Primer Design Dr Baha Eldin Khalid Elamin Ph

Primer Design Dr. Baha Eldin Khalid Elamin Ph. D Molecular Biology & Biotechnology

PCR Primer Design Guidelines 1. Primer Length 2. Primer Melting Temperature 3. Primer annealing temperature 4. GC Content 5. GC Clamp 6. Primer Secondary Structures 7. Repeats 8. Runs 9. 3' End Stability 10. Avoid Template secondary structure 11. Avoid Cross homology

1. Primer Length: It is generally accepted that the optimal length of PCR primers is 18 -22 bp. This length is long enough for adequate specificity, and short enough for primers to bind easily to the template at the annealing temperature.

2. Primer Melting Temperature (Tm) by definition is the temperature at which one half of the DNA duplex will dissociate to become single stranded and indicates the duplex stability. Primers with melting temperatures in the range of 52 -58 o C generally produce the best results. Primers with melting temperatures above 65 o. C have a tendency for secondary annealing. The GC content of the sequence gives a fair indication of the primer Tm. All our products calculate it using the nearest neighbor thermodynamic theory, accepted as a much superior method for estimating it, which is considered the most recent and best available.

3. Primer annealing temperature : The primer melting temperature is the estimate of the DNADNA hybrid stability and critical in determining the annealing temperature. Too high Ta will produce insufficient primer-template hybridization resulting in low PCR product yield. Too low Ta may possibly lead to non-specific products caused by a high number of base pair mismatches, . Mismatch tolerance is found to have the strongest influence on PCR specificity.

4. GC Content : The GC content (the number of G's and C's in the primer as a percentage of the total bases) of primer should be 40 -60%.

5. GC Clamp : The presence of G or C bases within the last five bases from the 3' end of primers (GC clamp) helps promote specific binding at the 3' end due to the stronger bonding of G and C bases. More than 3 G's or C's should be avoided in the last 5 bases at the 3' end of the primer.

6. Primer Secondary Structures : Presence of the primer secondary structures produced by intermolecular or intramolecular interactions can lead to poor or no yield of the product. They adversely affect primer template annealing and thus the amplification. They greatly reduce the availability of primers to the reaction. i) Hairpins : It is formed by intramolecular interaction within the primer and should be avoided. Optimally a 3' end hairpin with a ΔG of -2 kcal/mol and an internal hairpin with a ΔG of -3 kcal/mol is tolerated generally.

ii) Self Dimer : A primer self-dimer is formed by intermolecular interactions between the two (same sense) primers, where the primer is homologous to itself. Generally a large amount of primers are used in PCR compared to the amount of target gene. When primers form intermolecular dimers much more readily than hybridizing to target DNA, they reduce the product yield. Optimally a 3' end self dimer with a ΔG of -5 kcal/mol and an internal self dimer with a ΔG of -6 kcal/mol is tolerated generally.

iii) Cross Dimer : Primer cross dimers are formed by intermolecular interaction between sense and antisense primers, where they are homologous. Optimally a 3' end cross dimer with a ΔG of -5 kcal/mol and an internal cross dimer with a ΔG of -6 kcal/mol is tolerated generally.

7. Repeats : A repeat is a di-nucleotide occurring many times consecutively and should be avoided because they can misprime. For example: ATAT. A maximum number of di-nucleotide repeats acceptable in an oligo is 4 di-nucleotides.

8. Runs : Primers with long runs of a single base should generally be avoided as they can misprime. For example, AGCGGGGGATGGGG has runs of base 'G' of value 5 and 4. A maximum number of runs accepted is 4 bp.

9. 3' End Stability : It is the maximum ΔG value of the five bases from the 3' end. An unstable 3' end (less negative ΔG) will result in less false priming.

10. Avoid Template secondary structure : A single stranded Nucleic acid sequences is highly unstable and fold into conformations (secondary structures). The stability of these template secondary structures depends largely on their free energy and melting temperatures(Tm). Consideration of template secondary structures is important in designing primers, especially in q. PCR. If primers are designed on a secondary structures which is stable even above the annealing temperatures, the primers are unable to bind to the template and the yield of PCR product is significantly affected. Hence, it is important to design primers in the regions of the templates that do not form stable secondary structures during the PCR reaction. Our products determine the secondary structures of the template and design primers avoiding them.

