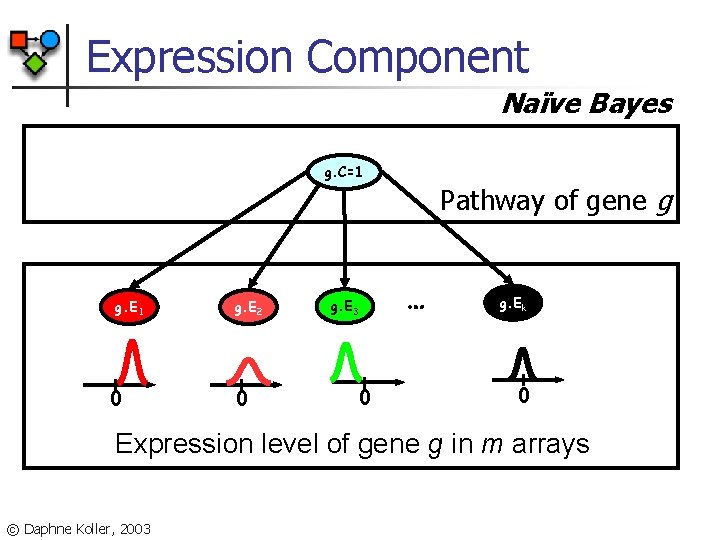
Expression Component Naïve Bayes g. C=1 g. E 1 0 g. E 2 0 g. E 3 0 Pathway of gene g … g. Ek 0 Expression level of gene g in m arrays © Daphne Koller, 2003
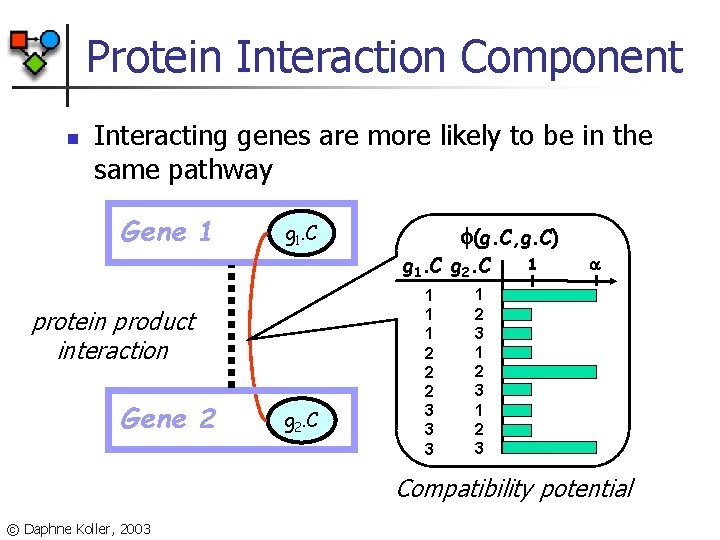
Protein Interaction Component n Interacting genes are more likely to be in the same pathway Gene 1 g 1. C (g. C, g. C) g 1. C g 2. C protein product interaction Gene 2 g 2. C 1 1 1 2 2 2 3 3 3 1 1 2 3 Compatibility potential © Daphne Koller, 2003
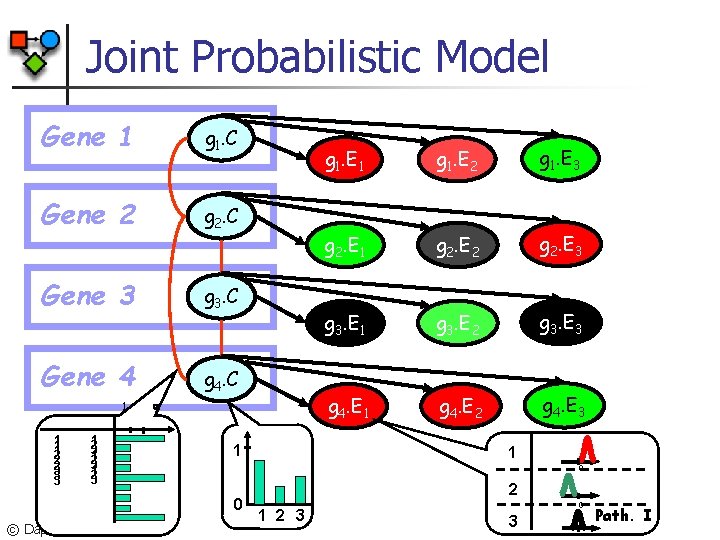
Joint Probabilistic Model Gene 1 g 1. C Gene 2 g 2. C Gene 3 g 3. C Gene 4 g 4. C 1 1 2 2 2 3 3 3 1 2 3 1 g 1. E 2 g 1. E 3 g 2. E 1 g 2. E 2 g 2. E 3 g 3. E 1 g 3. E 2 g 3. E 3 g 4. E 1 g 4. E 2 g 4. E 3 1 0 0 © Daphne Koller, 2003 g 1. E 1 2 3 0 3 Path. I
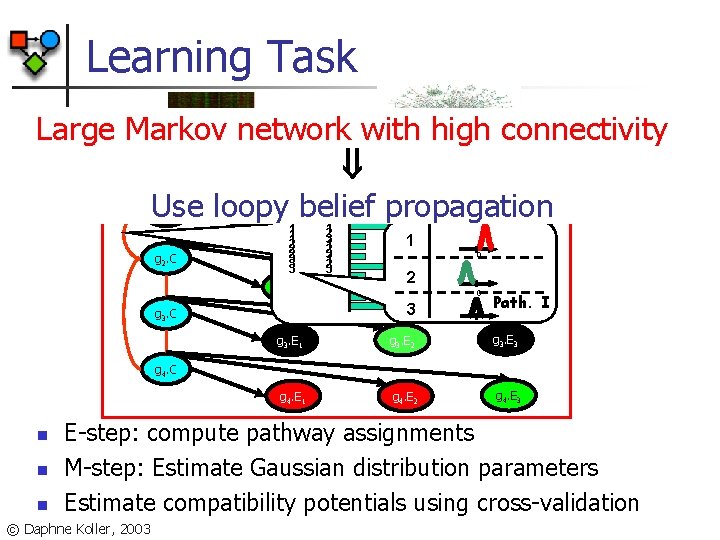
Learning Task Large Markov network with high connectivity Use loopy belief propagation 1 g 1. C g 2. C 1 g 1. E 1 1 1 2 2 2 3 3 3 g 2. E 1 1 2 3 1 0 2 g 2. E 2 3 g 3. C g 1. E 3 g 1. E 2 0 g 2. E 3 Path. I g 3. E 1 g 3. E 2 g 3. E 3 