Primary methods for dissociating peptides Collisionbased methods Ion
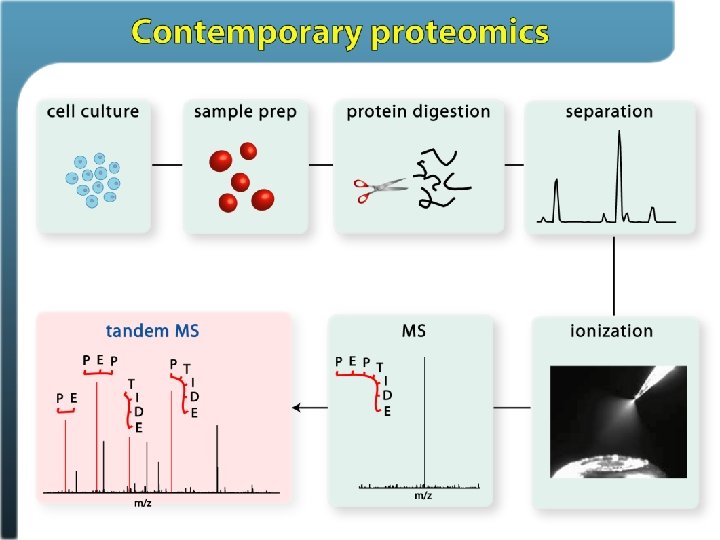
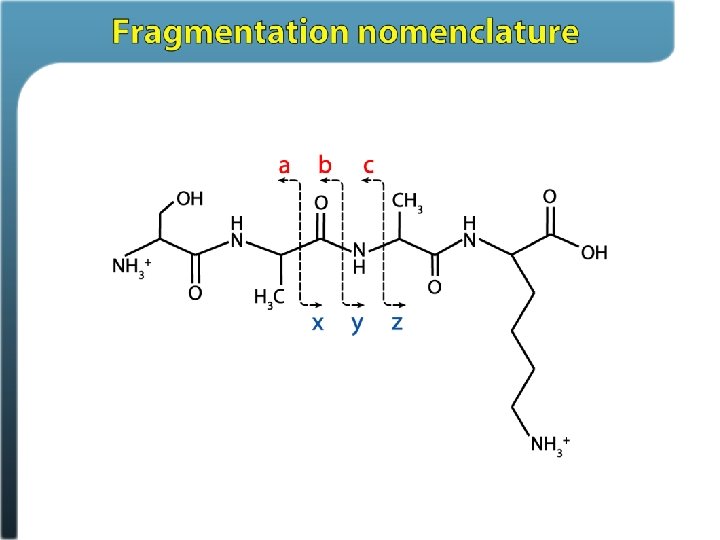
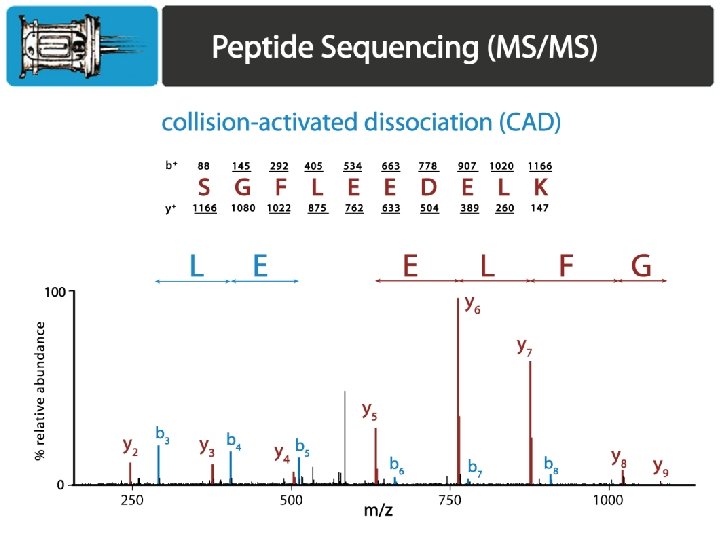

Primary methods for dissociating peptides Collision-based methods: Ion trap collisional activation – it. CAD Beam-type collisional activation – CAD aka (HCD) Electron-based methods: Electron capture dissociation (ECD) Electron transfer dissociation (ETD)
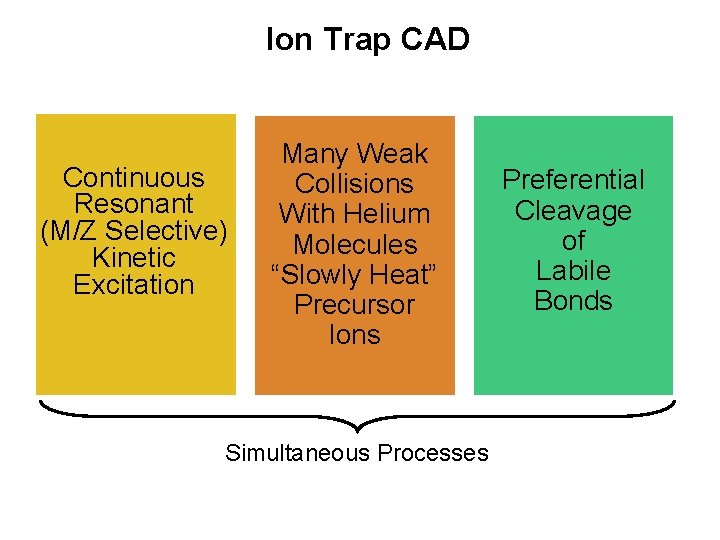
Ion Trap CAD Continuous Resonant (M/Z Selective) Kinetic Excitation Many Weak Collisions With Helium Molecules “Slowly Heat” Precursor Ions Simultaneous Processes Preferential Cleavage of Labile Bonds

Ion Trap CAD No Resonant (M/Z Selective) Kinetic Excitation Of Product Ions Many Weak Collisions With Helium Molecules “Cool” Product Ions Remain Intact Product Ions NOT Subject to Further Activation/Dissociation

RF ION TRAP ELECTRODE STRUCTURES LCQ-Type 3 D Quadrupole Trap LTQ-Type (2 D) Linear Quadrupole Trap
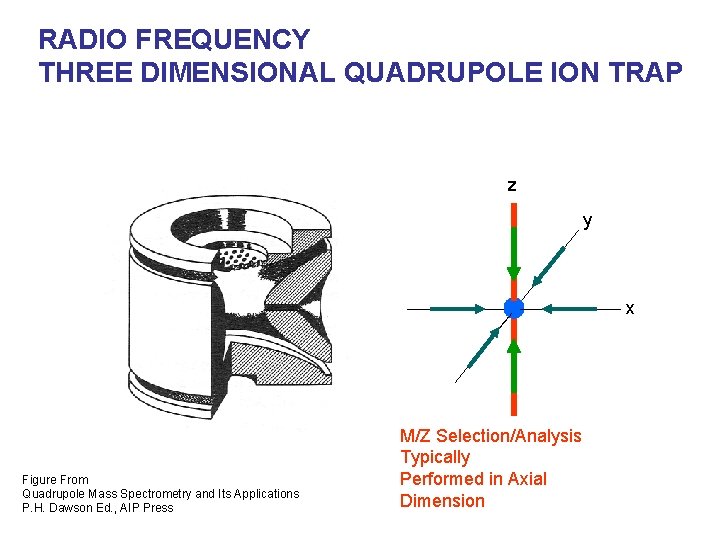