11. Avoid Cross homology : To improve specificity of the primers it is necessary to avoid regions of homology. Primers designed for a sequence must not amplify other genes in the mixture. Commonly, primers are designed and then BLASTed to test the specificity. Our products offer a better alternative. You can avoid regions of cross homology while designing primers. You can BLAST the templates against the appropriate non-redundant database and the software will interpret the results. It will identify regions significant cross homologies in each template and avoid them during primer search.
TAACATCATCAATAATATACCTTATTTTGGATTGAAGCCAATATGATAATGAGGGGGTGGAG TTTGTGACGTGGCGCGGGGCGTGGGAACGGGGCGGGTGACGTAGTAGTGTGGCGGAAGT GTGATGTTGCAAGTGTGGCGGAACACATGTAAGCGACGGATGTGGCAAAAGTGACGTTTTT GGTGTGCGCCGGTGTACACAGGAAGTGACAATTTTCGCGCGGTTTTAGGCGGATGTTGTA GTAAATTTGGGCGTAACCGAGTAAGATTTGGCCATTTTCGCGGGAAAACTGAATAAGAGGA AGTGAAATCTGAATAATTTTGTGTTACTCATAGCGCGTAATATTTGTCTAGGGCCGCGGGG ACTTTGACCGTTTACGTGGAGACTCGCCCAGGTGTTTTTCTCAGGTGTTTTCCGCGTTCCG GGTCAAAGTTGGCGTTTTATTATTATAGTCAGCTGACGTGTATTTATACCCGGTGAG TTCCTCAAGAGGCCACTCTTGAGTGCCAGCGAGTAGAGTTTTCTCCTCCGAGCCGCTCCGA CACCGGGACTGAAAATGAGACATATTATCTGCCACGGAGGTGTTATTACCGAAGAAATGGC CGCCAGTCTTTTGGACCAGCTGATCGAAGAGGTACTGGCTGATAATCTTCCACCTCCTAGC CATTTTGAACCACCTACCCTTCACGAACTGTATGATTTAGACGTGACGGCCCCCGAAGATC CCAACGAGGAGGCGGTTTCGCAGATTTTTCCCGACTCTGTAATGTTGGCGGTGCAGGAAG GGATTGACTTACTCACTTTTCCGCCGGCGCCCGGTTCTCCGGAGCCGCCTCACCTTTCCCG GCAGCCCGAGCAGCCGGAGCAGAGAGCCTTGGGTCCGGTTTCTATGCCAAACCTTGTACC GGAGGTGATCTTACCTGCCACGAGGCTTTCCACCCAGTGACGACGAGGATGA AGAGGGTGAGGAGTTTGTGTTAGATTATGTGGAGCACCCCGGGCACGGTTGCAGGTCTTG TCATTATCACCGGAGGAATACGGGGGACCCAGATATTATGTGTTCGCTTTGCTATATGAGG ACCTGTGGCATGTTTGTCTACAGTAAGTGAAAATTATGGGCAGTGGGTGATAGAGTGGTGG GTTTGGTGTGGTAATTTTTAATTTTTACAGTTTTGTGGTTTAAAGAATTTTGTATTGTG ATTTTTTTAAAAGGTCCTGTGTCTGAACCTGAGCCCGAGCCAGAACCGGAGCCTG CAAGACCTACCCGCCGTCCTAAAATGGCGCCTGCTATCCTGAGACGCCCGACATCACCTGT GTCTAGAGAATGCAATAGTAGTACGGATAGCTGTGACTCCGGTCCTTCTAACACACCTCCT GAGATACACCCGGTGGTCCCGCTGTGCCCCATTAAACCAGTTGCCGTGAGAGTTGGTGGG CGTCGCCAGGCTGTGGAATGTATCGAGGACTTGCTTAACGAGCCTGGGCAACCTTTGGAC TTGAGCTGTAAACGCCCCAGG
F AGCAGCCGCGGTAATTCCAGCT R R F CGGGACACTCAGCTAAGAGCATC GCACTATAAGGAACAGCTGCGCC CATTGCCGACAGGATGCA

primers were used : RAPD 1 F (5´-GGTTGGGTGAGAATTCACG-3΄) , RAPD 7 R (5´-GTAGGATGCGA--3΄) , and ERIC 2 R (5´AAGTGACTGGGGTGAGGCG-3΄). catcctacgct
Ø Methicilline Resistant Gene Ø Insulin gene Ø Tumor marker

Definition PCR primer design is the creation of short nucleotide sequences for use in amplifying a specific region of DNA.

Examples PCR primers are designed to: 1. Highly conserved DNA regions 2. Protein-coding regions with low degeneracy 3. More conserved regions that flank variable regions

Non-examples PCR primers are not designed to: 1. Repeat regions 2. Regions with secondary structure 3. Regions that can form primer-dimers 4. Regions with low G/C content

Lecture Outline ü The General Rules for PCR Primer Design ü Resources for General Purpose PCR Primer Design ü Resources for Real-Time q-PCR Primer Design ü Resources for Site-Directed Mutagenesis PCR Primer Design ü Resources for PCR Primers/Oligos Quality Analysis ü Resources for Multiplex PCR Primer Design ü Resources for Microarray Probes Design ü Resources for SNPs and Genotyping PCR Applications ü Resources for Degenerate PCR Primer Design ü Resources Methylation PCR Primer Design
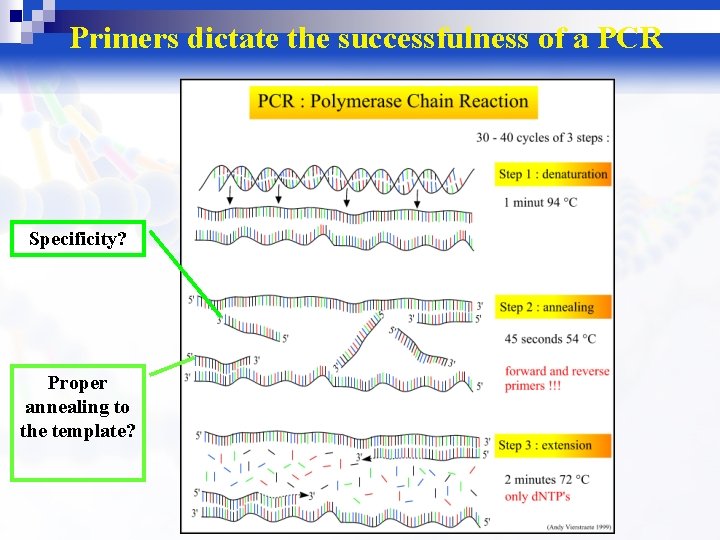
Primers dictate the successfulness of a PCR Specificity? Proper annealing to the template?

Before you design your own primers – Don’t reinvent the wheels!

