Preparing a cyanobacterial chassis for H 2 production


Preparing a cyanobacterial chassis for H 2 production: a synthetic biology approach Catarina Pacheco Cell and Applied Microbiology Group IBMC, INEB E 4. Genómica funcional e biologia sintética Encontro Nacional de Ciência - Ciência 2009 Fundação Calouste Gulbenkian, 30 th July 2009

Synthetic Biology is. . . the design and construction of new biological parts, devices and systems and the re-design of existing, natural biological systems for useful purposes. Synthetic Biology is the application of engineering concepts to biology
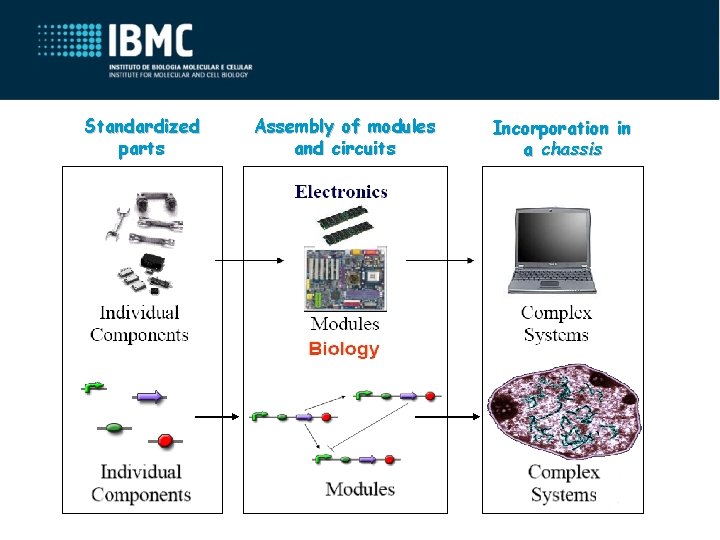
Standardized parts Assembly of modules and circuits Incorporation in a chassis
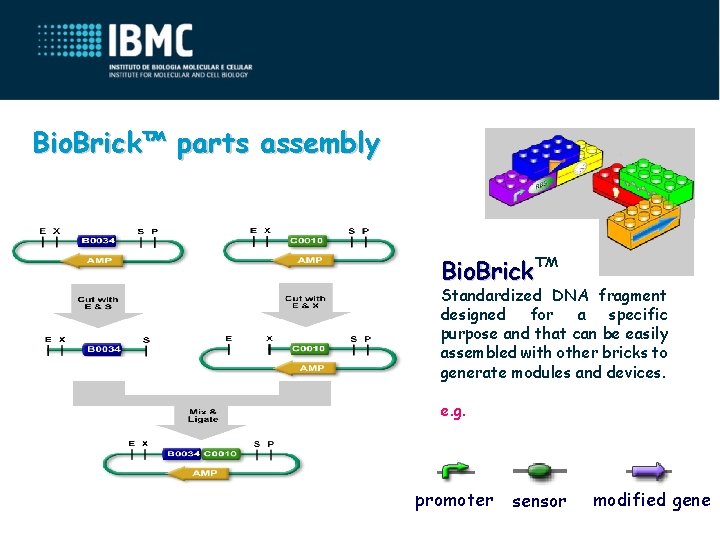
Bio. Brick™ parts assembly Bio. Brick™ Standardized DNA fragment designed for a specific purpose and that can be easily assembled with other bricks to generate modules and devices. e. g. promoter sensor modified gene
http: //partsregistry. org/

Chassis “The candidates for chassis should be well studied organisms with high throughput genomic and proteomic data available, minimalist in terms of the subset of genes that will allow retaining viability, and easy to engineer with the available molecular tools, becoming a versatile platform for multiple purpose applications” Escherichia coli Yeast Bacillus subtilis