g 4. E 1 g 4. E 2 g 4. E 3 g 4. C n n n E-step: compute pathway assignments M-step: Estimate Gaussian distribution parameters Estimate compatibility potentials using cross-validation © Daphne Koller, 2003
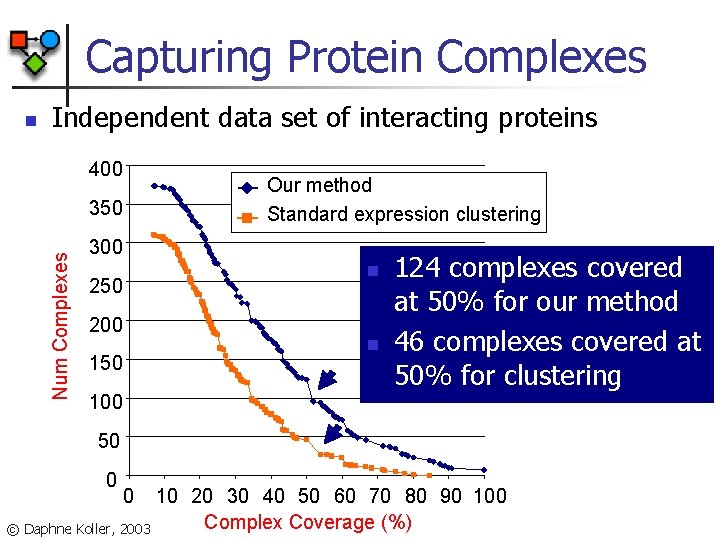
Capturing Protein Complexes n Independent data set of interacting proteins 400 Our method Standard expression clustering Num Complexes 350 300 n 250 200 n 150 124 complexes covered at 50% for our method 46 complexes covered at 50% for clustering 100 50 0 0 10 20 30 40 50 60 70 80 90 100 Complex Coverage (%) © Daphne Koller, 2003
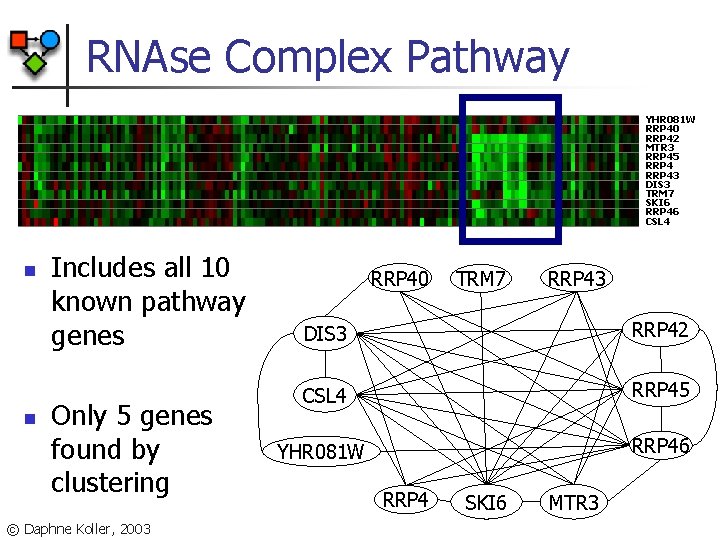
RNAse Complex Pathway YHR 081 W RRP 40 RRP 42 MTR 3 RRP 45 RRP 43 DIS 3 TRM 7 SKI 6 RRP 46 CSL 4 n n Includes all 10 known pathway genes Only 5 genes found by clustering © Daphne Koller, 2003 RRP 40 TRM 7 RRP 43 DIS 3 RRP 42 CSL 4 RRP 45 YHR 081 W RRP 46 RRP 4 SKI 6 MTR 3
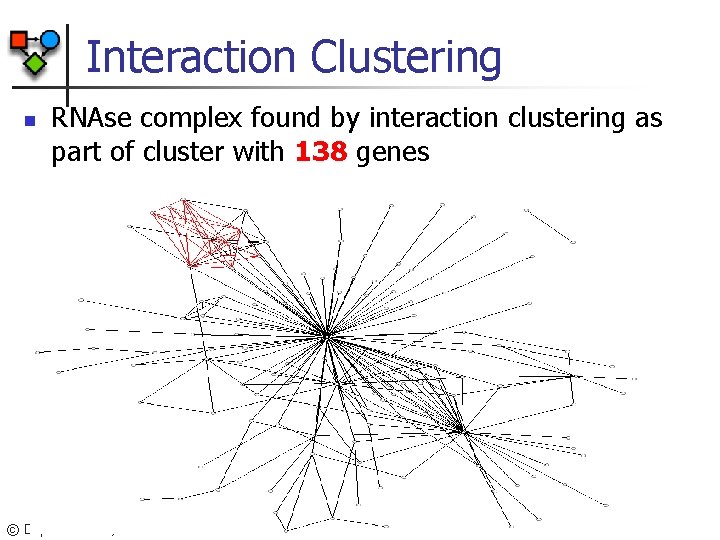
Interaction Clustering n RNAse complex found by interaction clustering as part of cluster with 138 genes © Daphne Koller, 2003