Before you start designing primers – Find and use the right resources! q What are the primers for? o General purpose amplification? o SNPs detection/validation? o Methylation study? o Real-time PCR? o Microarray probes? o Degenerate PCR? o Multiplex PCR? q What do you have to begin with? Ø Single DNA/protein sequence? Ø Multiple DNA/protein sequence files? Ø Gen. Bank ID/Gene Symbol/rs. SNP ID?

After you have your primers designed – Consider a second opinion! Ø Most likely your primers can be designed by several different software Ø Different software may vary significantly in: § § § Concepts and overall approaches Designing criteria and default settings Comprehensiveness Usability Accessibility and speed v Consider a second opinion when § § You are new to such design task/application You don’t have a lot of confidence in the initial result

General rules for primer design -- Primer and amplicon length v Primer length determines the specificity and significantly affect its annealing to the template Ø Ø Too short -- low specificity, resulting in non-specific amplification Too long -- decrease the template-binding efficiency at normal annealing temperature due to the higher probability of forming secondary structures such as hairpins. v Optimal primer length Ø 18 -24 bp for general applications Ø 30 -35 bp for multiplex PCR v Optimal amplicon size Ø 300 -1000 bp for general application, avoid > 3 kb Ø 50 -150 bp for real-time PCR, avoid > 400 bp

General rules for primer design -- Melting temperature (Tm) v Tm is the temperature at which 50% of the DNA duplex dissociates to become single stranded Ø Determined by primer length, base composition and concentration. Ø Also affected by the salt concentration of the PCR reaction mix Ø Working approximation: Tm=2(A+T)+4(G+C) (suitable only for 18 mer or shorter). v Optimal melting temperature Ø 52°C-- 60°C Ø Tm above 65°C should be generally avoided because of the potential for secondary annealing. Ø Higher Tm (75°C-- 80°C) is recommended for amplifying high GC content targets. v Primer pair Tm mismatch Ø Significant primer pair Tm mismatch can lead to poor amplification Ø Desirable Tm difference < 5°C between the primer pair

General rules for primer design -- Specificity and cross homology v Specificity Ø Determined primarily by primer length as well as sequence Ø The adequacy of primer specificity is dependent on the nature of the template used in the PCR reaction. v Cross homology Ø Cross homology may become a problem when PCR template is genomic DNA or consists of mixed gene fragments. Ø Primers containing highly repetitive sequence are prone to generate non-specific amplicons when amplifying genomic DNA. v Avoid non-specific amplification Ø BLASTing PCR primers against NCBI non-redundant sequence database is a common way to avoid designing primers that may amplify non-targeted homologous regions. Ø Primers spanning intron-exon boundaries to avoid non-specific amplification of g. DNA due to c. DNA contamination. Ø Primers spanning exon-exon boundaries to avoid non-specific amplification c. DNA due to g. DNA contamination.

General rules for primer design -- GC content; repeats and runs q Primer G/C content Ø Ø Optimal G/C content: 45 -55% Common G/C content range: 40 -60% q Runs (single base stretches) Ø Ø Long runs increases mispriming (non-specific annealing) potential The maximum acceptable number of runs is 4 bp q Repeats (consecutive di-nucleotide) Ø Ø Repeats increases mispriming potential The maximum acceptable number of repeats is 4 dinucleotide

General rules for primer design -- Primer secondary structures ü Hairpins Ø Ø Ø Formed via intra-molecular interactions Negatively affect primer-template binding, leading to poor or no amplification Acceptable ΔG (free energy required to break the structure): >-2 kcal/mol for 3’end hairpin; >-3 kcal/mol for internal hairpin; ü Self-Dimer (homodimer) Ø Ø Formed by inter-molecular interactions between the two same primers Acceptable ΔG: >-5 kcal/mol for 3’end self-dimer; >-6 kcal/mol for internal self-dimer; ü Cross-Dimer (heterodimer) Ø Ø Formed by inter-molecular interactions between the sense and antisense primers Acceptable ΔG: >-5 kcal/mol for 3’end cross-dimer; >-6 kcal/mol for internal cross-dimer;

General rules for primer design -- GC clamp and max 3’ end stability v GC clamp Ø Refers to the presence of G or C within the last 4 bases from the 3’ end of primers Ø Essential for preventing mis-priming and enhancing specific primer-template binding Ø Avoid >3 G’s or C’s near the 3’ end v Max 3’end stability Ø Refers to the maximum ΔG of the 5 bases from the 3’end of primers. Ø While higher 3’end stability improves priming efficiency, too higher stability could negatively affect specificity because of 3’-terminal partial hybridization induced nonspecific extension. Ø Avoid ΔG < -9.

General rules for primer design -- Annealing temperatures and other considerations ü Ta (Annealing temperature) vs. Tm Ø Ø Ta is determined by the Tm of both primers and amplicons: optimal Ta=0. 3 x Tm(primer)+0. 7 x Tm(product)-25 General rule: Ta is 5°C lower than Tm Higher Ta enhances specific amplification but may lower yields Crucial in detecting polymorphisms ü Primer location on template Ø Ø Dictated by the purpose of the experiment For detection purpose, section towards 3’ end may be preferred. ü When using composite primers Ø Ø Initial calculations and considerations should emphasize on the template-specific part of the primers Consider nested PCR
http: //www. hsls. pitt. edu/guides/genetics/obrc http: //www. usc. edu/hsc/nml/lib-services/bioinformatics/index. html
http: //search. hsls. pitt. edu/vivisimo/cgi-bin/query-meta? input-form=molbio-simple&query=pcr+primer&v%3 Asources=OBRC&v%3 Aproject=molbio

