Practical Protein Sequence Alignment With Algebraic Dynamic Programming

Practical Protein Sequence Alignment With Algebraic Dynamic Programming Lyle Kopnicky Pac. Soft Research Group Tim Sheard, Adviser

Bioinformatics GTTAGCGTGAATCTGTACTGAG • • • DNA, RNA and proteins are strings Strings contain information Some problems • • • Determine relatedness of strands of DNA Figure out how RNA folds on itself Identify proteins in a sample

Tools for bioinformatics • • • Written in a general-purpose programming language such as C Designed to solve a narrow range of problems When problem doesn’t fit tool: • • Tweak data to fit tool – awkward, inefficient, may not fully solve problem Write new tools – time consuming, errorprone, require maintenance
The disconnect #ifndef SS strncpy(pgm_name, "gsw", MAX_FN); #else strncpy(pgm_name, "ssw", MAX_FN); #endif standard_pam("BL 50", ppst); ppst->nsq = naa; ppst->nsqx = naax; for (i=0; i<=ppst->nsqx; i++) { ppst->sq[i]=aa[i]; /* sq = aa */ ppst->hsq[i]=haa[i]; /* hsq = haa */ ppst->sqx[i]=aax[i]; /* sq = aax */ ppst->hsqx[i]=haax[i]; /* hsq = haax */ } ppst->sq[ppst->nsqx+1] = ppst->sqx[ppst->nsqx+1] = '�'; memcpy(qascii, aascii, sizeof(qascii)); /* set up the c_nt[] mapping */ ppst->c_nt[0]=0; for (i=1; i<=nnt; i++) { ppst->c_nt[i]=gc_nt[i]; ppst->c_nt[i+nnt]=gc_nt[i]+nnt; } }

The disconnect • • • General purpose languages are easily available, efficient, BUT Biologists think: amino acids, matching, classification General-purpose languages provide characters, procedures, objects Domain experts may not be expert programmers A lot of design, development and maintenance time
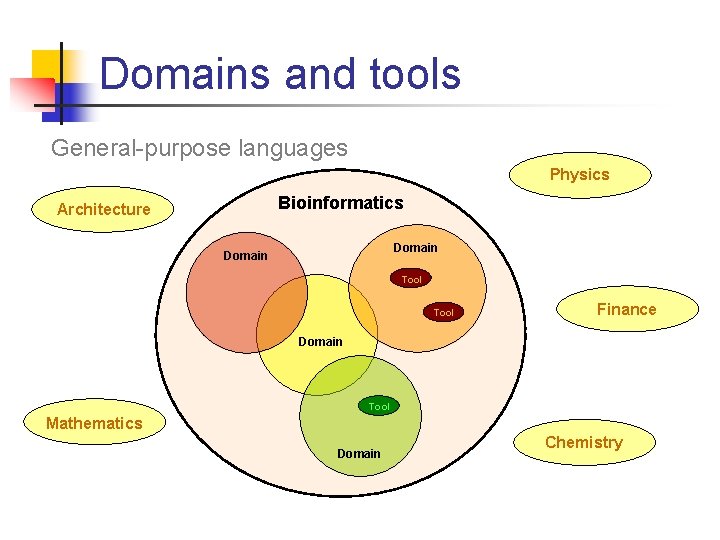
Domains and tools General-purpose languages Physics Bioinformatics Architecture Domain Tool Finance Domain Tool Mathematics Domain Chemistry

Libraries

Domain-specific languages • • • Represent a problem in the way domain experts think of it Examples: Excel, HTML, Matlab General enough to capture a domain, specific enough to reduce design time Small change in requirements = small change in program Easy to answer “what-if” questions Can be implemented efficiently using known techniques