Uncertainty about Domain Structure or PRMs are not just template BNs/MNs © Daphne Koller, 2003

Structural Uncertainty n Class uncertainty: n n Relational uncertainty: n n To which class does an object belong What is the relational (link) structure Identity uncertainty: n n Which “names” refer to the same objects Also covers data association © Daphne Koller, 2003

Relational Uncertainty Probability distribution over graph structures n n Link existence model n E. g. , hyperlinks between webpages n Each potential link is a separate event n Its existence is a random variable Link reference model n E. g. , instructors teaching a course n n Distribution over # of endpoints for each outgoing link Each link has distribution over link endpoint n n e. g. , instructor link for CS course likely to point to CS prof Many other models possible [Getoor, Friedman, K. Taskar, 2002] © Daphne Koller, 2003

Link Existence Model n Background knowledge is an object skeleton n n A set of entity objects PRM defines distribution over worlds n n Assignments of values to all attributes Existence of links between objects [Getoor, Friedman, K. Taskar, 2002] © Daphne Koller, 2003

Link Existence as a PRM n Define objects for any potential links n n n Potential link object for any pair of webpages w 1, w 2 Each potential link object has link existence attribute, denoting whether the link exists or not Link existence variables have probabilistic model Page-from Type Words Page-to Link [Getoor, Friedman, K. Taskar, 2002] © Daphne Koller, 2003 Exists Type Words From. Type To. Type False True Student Professor 0. 999 0. 995 0. 998 0. 999 0001 0005 0002 0001