Resources for General Purpose PCR Primer Design § § § Primer 3 Plus Primer. Z Perl. Primer Vector NTI Advantage 10
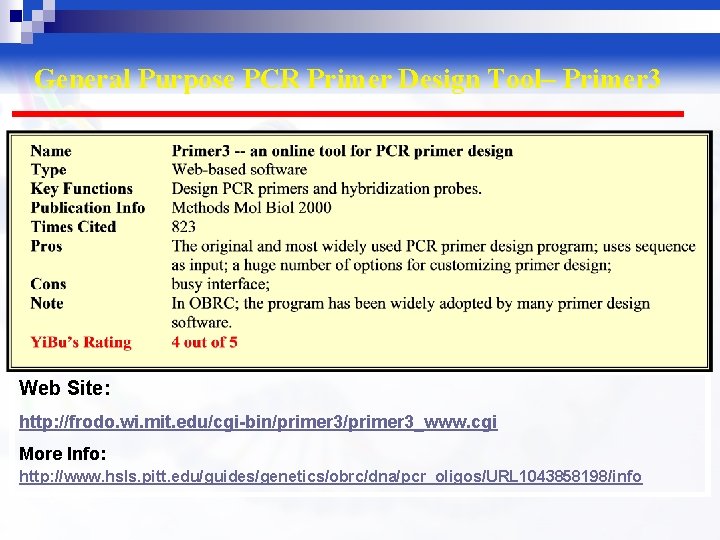
General Purpose PCR Primer Design Tool– Primer 3 Web Site: http: //frodo. wi. mit. edu/cgi-bin/primer 3_www. cgi More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1043858198/info
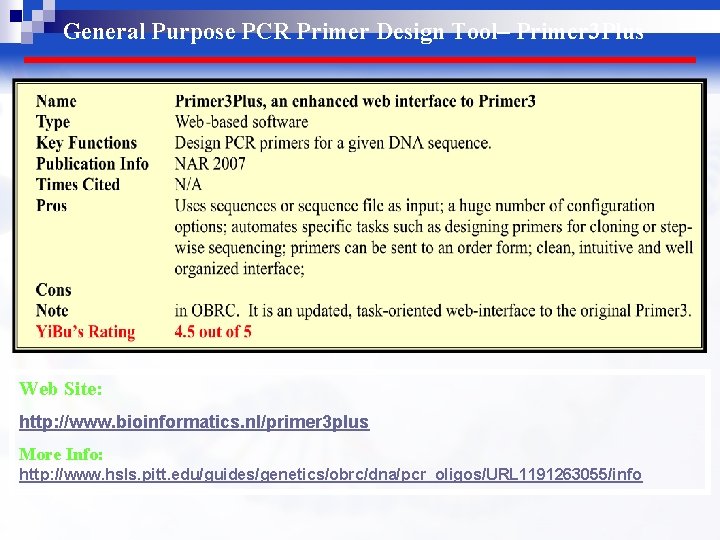
General Purpose PCR Primer Design Tool– Primer 3 Plus Web Site: http: //www. bioinformatics. nl/primer 3 plus More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1191263055/info

General Purpose PCR Primer Design Tool– Primer. Z Web Site: http: //genepipe. ngc. sinica. edu. tw/primerz/begin. Design. do More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1190992855/info

General Purpose PCR Primer Design Tool – Perl. Primer Web Site: http: //perlprimer. sourceforge. net/index. html Perl. Primer screenshots: http: //perlprimer. sourceforge. net/screenshots. html More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1167845497/info

General Purpose PCR Primer Design Tool– Vector NTI Advance 10 Web Site for NML Workshop: http: //www. usc. edu/hsc/nml/libservices/bioinformatics/vector_nti_advance_10_workshop. html More Info On Vector NTI Advance 10: http: //www. usc. edu/hsc/nml/lib-services/bioinformatics/vector_nti_advance_10. html

Primer Design Resources for Real-time PCR § § § NCBI Probe Database RTPrimer. DB Primer Bank q. Primer. Depot PCR-QPPD Perl. Primer

Public PCR Primers/Oligo Probes Repository – The NCBI Probe Database Web Site: Database Overview: http: //www. ncbi. nlm. nih. gov/genome/probe/doc/Overview. shtml Database Query Tips: http: //www. ncbi. nlm. nih. gov/genome/probe/doc/Query. Tips. shtml
http: //www. ncbi. nlm. nih. gov/sites/entrez? db=probe

Resources for real time PCR– RTPrimer. DB Web Site: http: //medgen. ugent. be/rtprimerdb/ More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1099597360/info

Resources for real time PCR– Primer Bank Web Site: http: //pga. mgh. harvard. edu/primerbank/ More Info: http: //www. ncbi. nlm. nih. gov/sites/entrez? Db=pubmed&Cmd=Show. Detail. View&Term. To. Search=14654707
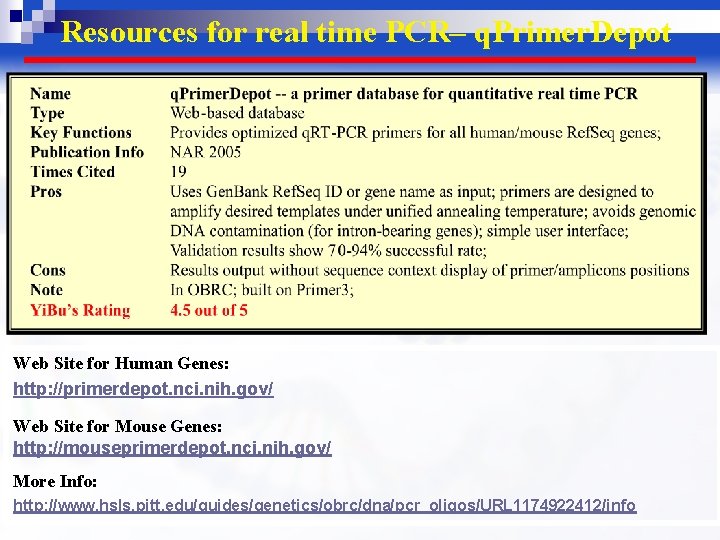
Resources for real time PCR– q. Primer. Depot Web Site for Human Genes: http: //primerdepot. nci. nih. gov/ Web Site for Mouse Genes: http: //mouseprimerdepot. nci. nih. gov/ More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1174922412/info