Jan. 07 - Jan. 10 Consortium members: Instituto de Biologia Molecular e Celular (Portugal) École Polytechnique (France) Universidad Politécnica de Valencia (Spain) Uppsala Universitet (Sweden) University of Sheffield (UK) Weizmann Institute of Science (Israel) FP 6 -2005 -NEST-PATH Contract no. : 043340
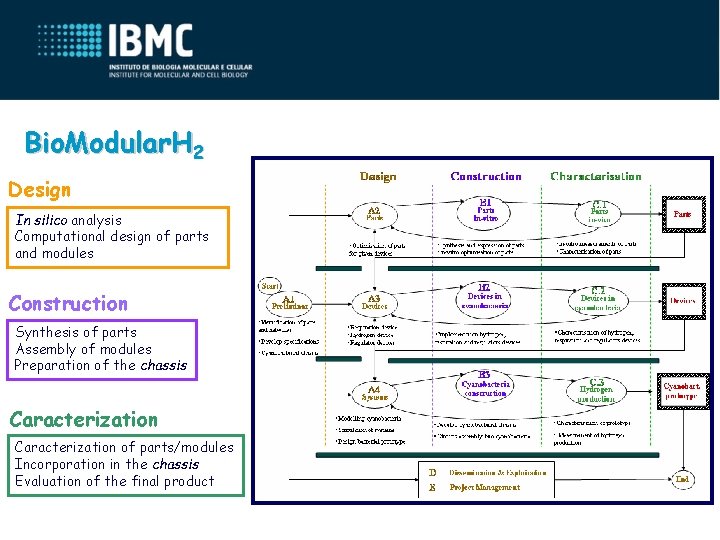
Bio. Modular. H 2 Design In silico analysis Computational design of parts and modules Construction Synthesis of parts Assembly of modules Preparation of the chassis Caracterization of parts/modules Incorporation in the chassis Evaluation of the final product
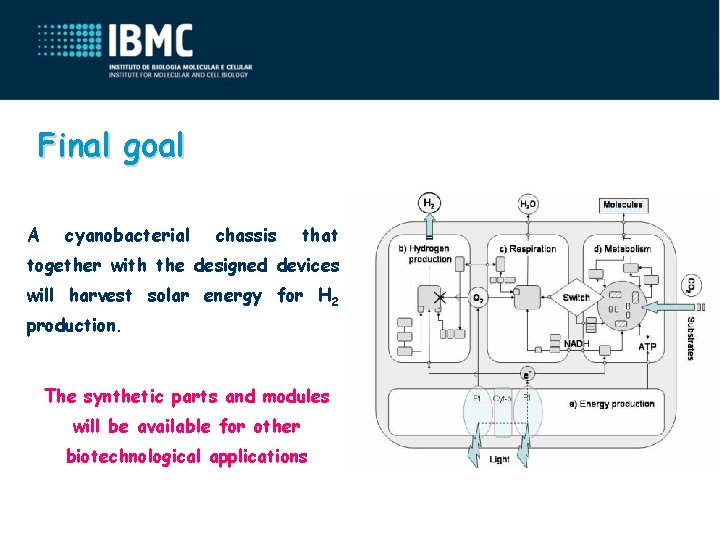
Final goal A cyanobacterial chassis that together with the designed devices will harvest solar energy for H 2 production. The synthetic parts and modules will be available for other biotechnological applications

Photoautotrophic chassis - Synechocystis sp. PCC 6803 Ø the most studied cyanobacteria Ø unicellular and non-N 2 -fixing Ø simple nutritional requirements Ø naturally transformable Ø molecular tools for manipulation available Ø small genome comprising a 3. 6 Mb genome and 7 plasmids (1 st cyano genome sequenced)
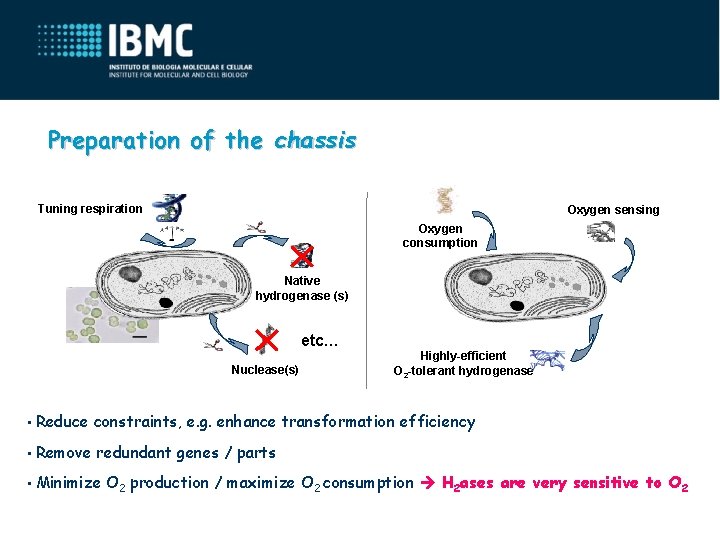
Preparation of the chassis Tuning respiration Oxygen sensing Oxygen consumption Native hydrogenase (s) etc… Nuclease(s) Highly-efficient O 2 -tolerant hydrogenase • Reduce constraints, e. g. enhance transformation efficiency • Remove redundant genes / parts • Minimize O 2 production / maximize O 2 consumption H 2 ases are very sensitive to O 2
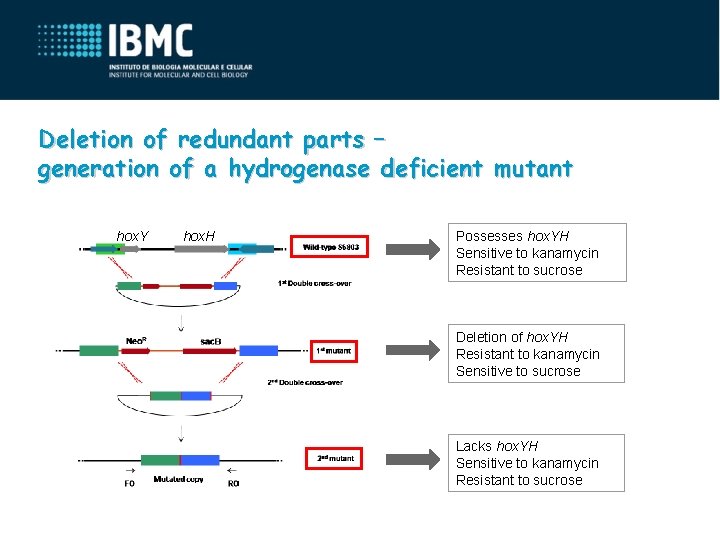
Deletion of redundant parts – generation of a hydrogenase deficient mutant hox. Y hox. H Possesses hox. YH Sensitive to kanamycin Resistant to sucrose Deletion of hox. YH Resistant to kanamycin Sensitive to sucrose Lacks hox. YH Sensitive to kanamycin Resistant to sucrose