Why Are Link Models Useful? n Predict which links exist in the world n n n Which professor teaches each course Which student will register for which course Use known links to infer values of attributes n n Given that student registered for a hard class, is she more likely to be a graduate student Given that one page points to another, is it more likely to be a faculty page? © Daphne Koller, 2003
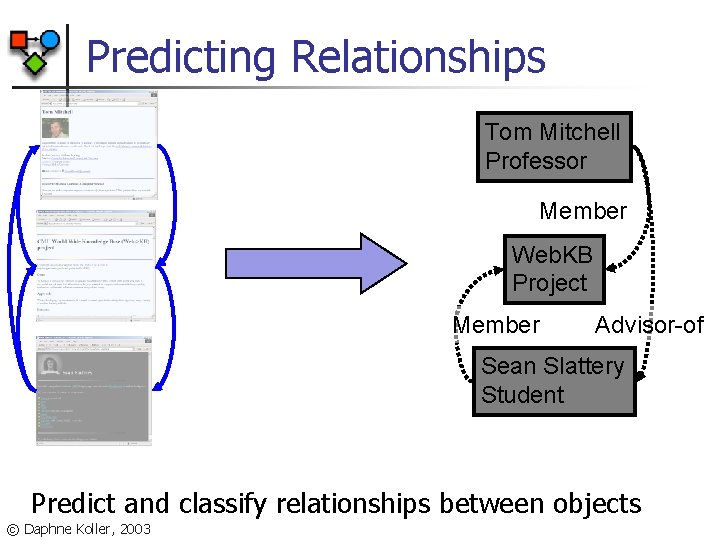
Predicting Relationships Tom Mitchell Professor Member Web. KB Project Member Advisor-of Sean Slattery Student Predict and classify relationships between objects © Daphne Koller, 2003
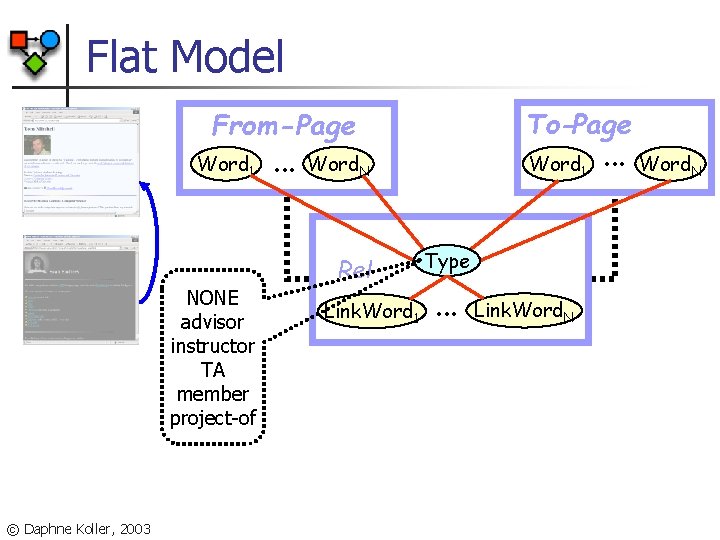
Flat Model To-Page From-Page Word 1 NONE advisor instructor TA member project-of © Daphne Koller, 2003 . . . Word 1 Word. N Rel Link. Word 1 Type . . . Link. Word. N . . . Word. N
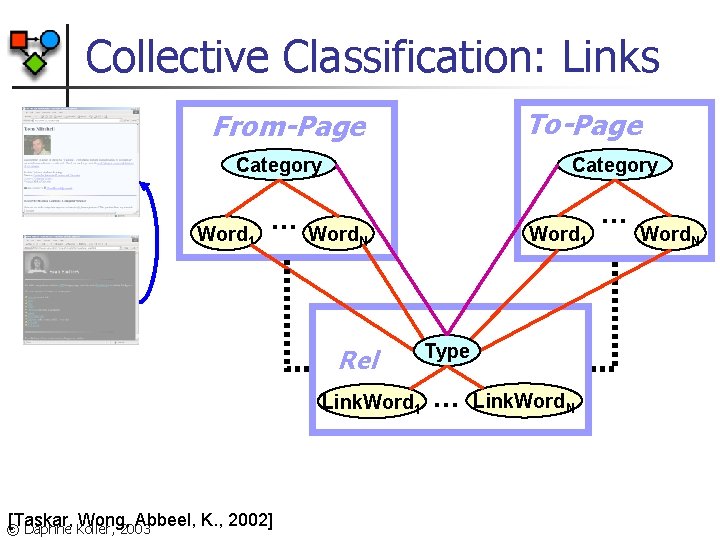
Collective Classification: Links To-Page From-Page Category . . . Word 1 Rel Link. Word 1 [Taskar, Wong, Abbeel, K. , 2002] © Daphne Koller, 2003 Word 1 Word. N Type . . . Link. Word. N
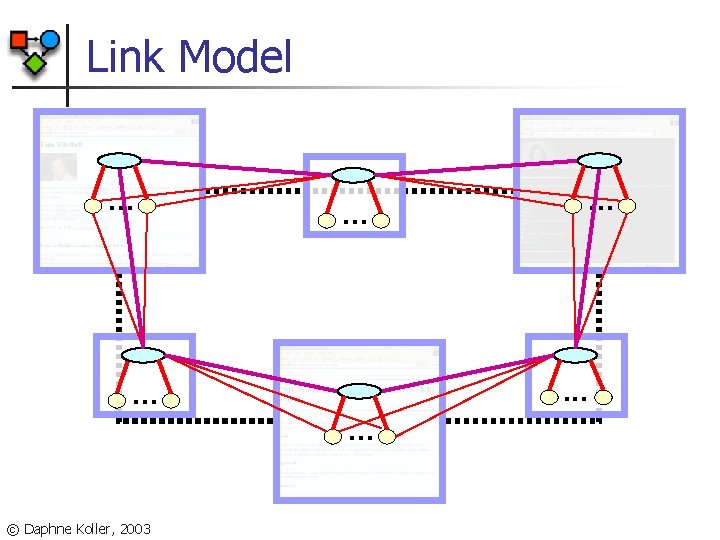
Link Model. . . . © Daphne Koller, 2003 . . .
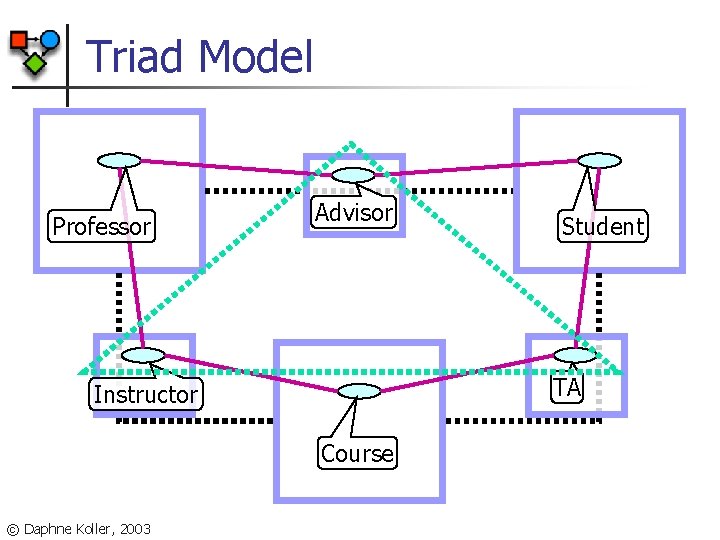
Triad Model Professor Advisor TA Instructor Course © Daphne Koller, 2003 Student
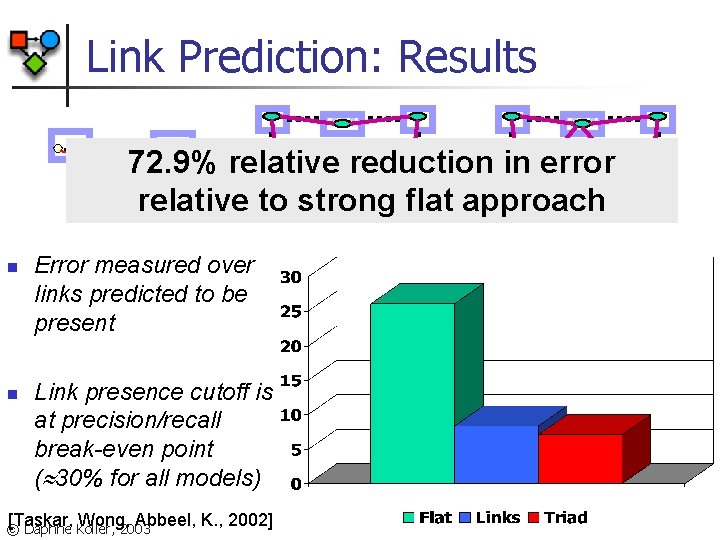
Link Prediction: Results. . . n n . . . 72. 9% relative reduction in error. . . relative to strong flat approach Error measured over links predicted to be present Link presence cutoff is at precision/recall break-even point ( 30% for all models) [Taskar, Wong, Abbeel, K. , 2002] © Daphne Koller, 2003