Resources for real time PCR– QPPD Web Site: http: //web. ncifcrf. gov/rtp/GEL/primerdb/default. asp More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1152117830/info

Resources for real time PCR– Perl. Primer Web Site: http: //perlprimer. sourceforge. net/index. html Perl. Primer screenshots: http: //perlprimer. sourceforge. net/screenshots. html More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1167845497/info
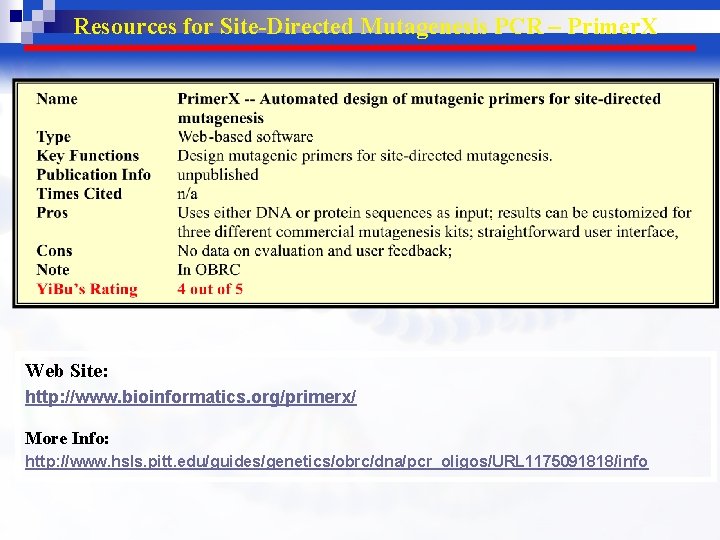
Resources for Site-Directed Mutagenesis PCR – Primer. X Web Site: http: //www. bioinformatics. org/primerx/ More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1175091818/info

Resources for PCR Primer or Oligo Analysis § § § Auto. Dimer IDT Oligo. Analyzer 3. 0 PUNS NCBI BLAST UCSC In-Silico PCR

Resources for PCR Primer or Oligo Analysis – Auto. Dimer Web Site: http: //www. cstl. nist. gov/div 831/strbase/Auto. Dimer. Homepage/Auto. Dimer. Program. Homepage. htm More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1154964478/info

Resources for PCR Primer or Oligo Analysis –IDT Oligo. Analyzer 3. 0 Web Site: http: //www. idtdna. com/analyzer/Applications/Oligo. Analyzer/ Online Instruction: http: //www. idtdna. com/Analyzer/Applications/Instructions/Default. aspx? Analyzer. Instructions=true

Resources for PCR Primer Specificity Analysis – PUNS Web Site: http: //okeylabimac. med. utoronto. ca/PUNS/ More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1175092441/info
Resources for PCR Primer Specificity Analysis – NCBI BLAST http: //www. ncbi. nlm. nih. gov/blast/Blast. cgi? PAGE=Nucleotides&PROGRAM=blastn&MEGABLAST=on&BLAST_PROGRAMS=mega. Blast&PAGE_TYPE=Blast. Search&SHOW_DEFAULTS=on
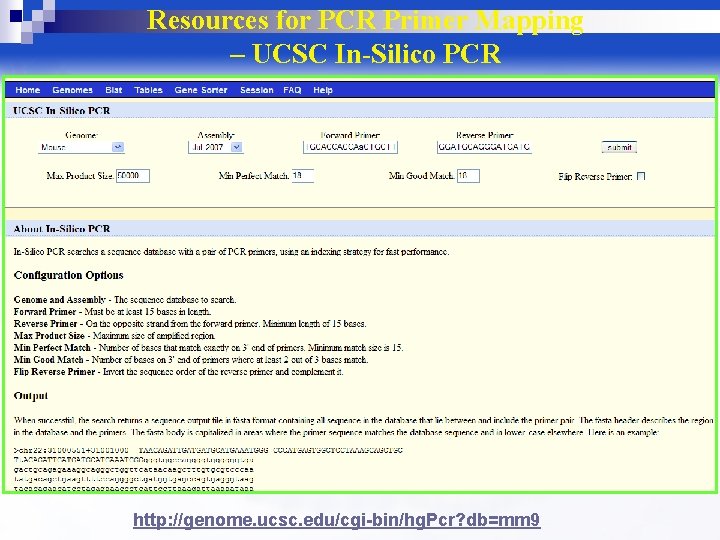
Resources for PCR Primer Mapping – UCSC In-Silico PCR http: //genome. ucsc. edu/cgi-bin/hg. Pcr? db=mm 9
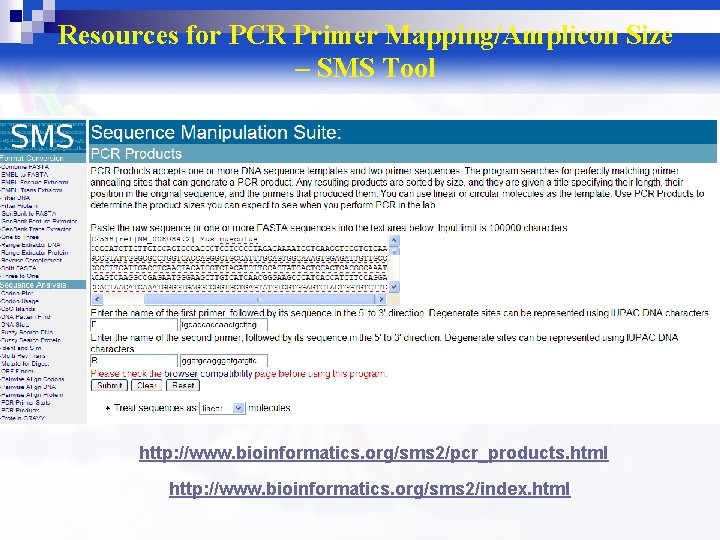
Resources for PCR Primer Mapping/Amplicon Size – SMS Tool http: //www. bioinformatics. org/sms 2/pcr_products. html http: //www. bioinformatics. org/sms 2/index. html