RADIO FREQUENCY THREE DIMENSIONAL QUADRUPOLE ION TRAP z y x Figure From Quadrupole Mass Spectrometry and Its Applications P. H. Dawson Ed. , AIP Press M/Z Selection/Analysis Typically Performed in Axial Dimension
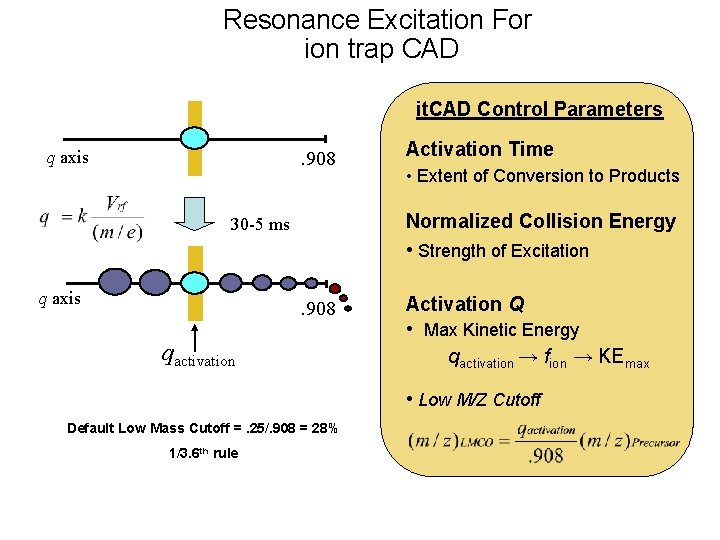
Resonance Excitation For ion trap CAD it. CAD Control Parameters q axis . 908 • Extent of Conversion to Products Normalized Collision Energy • Strength of Excitation 30 -5 ms q axis Activation Time . 908 qactivation Activation Q • Max Kinetic Energy qactivation → fion → KEmax • Low M/Z Cutoff Default Low Mass Cutoff =. 25/. 908 = 28% 1/3. 6 th rule
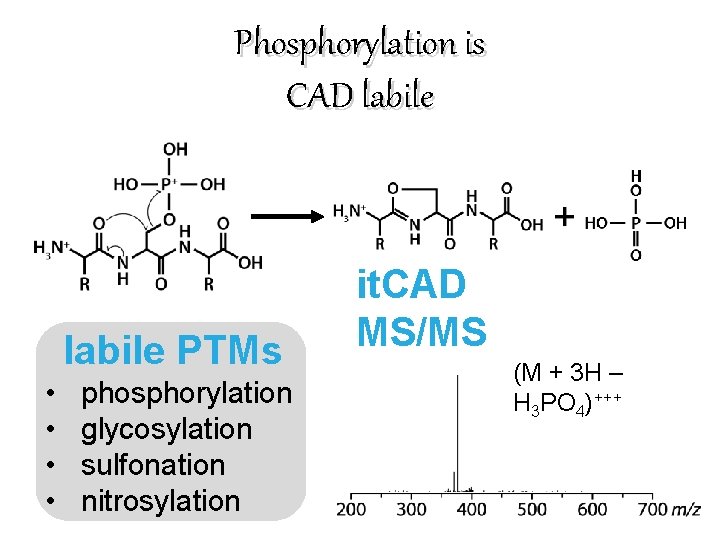
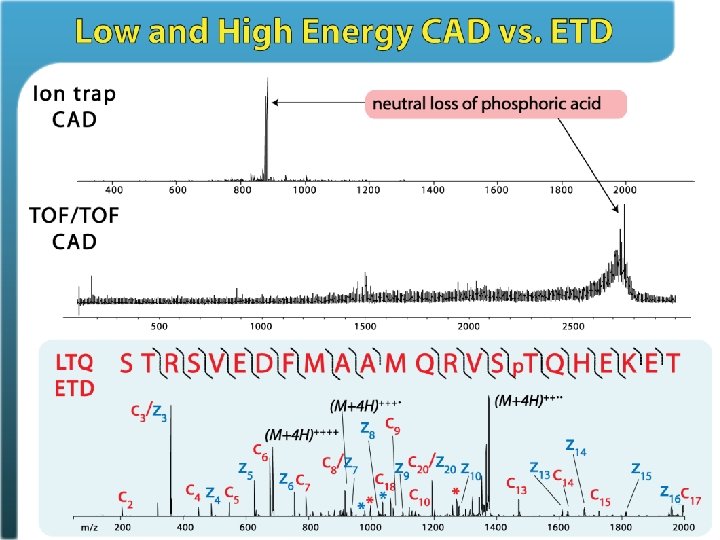
Phosphorylation is CAD labile PTMs • • phosphorylation glycosylation sulfonation nitrosylation it. CAD MS/MS (M + 3 H – H 3 PO 4)+++

Also known as Multi-Stage Activation (MSA)
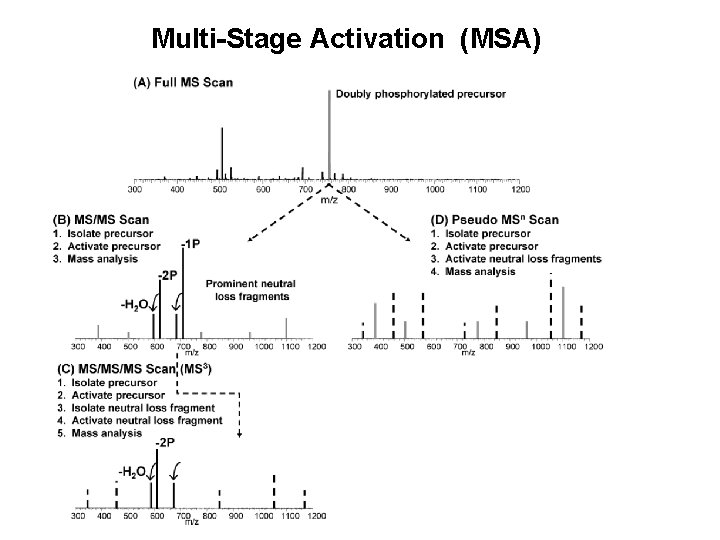
Multi-Stage Activation (MSA)
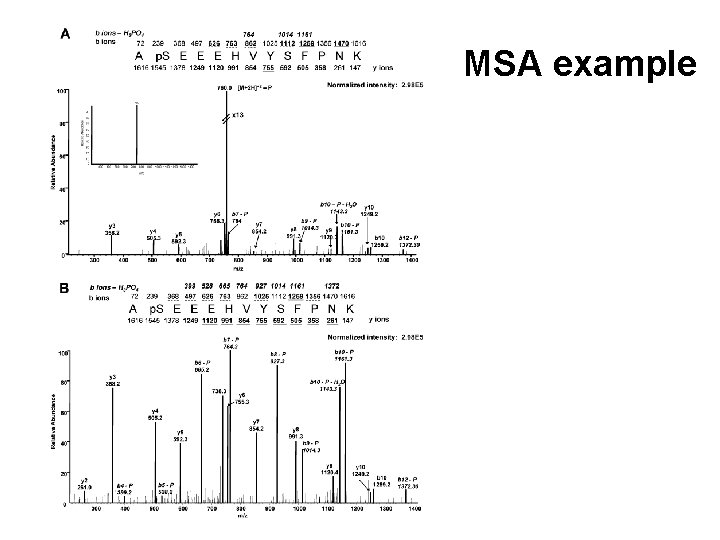
MSA example
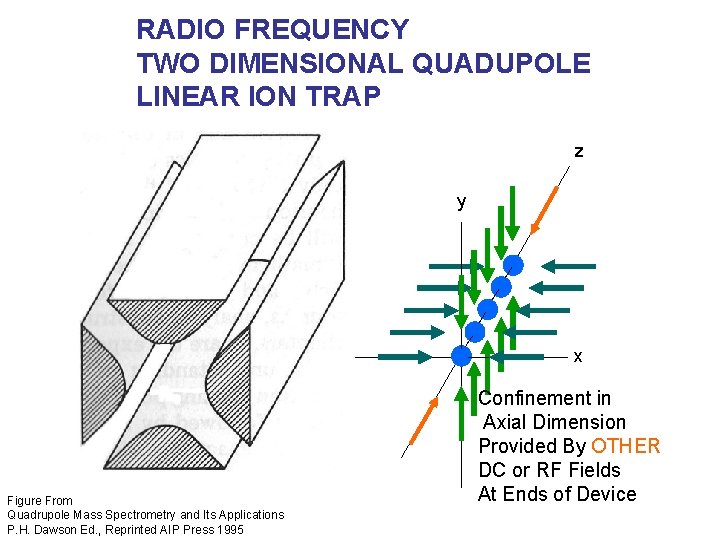
