Practical medical mycology Lab 10 Nonconventional identification techniques

ﺑﺴﻢ ﺍﻟﻠﻪ ﺍﻟﺮﺣﻤﻦ ﺍﻟﺮﺣﻴﻢ Practical medical mycology Lab 10 Non-conventional identification techniques

Introduction • Traditional identification techniques lack sensitivity & specificity, are slow, labor-intensive and require skilled personnel. • Many non-conventional approach have been developed to overcome the time and sensitivity issues; including molecular and serological techniques.

Conventional Fungal Identification Techniques Identification Method Concerns Culture ID - only in 26% positive if < 1 cfu/ml blood Germ-tube Formation - useful screening test only for C. albicans (does not differentiate C. ubliniensis) Carbohydrate Assimilation - limited species in the databases (e. g. Vitek & API strips) - can misidentify certain pathogenic species Morphological Characters - subjective measure, - high degree of skills required
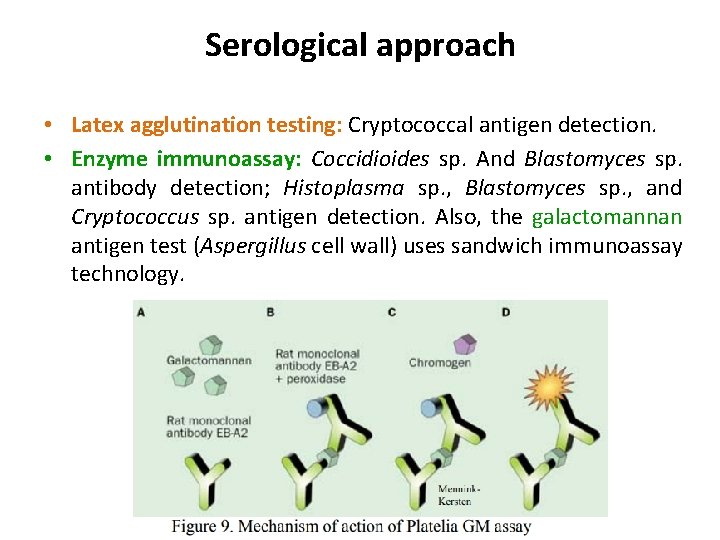
Serological approach • Latex agglutination testing: Cryptococcal antigen detection. • Enzyme immunoassay: Coccidioides sp. And Blastomyces sp. antibody detection; Histoplasma sp. , Blastomyces sp. , and Cryptococcus sp. antigen detection. Also, the galactomannan antigen test (Aspergillus cell wall) uses sandwich immunoassay technology.

Galactomannan antigen test • Invasive aspergillosis (IA) is a severe infection that occurs in patients with transplantation prolonged or in neutropenia, conjunction with following aggressive immunosuppressive regimens (eg, prolonged corticosteroid usage, chemotherapy). • IA has an extremely high mortality rate of 50% to 80% due in part to the rapid progression of the infection (ie, 1 -2 weeks from onset to death). • Approximately 30% of cases remain undiagnosed and untreated at death.

• Definitive diagnosis of IA requires histopathological evidence of deep-tissue invasion or a positive culture. • However, this evidence is often difficult to obtain due to the critically ill nature of the patient and the fact that severe thrombocytopenia often precludes the use of invasive procedures to obtain a quality specimen. • The sensitivity of culture in this setting also is low, reportedly ranging from 30% to 60% for Broncho alveolar lavage fluid.

• Accordingly, the diagnosis is often based on: • nonspecific clinical symptoms (unexplained fever, cough, chest pain, dyspnea). • in conjunction with radiologic evidence (computed tomography: CT scan); • and definitive diagnosis is often not established before fungal proliferation refractory to therapy. becomes overwhelming and

• Recently, a serologic assay was approved by the FDA for the detection of galactomannan, a molecule found in the cell wall of Aspergillus species. • Serum galactomannan can often be detected a mean of 7 to 14 days before other diagnostic clues become apparent. • Monitoring of galactomannan can potentially allow initiation of preemptive antifungal therapy before lifethreatening infection occurs.

Serological approach • Immunodiffusion: Histoplasma sp. And Blastomyces sp. Antibody detection. • Immunofluorescence assay: CAGTAa assay is an indirect IFA. • ( Testing for Candida albicans germ-tube antibody IFA Ig G assay) • Indirect immunofluorescent assay kit to test specific Ig. G antibodies against antigens located on the cell wall surface of of Candida albicans in human serum/plasma.