Collaboration with OHSU lab • • Dr. Srinivasa Nagalla’s laboratory Goals • • Gain domain knowledge Discover goals of biologists Potential users of DSL Began with protein identification problem
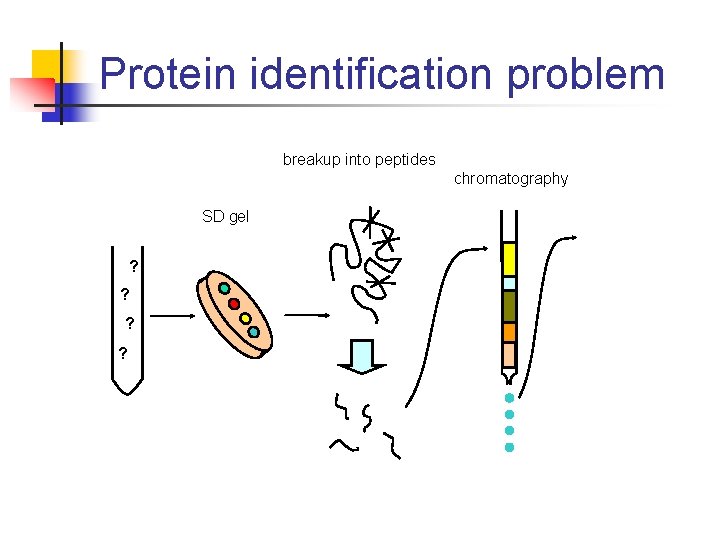
Protein identification problem breakup into peptides chromatography SD gel ? ?
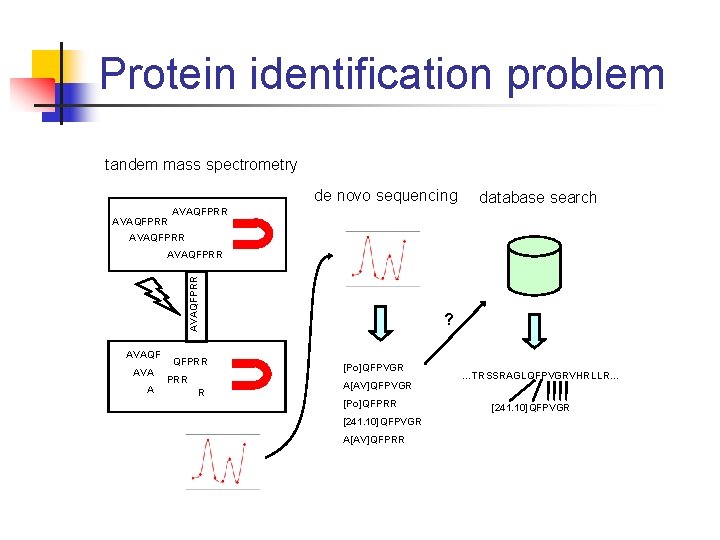
Protein identification problem tandem mass spectrometry de novo sequencing AVAQFPRR database search AVAQFPRR AVAQF AVA A QFPRR R ? [Po]QFPVGR A[AV]QFPVGR [Po]QFPRR [241. 10]QFPVGR A[AV]QFPRR …TRSSRAGLQFPVGRVHRLLR… [241. 10]QFPVGR

Database search • • Not an exact match Mutations – substitutions, insertions, deletions Modifications – amino acids altered by another molecule De novo sequencer outputs a list of ambiguous queries, e. g. [229. 07]Hh. Ny. G[PS][198. 1]QHADD[ep]VD[Rz]R unidentified mass unordered pair

Sequence alignment • • • Find the best match between query string and target (database) string Each match is also called an alignment Alignments are scored Target string WTABRRFCWGYPD KWGGSCASPNE F WT PDPYK Query string

Alignment tools • Smith-Waterman dynamic programming algorithm most common • • O((n+m)2), where n and m are length of strings Run on every query-target pair, and data sets are large Requires precise query string FASTA, BLAST address speed problem • • First look for runs of exact matches Align on localized area

Fitting queries into FASTA • • Generate all possible exact queries Could be dozens of possibilities [241. 10]NEM[NP]YR NQ QN GG GAN GNA GQG GGQ AGGG • • NP PN Each one must be aligned – takes time Output must be converted back to original query string

Algebraic Dynamic Programming • • • Robert Giegerich, 2000 Generic approach to solving dynamic programming problems using parsing Domain-specific embedded language in Haskell Ambiguous queries can be represented directly as a grammar Searching with an ambiguous query slower, but only one search instead of dozens Still takes O((n+m)2) time
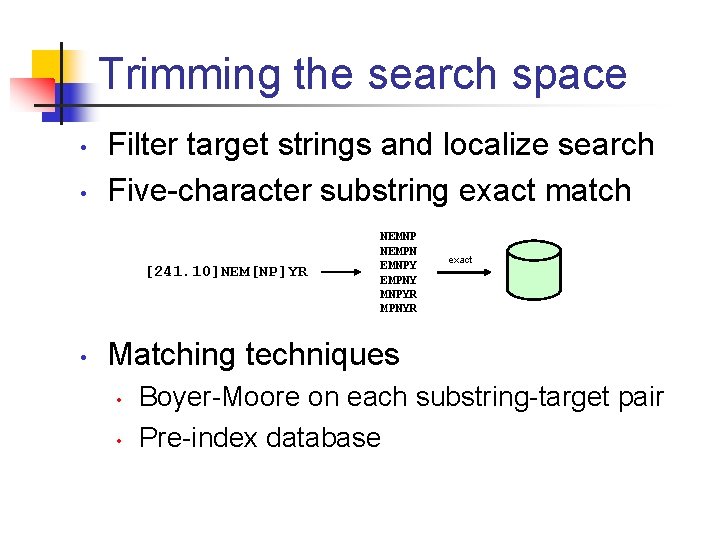
Trimming the search space • • Filter target strings and localize search Five-character substring exact match [241. 10]NEM[NP]YR • NEMNP NEMPN EMNPY EMPNY MNPYR MPNYR exact Matching techniques • • Boyer-Moore on each substring-target pair Pre-index database