RADIO FREQUENCY TWO DIMENSIONAL QUADUPOLE LINEAR ION TRAP z y x Figure From Quadrupole Mass Spectrometry and Its Applications P. H. Dawson Ed. , Reprinted AIP Press 1995 Confinement in Axial Dimension Provided By OTHER DC or RF Fields At Ends of Device
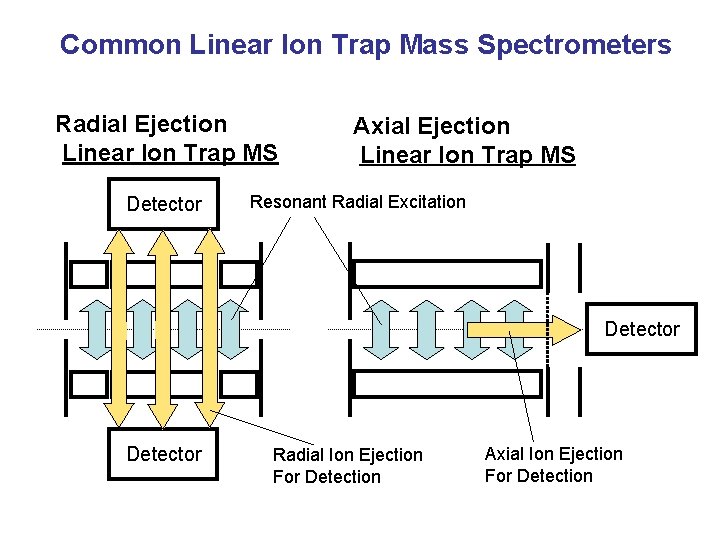
Common Linear Ion Trap Mass Spectrometers Radial Ejection Linear Ion Trap MS Detector Axial Ejection Linear Ion Trap MS Resonant Radial Excitation Detector Radial Ion Ejection For Detection Axial Ion Ejection For Detection
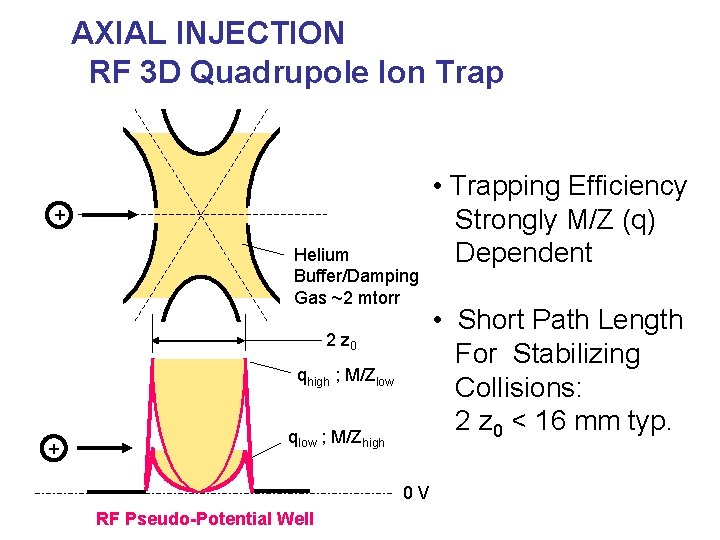
AXIAL INJECTION RF 3 D Quadrupole Ion Trap + Helium Buffer/Damping Gas ~2 mtorr 2 z 0 qhigh ; M/Zlow + qlow ; M/Zhigh 0 V RF Pseudo-Potential Well • Trapping Efficiency Strongly M/Z (q) Dependent • Short Path Length For Stabilizing Collisions: 2 z 0 < 16 mm typ.
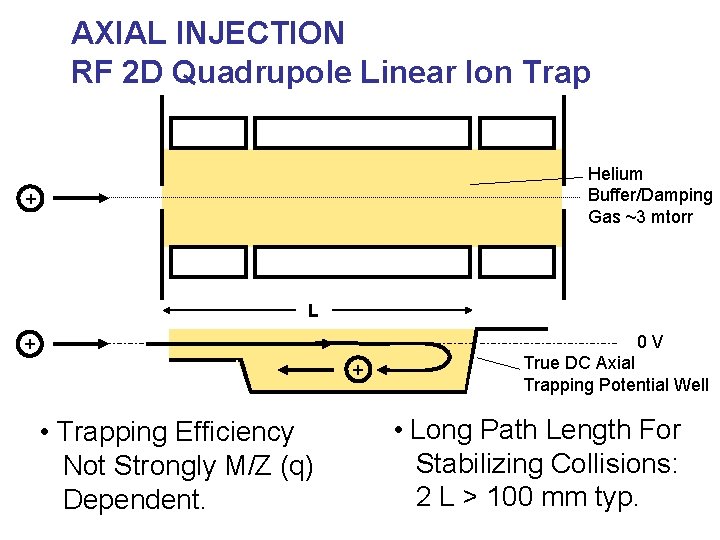
AXIAL INJECTION RF 2 D Quadrupole Linear Ion Trap Helium Buffer/Damping Gas ~3 mtorr + L 0 V + + • Trapping Efficiency Not Strongly M/Z (q) Dependent. True DC Axial Trapping Potential Well • Long Path Length For Stabilizing Collisions: 2 L > 100 mm typ.