Hydrogen Producing Device (HPD) Hyd. A 1_Fd Hydrogenase module Fe-only hydrogenase fused to ferredoxin – Chlamydomonas reinhardtii Maturation module Hyd. EF + Hyd. G – Chlamydomonas reinhardtii Homology models based on Chang et al. 2007 (Biophys J, 93: 3034 -45)

Identification of neutral sites for the insertion of synthetic modules Genes encoding proteins: - unknown or hypothetical - with maximum length of 300 a. a. - without predicted transmembrane domains (TMHMM Server v. 2. 0) - primary or secondary structure without relevant homologues - that do not interact with other proteins in two-hybrid system (Cyano. Base data) 16 potential neutral sites identified
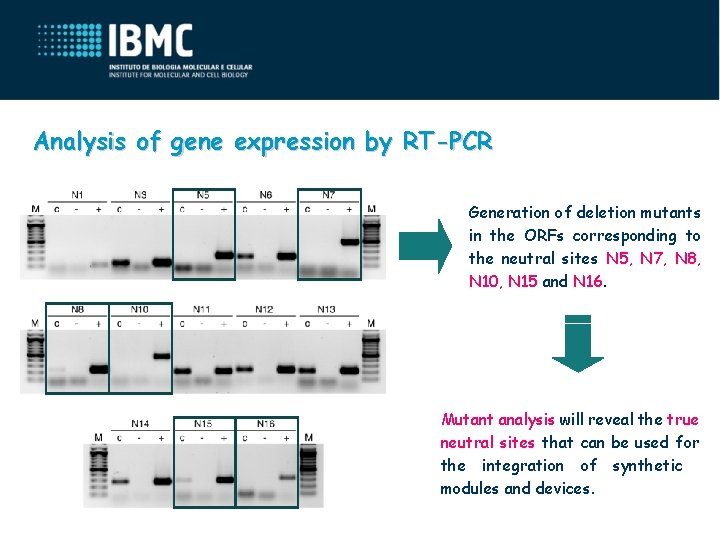
Analysis of gene expression by RT-PCR Generation of deletion mutants in the ORFs corresponding to the neutral sites N 5, N 7, N 8, N 10, N 15 and N 16. Mutant analysis will reveal the true neutral sites that can be used for the integration of synthetic modules and devices.

Design and characterization of parts for H 2 production Oxygen Consuming Device (OCD) SINGLE-protein modules O 2 H 2 O - A-type flavoprotein (ATF) – Synechocystis sp. PCC 6803 - Laccase – Escherichia coli TWO-protein module O 2 H 2 O 2 ½ O 2 + H 2 O - Glucose oxidase – Penicillium amagasakiense + Catalase – Synechocystis sp. PCC 6803
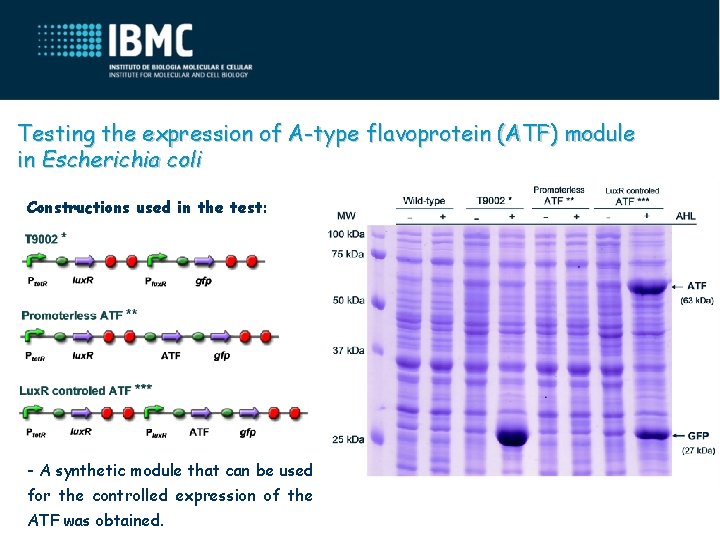
Testing the expression of A-type flavoprotein (ATF) module in Escherichia coli Constructions used in the test: - A synthetic module that can be used for the controlled expression of the ATF was obtained.

The Cellular and Applied Microbiolgy group Thank you for your attention
- Slides: 19