Boyer-Moore • • • Finds an exact occurrence of one string inside another Doesn’t check every position – knows when to skip ahead Can run in sublinear time SYNSNTLNNDIMLIKLKSAASLN x SAASL
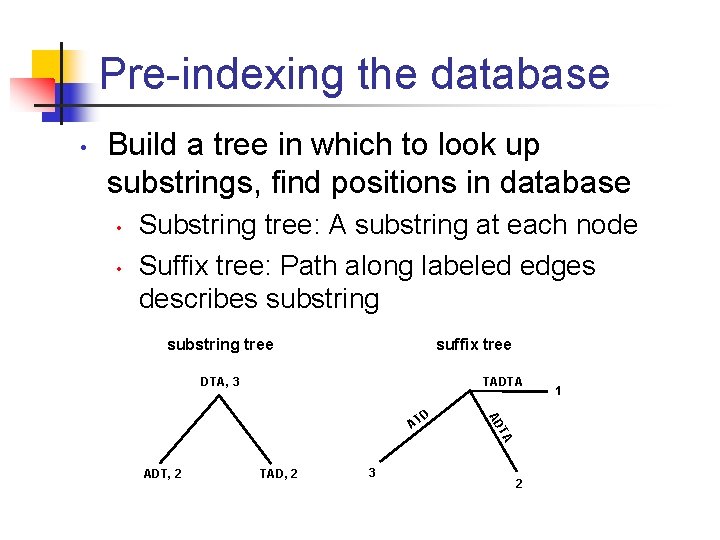
Pre-indexing the database • Build a tree in which to look up substrings, find positions in database • • Substring tree: A substring at each node Suffix tree: Path along labeled edges describes substring tree suffix tree DTA, 3 TADTA TAD, 2 3 TA ADT, 2 AD D AT 2 1
![Sample output Query string: [229. 07]Hh. Ny. G[PS][198. 1]QHADD[EP]VD[Rz]R: {21} Target string: >MK 14_HUMAN Sample output Query string: [229. 07]Hh. Ny. G[PS][198. 1]QHADD[EP]VD[Rz]R: {21} Target string: >MK 14_HUMAN](http://slidetodoc.com/presentation_image_h2/92108f4000f687b1d0d2312dcd0d78e7/image-20.jpg)
Sample output Query string: [229. 07]Hh. Ny. G[PS][198. 1]QHADD[EP]VD[Rz]R: {21} Target string: >MK 14_HUMAN (Q 16539) Mitogen-activated protein kinase 14 MSQERPTFYRQELNKTIWEVPERYQNLSPVGSGAYGSVCAAFDTKTGLRVAVKKLSRPFQSIIH AKRTYRELRLLKHMKHENVIGLLDVFTPARSLEEFNDVYLVTHLMGADLNNIVKCQKLTDDHVQ FLIYQILRGLKYIHSADIIHRDLKPSNLAVNEDCELKILDFGLARHTDDEMTGYVATRWYRAPE IMLNWMHYNQTVDIWSVGCIMAELLTGRTLFPGTDHIDQLKLILRLVGTPGAELLKKISSESAR NYIQSLTQMPKMNFANVFIGANPLAVDLLEKMLVLDSDKRITAAQALAHAYFAQYHDPDDEPVA DPYDQSFESRDLLIDEWKSLTYDEVISFVPPPLDQEEMES: {360} Target range: 290 -326 LDSDKRITAAQALAHA YFAQYHDPDDEPVADPYDQSF : |Xvvv. XX: |^|X^||//|^|XXX -Hh. Ny. GPS-Q HA DDEPV DRz. R Score: 31 Time: 0. 045 secs
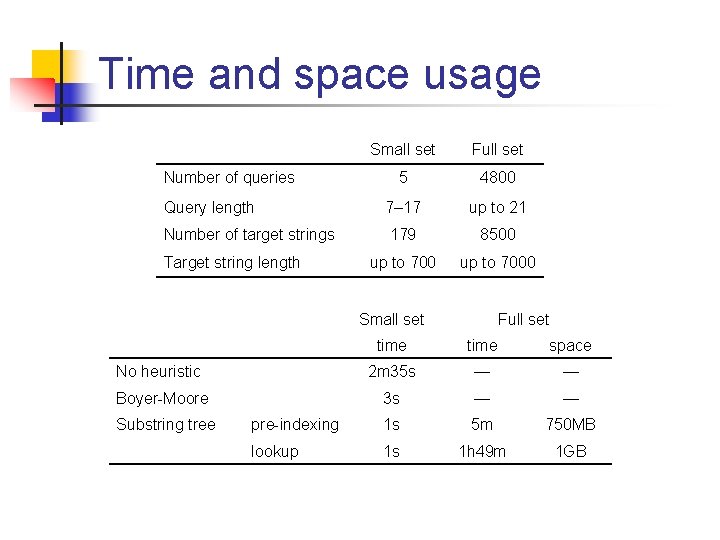
Time and space usage Small set Full set 5 4800 Query length 7– 17 up to 21 Number of target strings 179 8500 up to 7000 Number of queries Target string length Small set Full set time space No heuristic 2 m 35 s — — Boyer-Moore 3 s — — pre-indexing 1 s 5 m 750 MB lookup 1 s 1 h 49 m 1 GB Substring tree