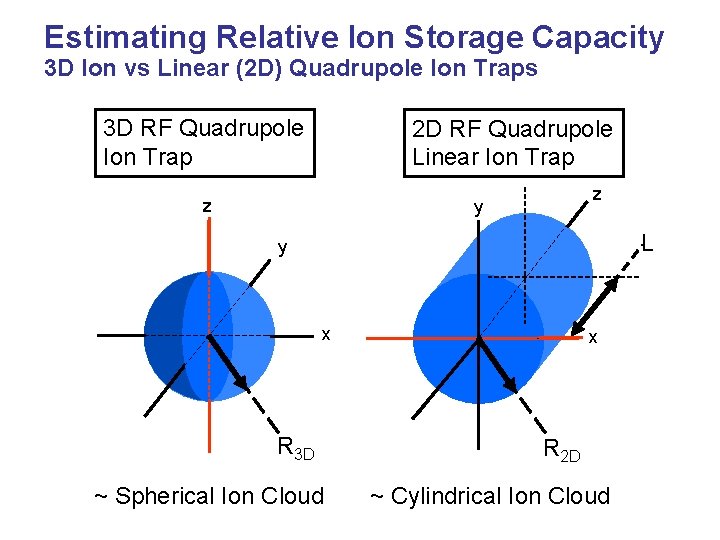
Estimating Relative Ion Storage Capacity 3 D Ion vs Linear (2 D) Quadrupole Ion Traps 3 D RF Quadrupole Ion Trap 2 D RF Quadrupole Linear Ion Trap z z y L y x R 3 D ~ Spherical Ion Cloud x R 2 D ~ Cylindrical Ion Cloud
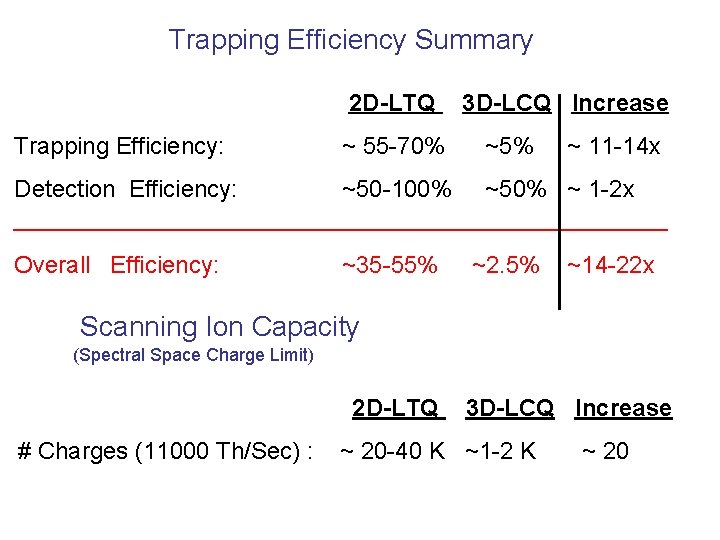
Trapping Efficiency Summary 2 D-LTQ Trapping Efficiency: ~ 55 -70% 3 D-LCQ Increase ~5% ~ 11 -14 x Detection Efficiency: ~50 -100% ~50% ~ 1 -2 x _________________________ Overall Efficiency: ~35 -55% ~2. 5% ~14 -22 x Scanning Ion Capacity (Spectral Space Charge Limit) 2 D-LTQ # Charges (11000 Th/Sec) : 3 D-LCQ Increase ~ 20 -40 K ~1 -2 K ~ 20