Primer Design Tools for Multiplex PCR Ø Multi. PLX Ø Primer. Station
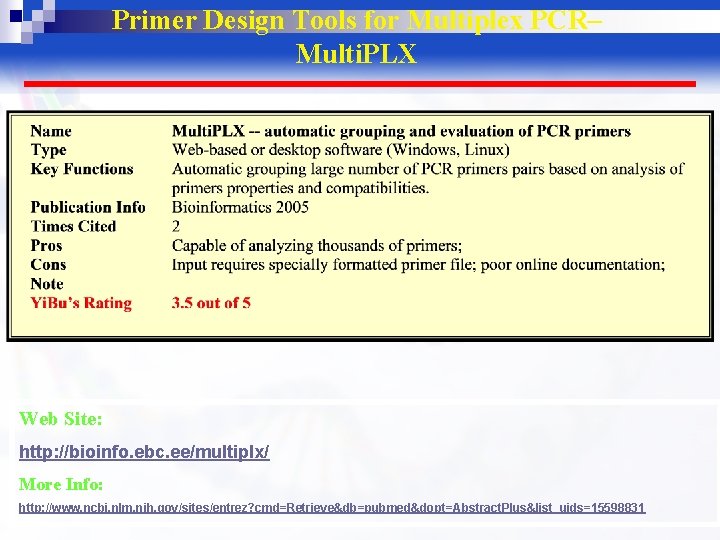
Primer Design Tools for Multiplex PCR– Multi. PLX Web Site: http: //bioinfo. ebc. ee/multiplx/ More Info: http: //www. ncbi. nlm. nih. gov/sites/entrez? cmd=Retrieve&db=pubmed&dopt=Abstract. Plus&list_uids=15598831
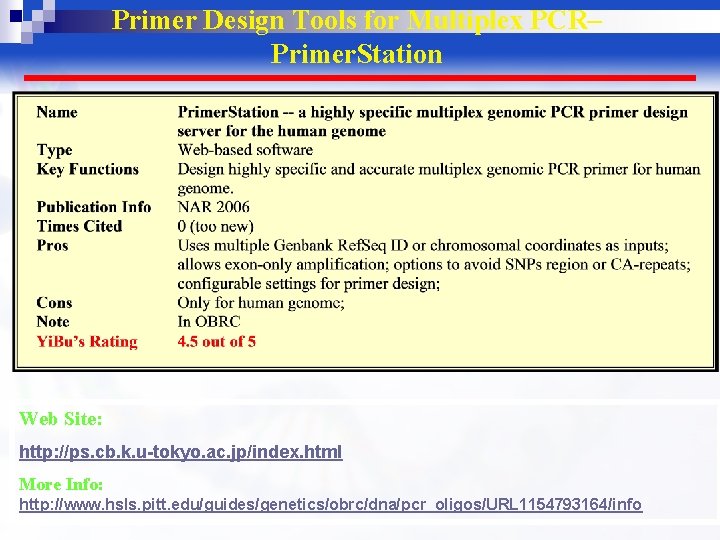
Primer Design Tools for Multiplex PCR– Primer. Station Web Site: http: //ps. cb. k. u-tokyo. ac. jp/index. html More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1154793164/info

Resources for Microarray Probe Design § § NCBI Probe Database Oligo. Wiz 2. 0 ROSO YODA

Resources for Microarray Probe Design –Oligo. Wiz 2. 0 Web Site: http: //www. cbs. dtu. dk/services/Oligo. Wiz 2/ More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/gene_expression/microarray_design_probes/URL 11187 67631/info

Resources for Microarray Probe Design –ROSO Web Site: http: //pbil. univ-lyon 1. fr/roso/Home. php More Info: http: //www. ncbi. nlm. nih. gov/sites/entrez? Db=pubmed&Cmd=Show. Detail. View&Term. To. Search=14734320

Resources for Microarray Probe Design –YODA Web Site: http: //pathport. vbi. vt. edu/YODA/ More Info: http: //bioinformatics. oxfordjournals. org/cgi/content/full/21/8/1365

PCR Primer Design Resources for SNPs and Genotyping Purposes § § NCBI Probe Database Primer. Z Mu. Plex SNPBox