Smith-Waterman (1981) • • Local alignment problem Dynamic programming algorithm Pathways through table represent alignments Entry represents best score of an alignment starting here, ending anywhere
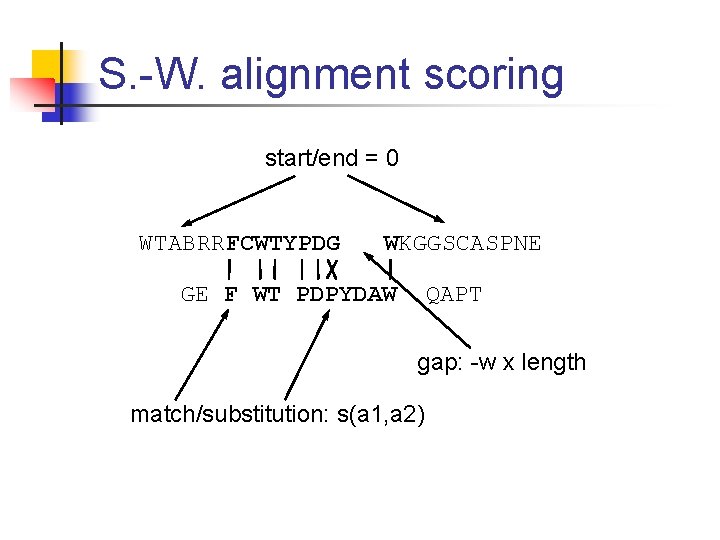
S. -W. alignment scoring start/end = 0 WTABRRFCWTYPDG WKGGSCASPNE GE F WT PDPYDAW QAPT gap: -w x length match/substitution: s(a 1, a 2)
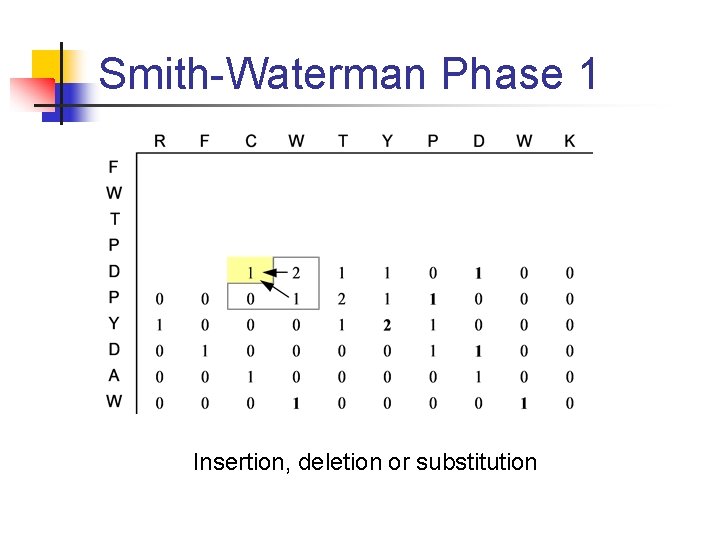
Smith-Waterman Phase 1 Insertion, deletion or substitution
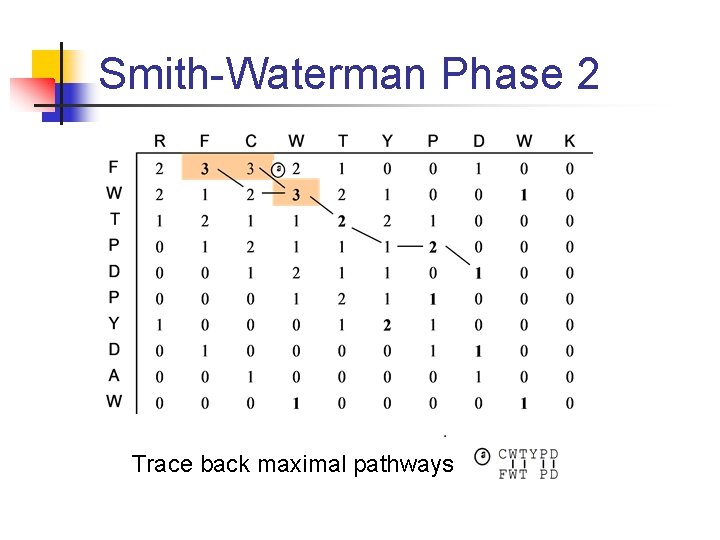
Smith-Waterman Phase 2 Trace back maximal pathways