Introduction of the linear ion trap improved it. CAD performance for phosphopeptide identification. This is primarily because it offered ~ 20 X boost in ion capacity so that the low level fragment ions are more often detectable, even if at low abundance


Roman Zubarev Neil Kelleher Fred Mc. Lafferty
Roman Zubarev

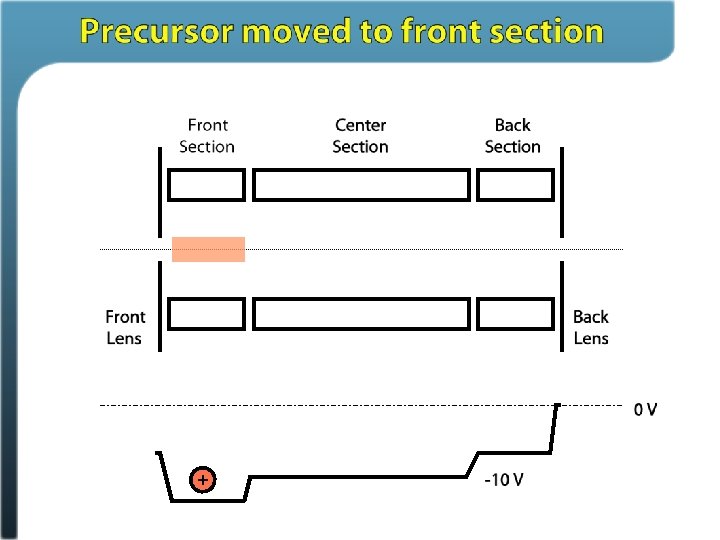
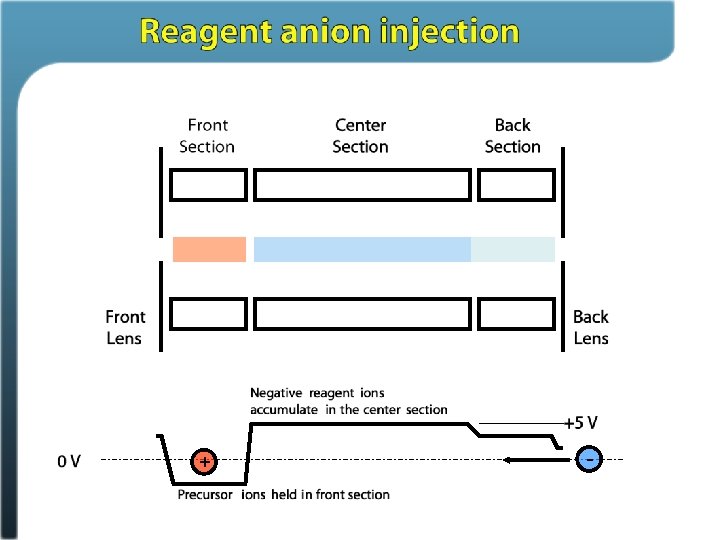
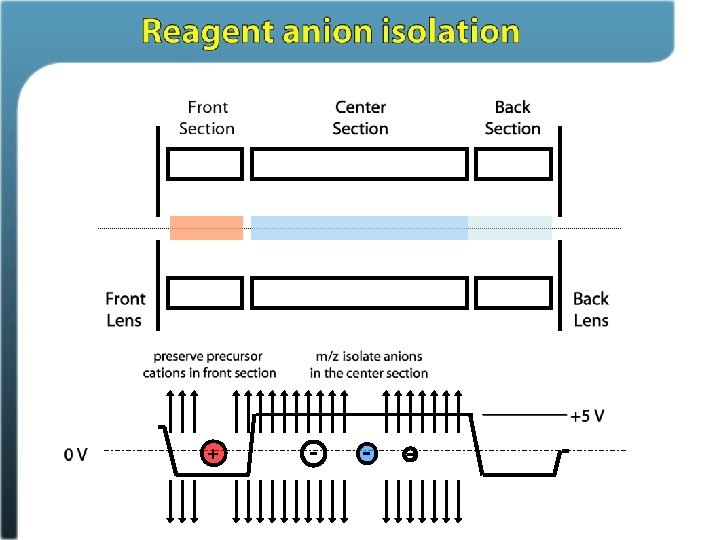
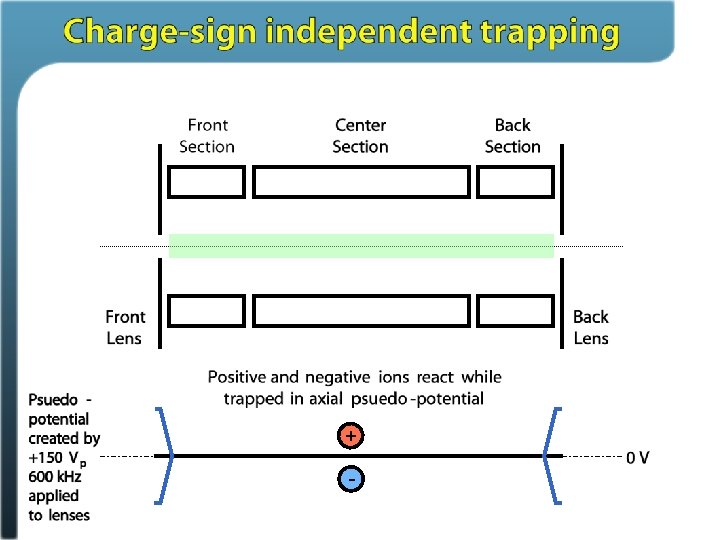
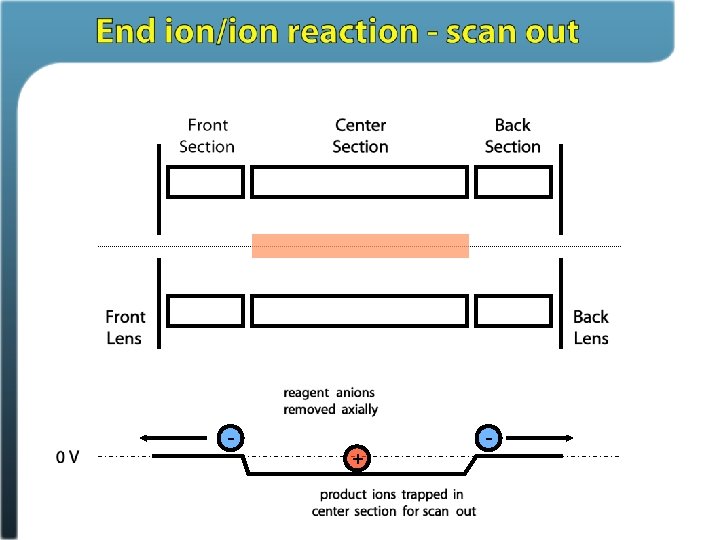
Ion/ion reactions in ion traps Proton transfer (M + 3 H)3+ + A– (M + 2 H)2+ + HA Anion attachment (M + 3 H)3+ + A– (M + 3 H + Y)2+ Electron transfer (M + 3 H)3+ + A– • (M + 3 H)2+ • + A Stephenson and Mc. Luckey, JACS, 1996 Mc. Luckey and Stephenson, Mass Spec Reviews, 1998
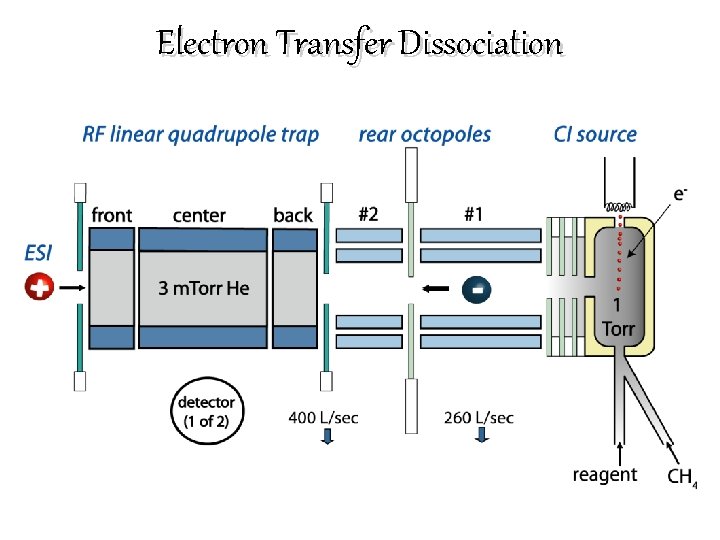