Identity Uncertainty Model n Background knowledge is an object universe n n A set of potential objects PRM defines distribution over worlds n n n Assignments of values to object attributes Partition of objects into equivalence classes Objects in same class have same attribute values [Pasula, Marthi, Milch, Russell, Shpitser, 2002] © Daphne Koller, 2003
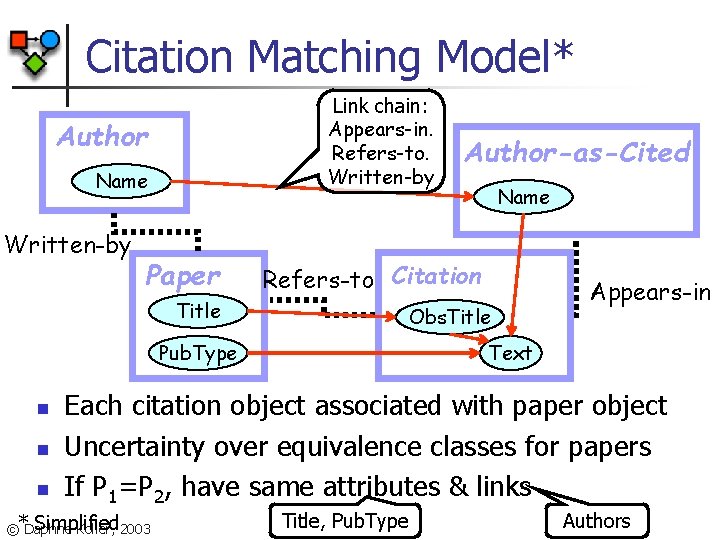
Citation Matching Model* Link chain: Appears-in. Refers-to. Written-by Author Name Written-by Paper Obs. Title Pub. Type n n Name Refers-to Citation Title n Author-as-Cited Appears-in Text Each citation object associated with paper object Uncertainty over equivalence classes for papers If P 1=P 2, have same attributes & links Simplified ©*Daphne Koller, 2003 Title, Pub. Type Authors