Public PCR Primers/Oligo Probes Repository – The NCBI Probe Database Web Site: Database Overview: http: //www. ncbi. nlm. nih. gov/genome/probe/doc/Overview. shtml Database Query Tips: http: //www. ncbi. nlm. nih. gov/genome/probe/doc/Query. Tips. shtml

PCR Primer Design Resources for SNPs and Genotyping Purposes – Primer. Z Web Site: http: //genepipe. ngc. sinica. edu. tw/primerz/begin. Design. do More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1190992855/info
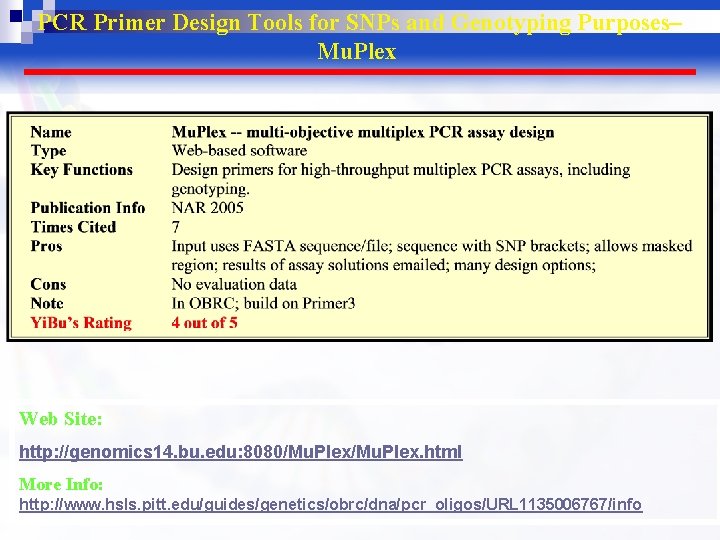
PCR Primer Design Tools for SNPs and Genotyping Purposes– Mu. Plex Web Site: http: //genomics 14. bu. edu: 8080/Mu. Plex. html More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1135006767/info
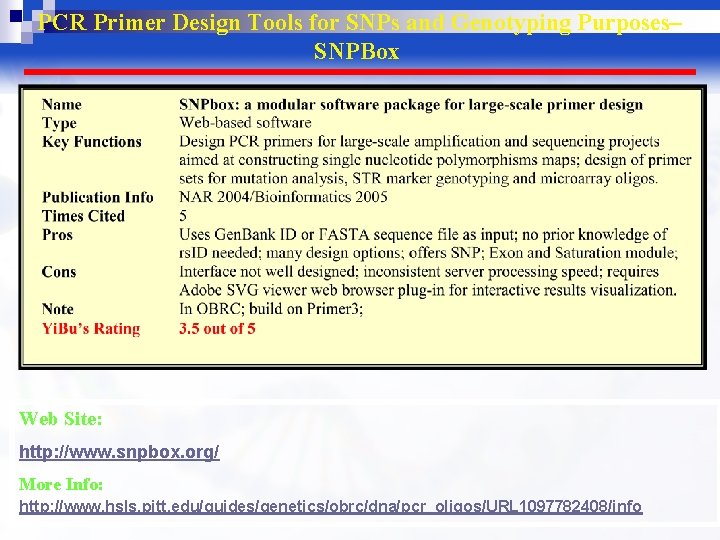
PCR Primer Design Tools for SNPs and Genotyping Purposes– SNPBox Web Site: http: //www. snpbox. org/ More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1097782408/info

Primer Design Tools for Degenerate PCR § Primaclade § Gene. Fisher 2 § CODEHOP

Primer Design Tools for Degenerate PCR– Primaclade Web Site: http: //www. umsl. edu/services/kellogg/primaclade. html More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1167846864/info

Primer Design Tools for Degenerate PCR– Gene. Fisher 2 Web Site: http: //bibiserv. techfak. uni-bielefeld. de/genefisher 2/ More Info: http: //www. ncbi. nlm. nih. gov/sites/entrez? Db=pubmed&Cmd=Show. Detail. View&Term. To. Search=8877506
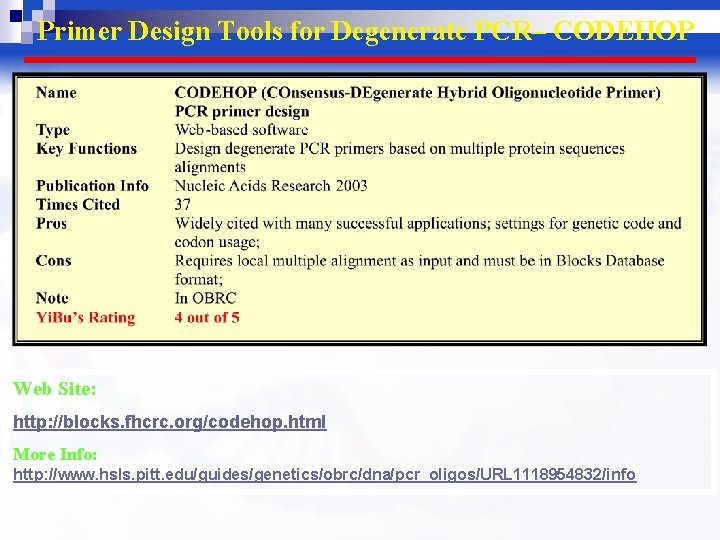
Primer Design Tools for Degenerate PCR– CODEHOP Web Site: http: //blocks. fhcrc. org/codehop. html More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1118954832/info