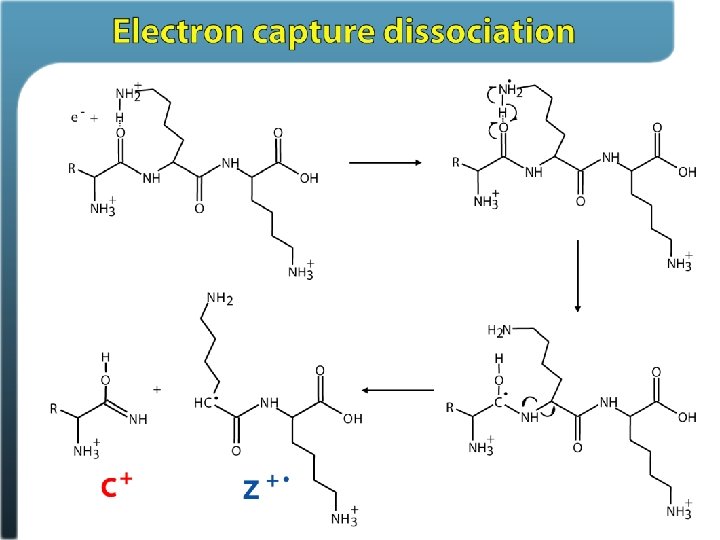
Electron Transfer Dissociation
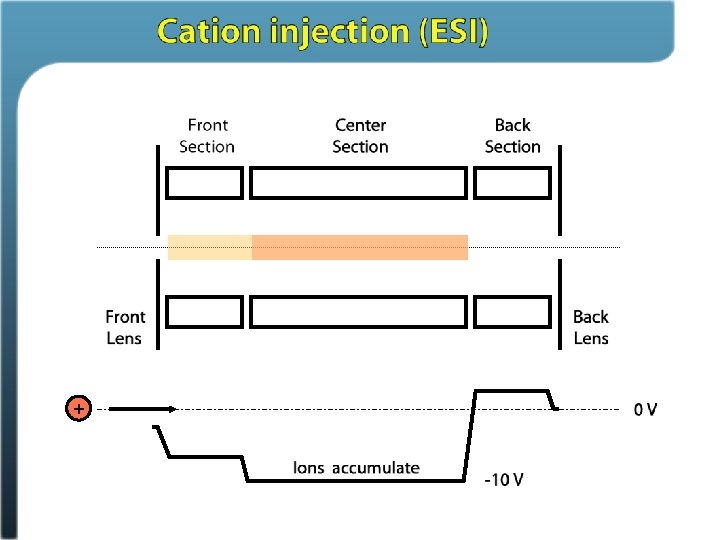
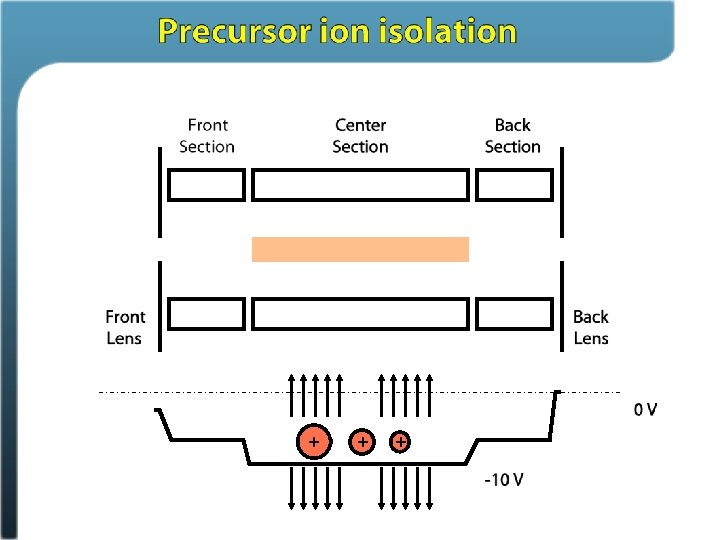
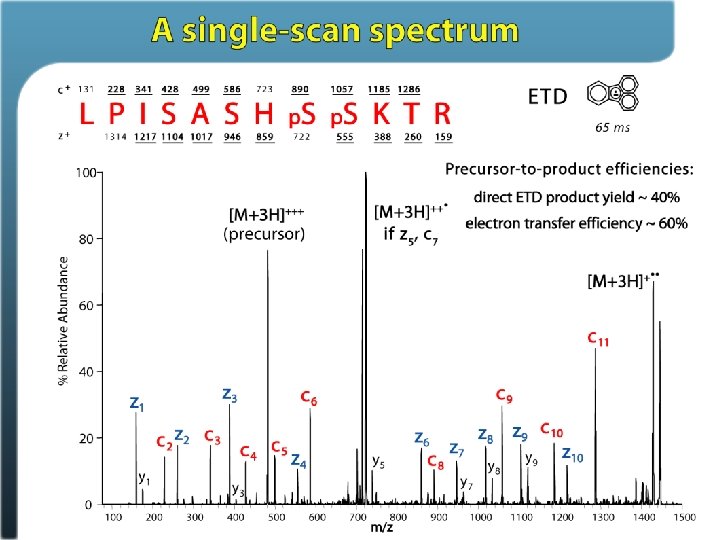
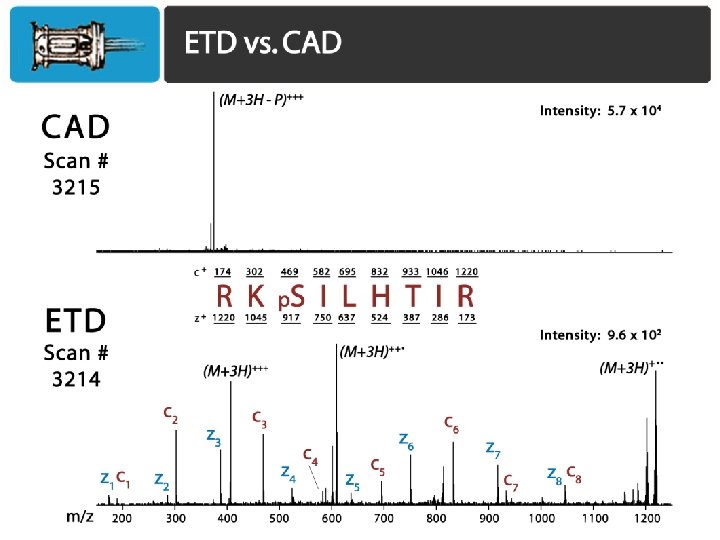
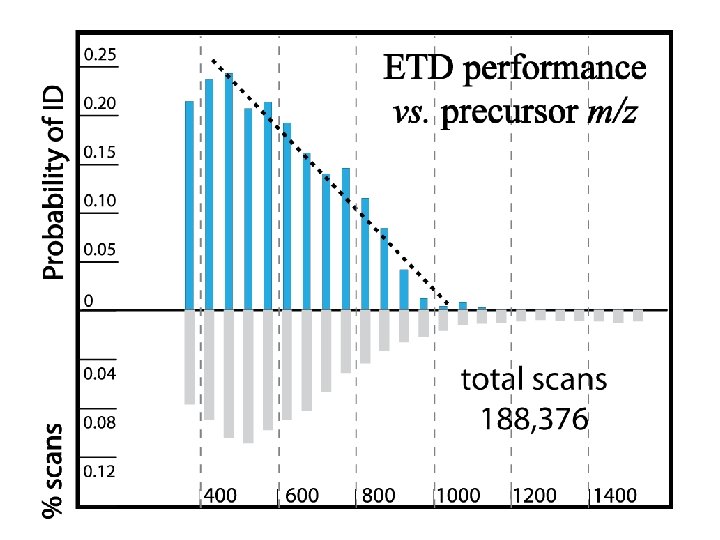
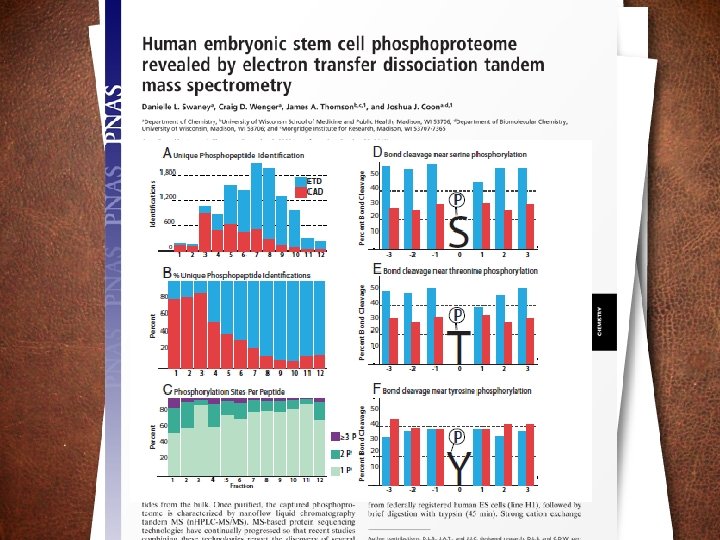
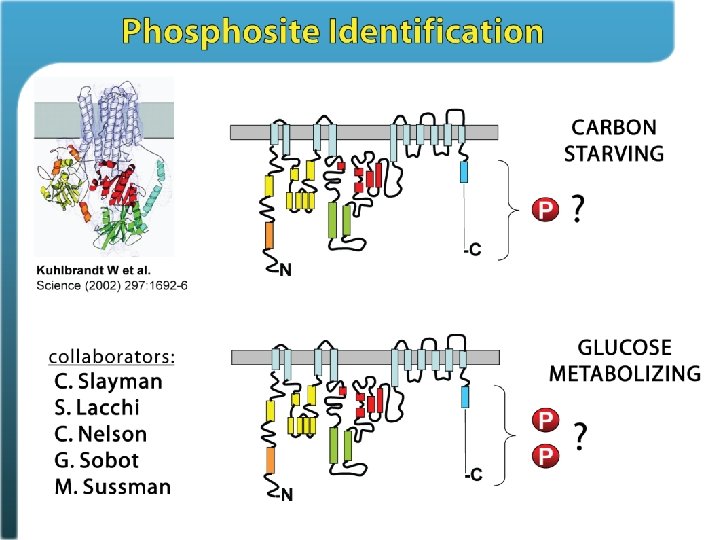
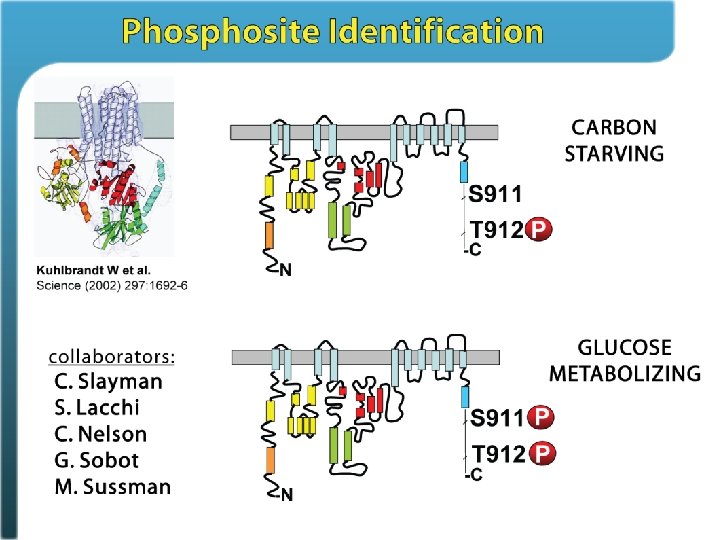
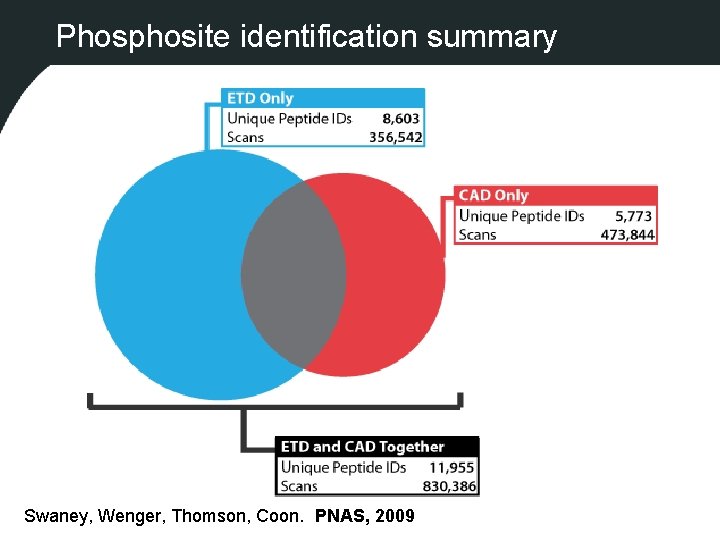
Phosphosite identification summary Swaney, Wenger, Thomson, Coon. PNAS, 2009

Probability of bond cleavage for CAD and ETD
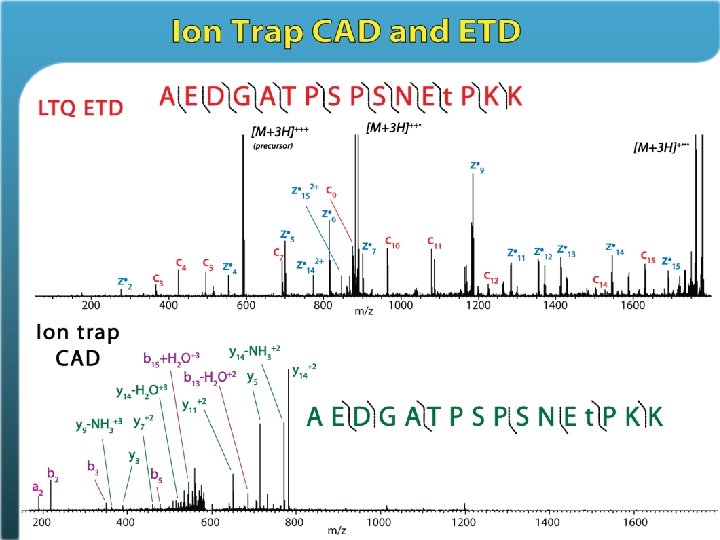
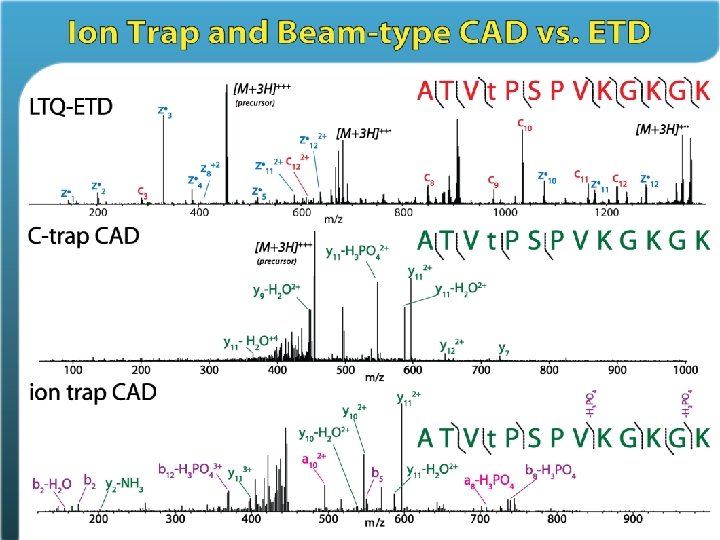
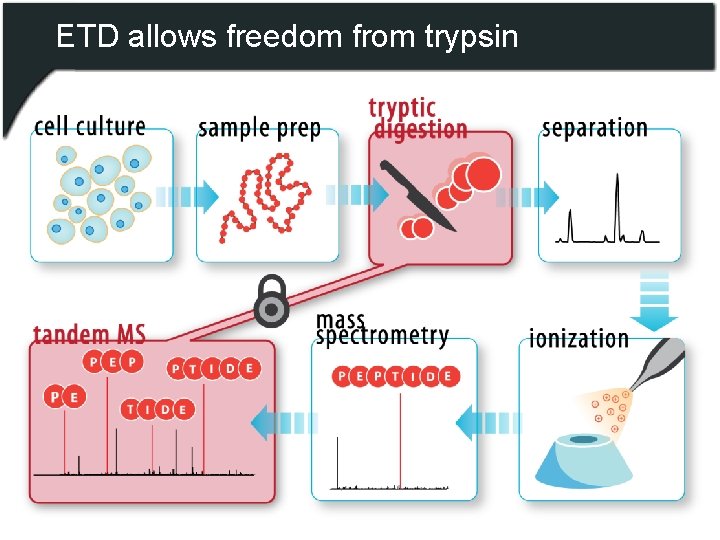
ETD allows freedom from trypsin
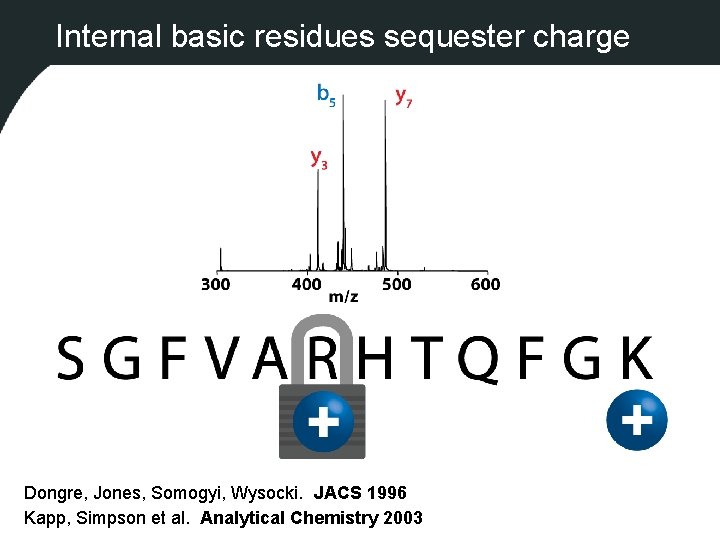