Today’s problems one-to-many W NAC transpositions AN NA endpoint scoring FGAK +5 dual representations AGNCF. . . 85 116 39 85 100 Trying to fit new data into old tools… We need a way to model new problems quickly

Recurrence relations • • Basis of traditional DP Hard to design and understand Mixes together search space, scoring, and order of evaluation Subscript errors are common in implementation Smith-Waterman recurrence relation

Algebraic Dynamic Programming • • Giegerich, 2000 Grammars describe search space Set of functions (evaluation algebra) specifies scoring Solution space is pared down by an objective function first string $ gnirts dnoces

Grammar & eval. algebra localign = start <<< string ~~~ internal ~~~ string. . . h internal = subst delete insert end. . . h <<< aa ~~~ internal ~~~ aa ||| <<< aa ~~~ internal ||| <<< internal ~~~ aa ||| <<< string ~~~ ’$’ ~~~ string subst(a 1, score, a 2) insert(score, a) delete(a, score) end(str 1, $, str 2) h[score 1, . . . , scorek] = = = score + s(a 1, a 2) score – w 0 max[score 1, . . . , scorek]
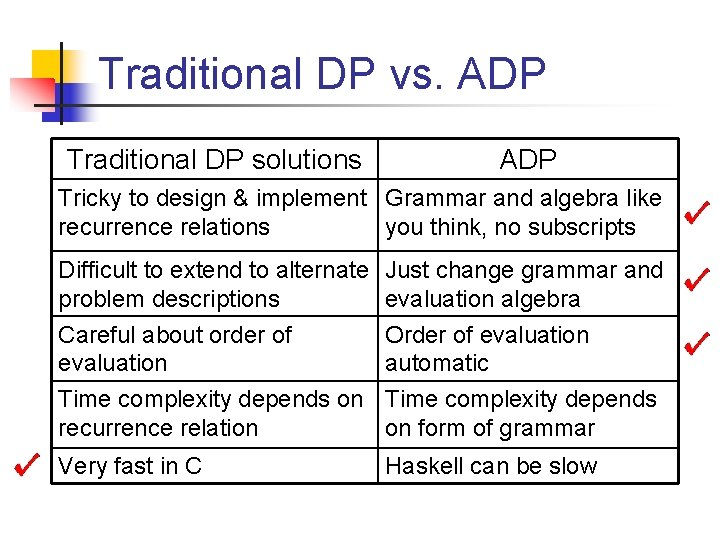
Traditional DP vs. ADP Traditional DP solutions ADP Tricky to design & implement Grammar and algebra like recurrence relations you think, no subscripts Difficult to extend to alternate Just change grammar and problem descriptions evaluation algebra Careful about order of evaluation Time complexity depends on recurrence relation Order of evaluation automatic Time complexity depends on form of grammar Very fast in C Haskell can be slow

Design of a DSL • Implement sample applications • • Abstract out common themes • • • Data structures Operations Decide how to handle errors • • • Use a flexible, higher-order language Run-time errors Type system Speed up implementation by generating C • • From embedded language, like PAN Using standalone compiler

The next steps • • • Compare our alignments with biologists’ Accumulate protein scores Try 2 -3 character statistical matches Pre-index using suffix trees Reduce memory usage Look for runs of exact matches as in FASTA

Our contributions • • • Built a working relationship with bio lab to understand domain Demonstrated feasibility of aligning large data sets in Haskell Constructed a framework for experimenting with representations for alignments, and heuristics for filtering target strings, localizing searches
- Slides: 33