Identity Uncertainty n Depending on choice of equivalence classes: n n Number of objects changes Dependency structure changes n No “nice” corresponding ground BN n Algorithm: n n n Each partition hypothesis defines simple BN Use MCMC over equivalence class partition Exact inference over resulting BN defines acceptance probability for Markov chain [Pasula, Marthi, Milch, Russell, Shpitser, 2002] © Daphne Koller, 2003

Identity Uncertainty Results 61. 5% relative reduction in error relative to state of the art Accuracy of citation recovery: % of actual citation clusters recovered perfectly [Pasula, Marthi, Milch, © Daphne Koller, 2003 Russell, Shpitser, 2002]

Summary: PRMs … n Inherit the advantages of graphical models: n n n Coherent probabilistic semantics Exploit structure of local interactions Allow us to represent the world in terms of: n n Objects Classes of objects Properties of objects Relations © Daphne Koller, 2003

So What Do We Gain? n Convenient language for specifying complex models n “Web of influence”: subtle & intuitive reasoning n A mechanism for tying parameters and structure n n n within models across models Framework for learning from relational and heterogeneous data © Daphne Koller, 2003

So What Do We Gain? New way of thinking about models & problems n Incorporating heterogeneous data by connecting related entities n New problems: n n n Collective classification Relational clustering Uncertainty about richer structures: n n Link graph structure Identity © Daphne Koller, 2003

But What Do We Really Gain? n Simple PRMs relational logic w. fixed domain and only n n Induce a “propositional” BN Can augment language with additional expressivity Existentialjust quantifiers & functions Aren PRMs a convenient language for n Equality specifying attribute-based graphical models? n Resulting language is inherently more expressive, allowing us to represent distributions over n n worlds where dependencies vary significantly [Getoor et al. , Pasula et al. ] worlds with different numbers of objects [Pfeffer et al. , Pasula et al. ] worlds with infinitely many objects [Pfeffer & K. ] Big questions: Inference & Learning © Daphne Koller, 2003

§ § § Pieter Abbeel Nir Friedman Lise Getoor Carlos Guestrin Brian Milch Avi Pfeffer § § § Eran Segal Ken Takusagawa Ben Taskar Haidong Wang Ming-Fai Wong http: //robotics. stanford. edu/~koller/ http: //dags. stanford. edu/ © Daphne Koller, 2003
- Slides: 127