Primer Design Resources for Methylation PCR § Meth. Primer § meth. BLAST and meth. Primer. DB § Bi. Search § Perl. Primer
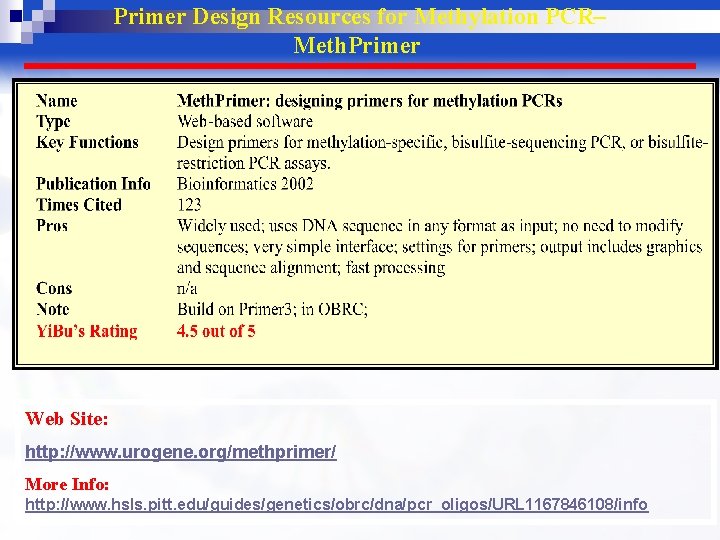
Primer Design Resources for Methylation PCR– Meth. Primer Web Site: http: //www. urogene. org/methprimer/ More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1167846108/info

Primer Design Tools for Methylation PCR– meth. BLAST and meth. Primer. DB Web Site: http: //medgen. ugent. be/methprimerdb/ meth. BLAST Web Site: http: //medgen. ugent. be/meth. BLAST/ More Info: http: //www. biomedcentral. com/1471 -2105/7/496
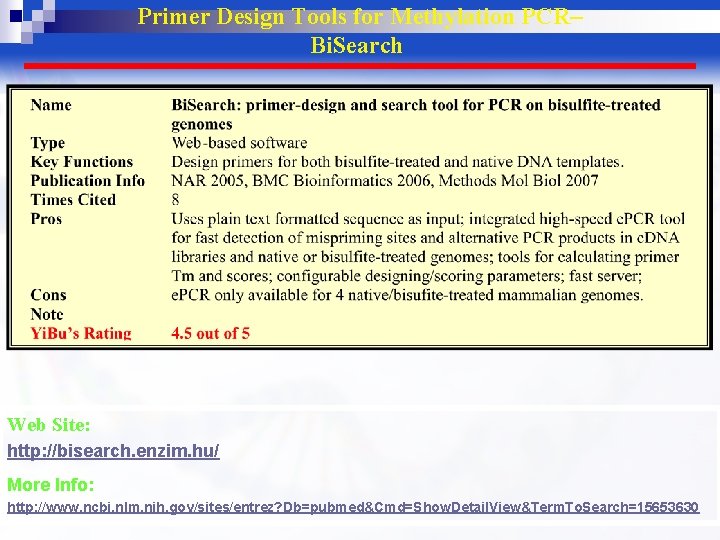
Primer Design Tools for Methylation PCR– Bi. Search Web Site: http: //bisearch. enzim. hu/ More Info: http: //www. ncbi. nlm. nih. gov/sites/entrez? Db=pubmed&Cmd=Show. Detail. View&Term. To. Search=15653630
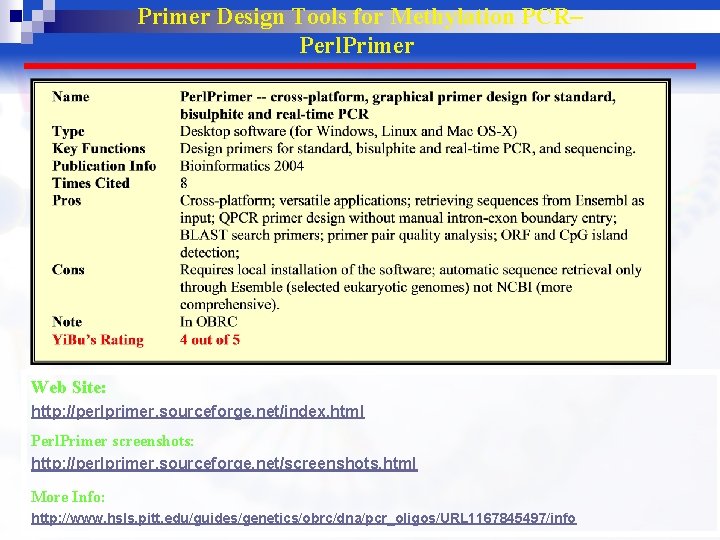
Primer Design Tools for Methylation PCR– Perl. Primer Web Site: http: //perlprimer. sourceforge. net/index. html Perl. Primer screenshots: http: //perlprimer. sourceforge. net/screenshots. html More Info: http: //www. hsls. pitt. edu/guides/genetics/obrc/dna/pcr_oligos/URL 1167845497/info
Useful web sites for design degenerate PCR primers http: //boneslab. bio. ntnu. no/degpcrshortguide. htm http: //info. med. yale. edu/mbb/koelle/protocols/protocol_degenerate_PCR. html http: //www. mcb. uct. ac. za//pcroptim. htm#Degenerate http: //www. protocol-online. org/prot/Molecular_Biology/PCR/Degenerate_PCR/ http: //cgat. ukm. my/protease/degpcr. html
- Slides: 80