Internal basic residues sequester charge Dongre, Jones, Somogyi, Wysocki. JACS 1996 Kapp, Simpson et al. Analytical Chemistry 2003

Sequence coverage - trypsin

Sequence coverage – 5 enzymes

Collision Activated Dissociation aka HCD Kinetic Excitation Collisions Convert Kinetic Energy to Vibrational Energy Elevated Vibrational Energy Causes Bond Cleavage
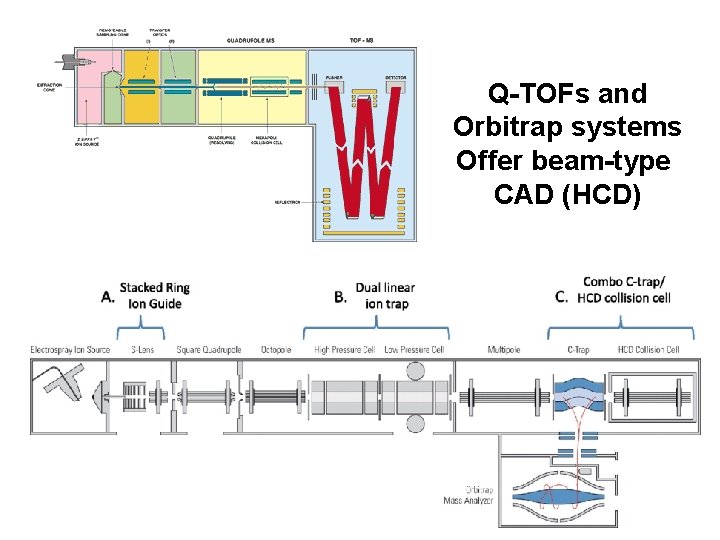
Q-TOFs and Orbitrap systems Offer beam-type CAD (HCD)
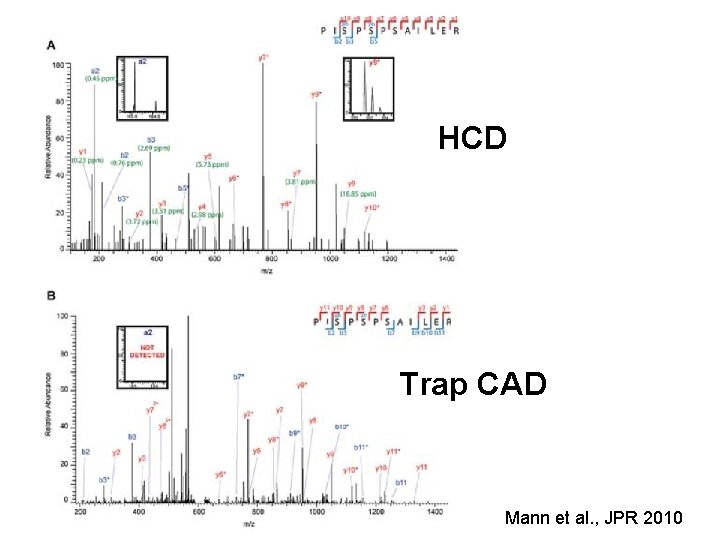
HCD Trap CAD Mann et al. , JPR 2010
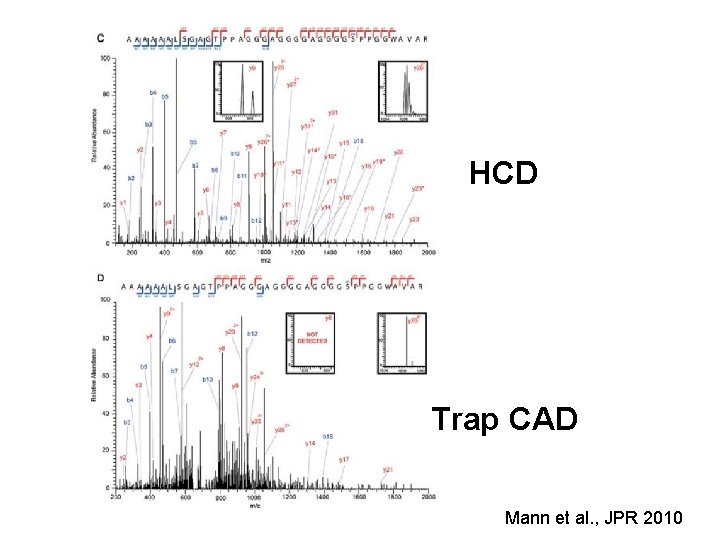
HCD Trap CAD Mann et al. , JPR 2010

Which dissociation method is best for phosphoproteomics? Depends on who you ask. Excellent results can be achieved with any of these methods The deepest coverage is achieved by using all three