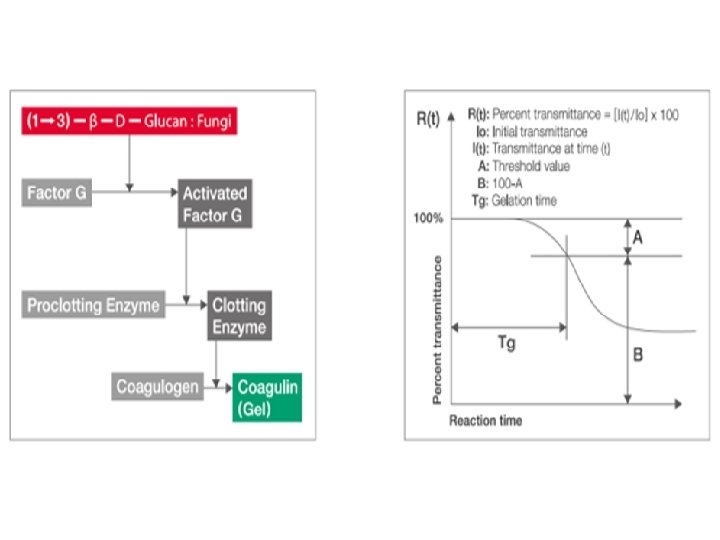
• Complement fixation: Histoplasma sp. , Blastomyces sp. , and Coccidioides sp. antibody detection. • lateral-flow assay: Point-of-care diagnostic tests for Cryptococcus sp. And Aspergillus sp. Antigen detection. • G test: is specific to beta-glucan detection. Factor G is a proclotting factor that is highly sensitive to beta-glucan. • When a patient’s sample containing beta-glucan is added to a mix containing factor G, it activates the factor, thus initiating an enzymatic cascade that results in a color or optical density change of the mixture, which can be detected with colorimetric or turbid metric methods.

Applications of molecular methods 1. Organism detection in blood, body fluids and tissue 2. Identification of the fungal agent 3. Quantification of the fungal load 4. Monitoring of antifungal treatment

Molecular approaches DNA probes PCR primers • Southern Hybridization • Conventional • In-situ Hybridization • Panfungal PCR • Microarray and Macroarray • Multiplex PCR • T 2 nuclear magnetic resonance • Nested PCR • MALDI-TOF • Real Time PCR • Sequencing • PCR-RFLP Karyotyping

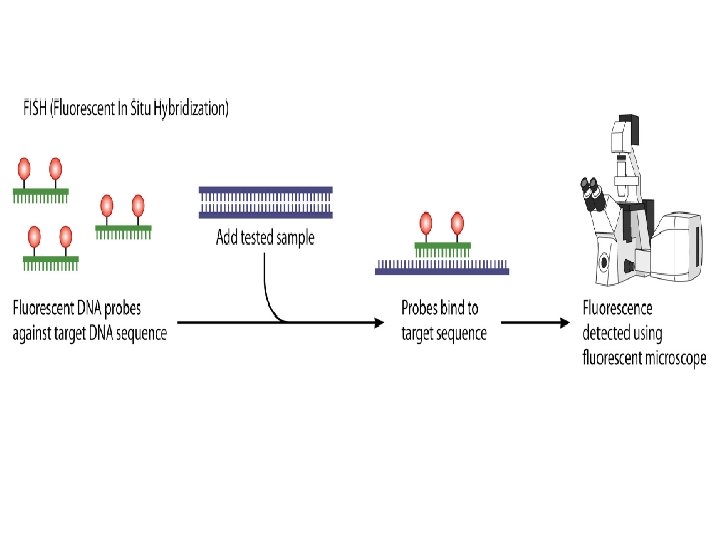
• Southern blotting: • It is a hybridization technique for identification of particular size of DNA from the mixture of other similar molecules. • This technique is based on the principle of separation of DNA fragments by gel electrophoresis and identified by labelled probe hybridization. • In situ hybridization: • (ISH) is a type of hybridization that uses a labeled complementary DNA, RNA or modified nucleic acids strand (i. e. , probe) to localize a specific DNA or RNA sequence in a portion or section of tissue (in situ)

Primer targets • Universal fungal primers for: - Multi-copy Genes r. DNA gene cluster 18 S, ITS 1/2, 5. 8 S, 28 S, 5 S, IGS - Single-copy Genes Actin, Alkaline Protease (ALP), Chitin Synthase, GP 43, Lanosterol - - demethylase (LIA 1), URA 5, Secreted Aspartic Protease (SAP), Beta glucan synthetase (FKS), Histone, etc. • Genus- or species-specific primers 18 S, ITS 1/2, 28 S r. DNA, Mitochondrial DNA, Histone e. g. Candida, Cryptococcus, Aspergillus