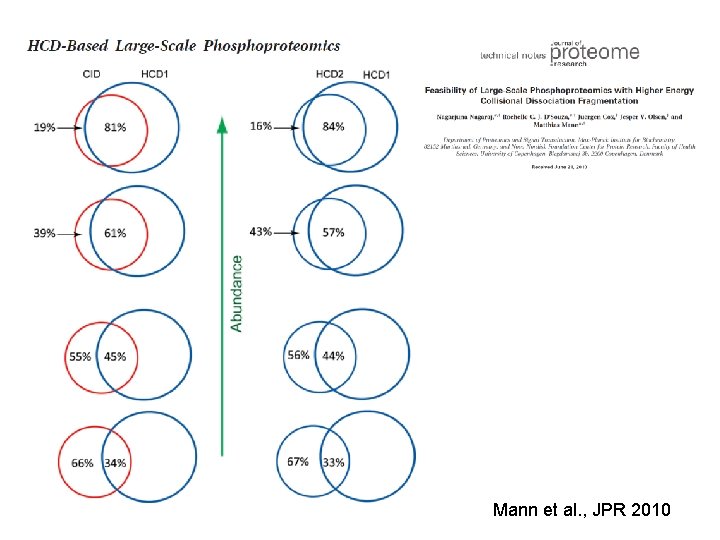
Mann et al. , JPR 2010

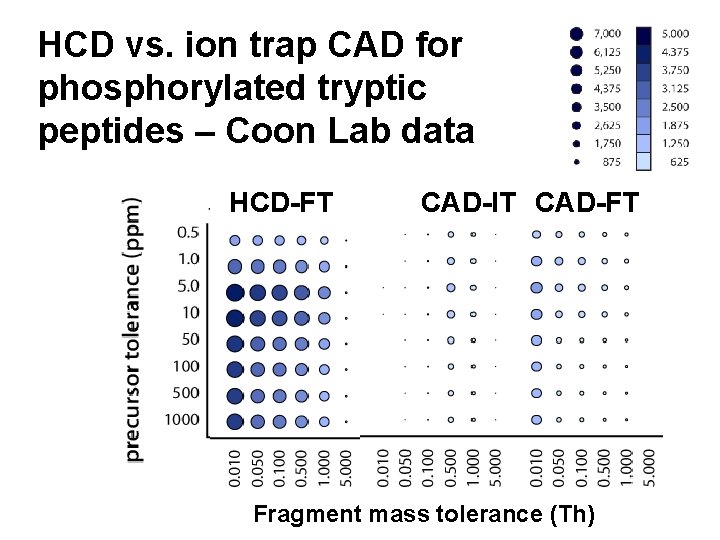
HCD vs. ion trap CAD for phosphorylated tryptic peptides – Coon Lab data HCD-FT CAD-IT CAD-FT Fragment mass tolerance (Th)

Why the varied results? I believe it’s a matter of comfort/compatibility with a specific method • Dissociation parameters can be highly optimized (e. g. , AGC, inject time, etc. ) • Database searching algorithm can make very large differences • Site localization methods
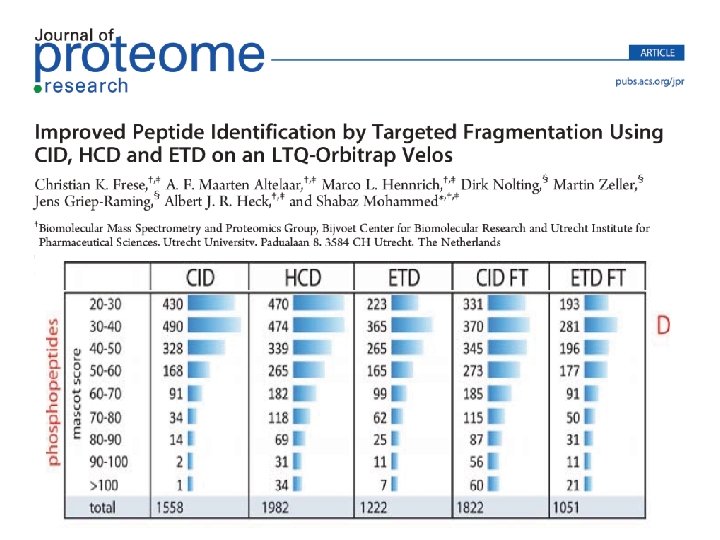
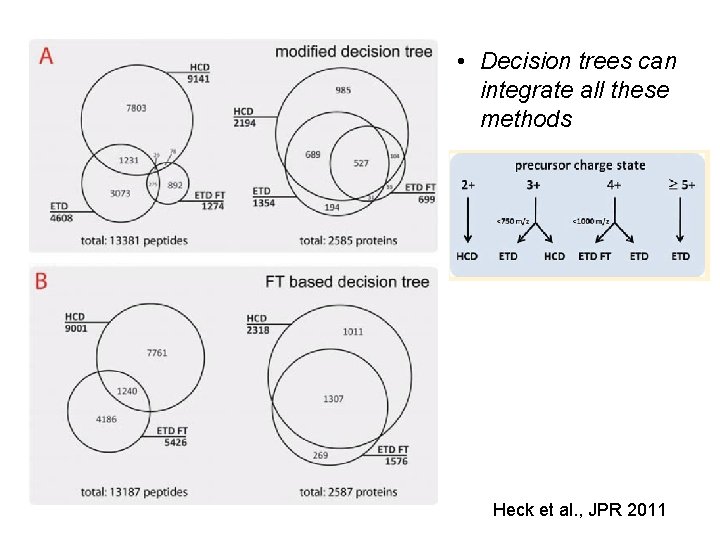
• Decision trees can integrate all these methods Heck et al. , JPR 2011
- Slides: 59