Commercial molecular assays for the detection of fungal pathogens. Assay Vendor (location) Yeast Traffic Light® Advan. Dx (MA, USA) Myc. Assay™ Pneumocystis Trinity Biotech (County Wicklow, Real-time PCR Ireland) Myc. Assay™ Aspe Trinity Biotech rgillus Luminex x. TAG®fungal assay Method Target PNA–FISH 26 S Blood (from positive bloodr. RNA; Candida s culture bottles) p. Real-time PCR Specimen type ml. SU; Pneumoc BAL; sputum ystis jirovecii 18 S r. RNA; Aspergill BAL; serum us sp. Luminex Molecular Multiplex PCR coupled with Up to 23 fungi BAL; blood Diagnostics (ON, bead probe fluid array Canada) IBIS Plex-ID IBIS Biosciences fungal spectrum (CA, USA) assay Multiplex PCR coupled with Large-subunit electrospray ionization r. RNA; up to 75 BAL; blood mass spectrometry fungi BAL: Bronchoalveolar lavage fluid; ITS: Internal transcribed spacer; ml. SU: Mitochondrial large subunit; PNA: Peptide nucleic acid.
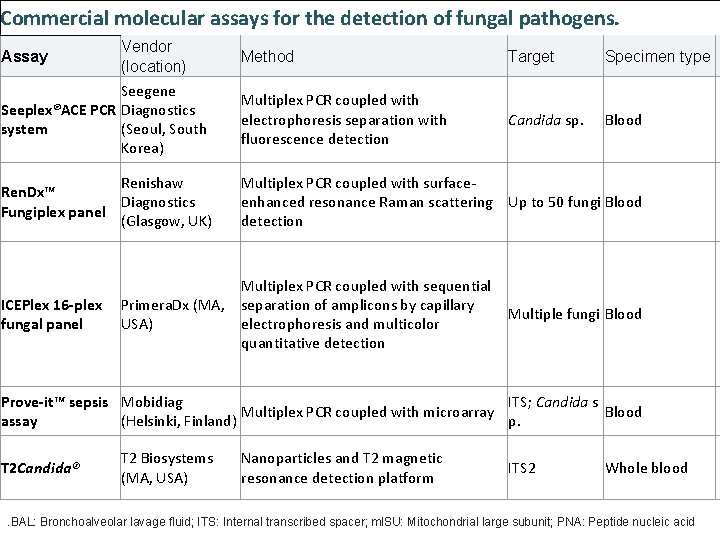
Commercial molecular assays for the detection of fungal pathogens. Assay Vendor (location) Method Target Specimen type Seegene Seeplex®ACE PCR Diagnostics system (Seoul, South Korea) Multiplex PCR coupled with electrophoresis separation with fluorescence detection Candida sp. Blood Renishaw Ren. Dx™ Diagnostics Fungiplex panel (Glasgow, UK) Multiplex PCR coupled with surfaceenhanced resonance Raman scattering Up to 50 fungi Blood detection ICEPlex 16 -plex fungal panel Multiplex PCR coupled with sequential separation of amplicons by capillary electrophoresis and multicolor quantitative detection Primera. Dx (MA, USA) Multiple fungi Blood Prove-it™ sepsis Mobidiag ITS; Candida s Multiplex PCR coupled with microarray Blood assay (Helsinki, Finland) p. T 2 Candida® T 2 Biosystems (MA, USA) Nanoparticles and T 2 magnetic resonance detection platform ITS 2 Whole blood . BAL: Bronchoalveolar lavage fluid; ITS: Internal transcribed spacer; ml. SU: Mitochondrial large subunit; PNA: Peptide nucleic acid

PCR Assay Sensitivity • Multi-copy Genes: e. g. 18 S r. RNA 1 -5 CFU ITS 10 CFU/ml blood • Single-copy Genes: e. g. Actin 1 -10 CFU 1. 4 - lanosterol demethylase 10 -100 CFU/ml blood Heat-shock protein 90 10 -100 CFU

Issues regarding the use of PCR for fungal diagnostics 1. Choice of sample 2. DNA extraction 3. Primer selection 4. Type of PCR 5. In vitro validation of a certain PCR 6. Co-infection by multiple microbial species

1 - Choice of sample • The best sample varies depending on the target pathogen and the site where it is preferentially accumulated. • Tissue samples can be ideal to evaluate a new test, as they can differentiate between colonization and infection. • However, less invasive samples, such as BAL fluid, serum, and whole blood, are favorable because they can be utilized as screening tests. • Serum also allows for multiple tests to be performed with the sample.

2 - DNA extraction • The general idea is to maximize and concentrate the amount of fungal cells or free fungal DNA in the tested sample. • However, even with perfect DNA extraction, some fungal species may be found in the circulation only transiently when they establish deep-seated infections. • The rigidity of fungal cell wall would increase time and steps needed for cell lysis as will as the contamination chances

3 - Primer selection • The target amplification sequence should be found in multiple repeats and should differ from the respective host sequence. • r. DNA seems to be superior to mitochondrial DNA for diagnosis of aspergillosis. 4 - Type of PCR • Nested PCR: • It is a modification of PCR that was designed to improve sensitivity and specificity.

• Nested PCR involves the use of two primer sets and two successive PCR reactions. The first set of primers are designed to anneal to sequences upstream from the second set of primers and are used in an initial PCR reaction. • It requires additional time and might be more prone to contamination due to the additional amplification step. • Real-time PCR is a technique used to monitor the progress of a PCR reaction in real time. At the same time, a relatively small amount of PCR product (DNA, c. DNA or RNA) can be quantified. • It allows for quantitation of the amplified DNA and thus could help to differentiate infection from colonization.

5. In vitro validation of a certain PCR • These methods are of paramount importance for the accurate evaluation of the sensitivity of any PCR and allow for inter-laboratory comparisons of the results. 6. Co-infection by multiple microbial species • This technique allows for the simultaneous identification of multiple microbial pathogens from the sample. • The broadest method that has been proposed is the multiplex Septi. Fast PCR (ROCHE).
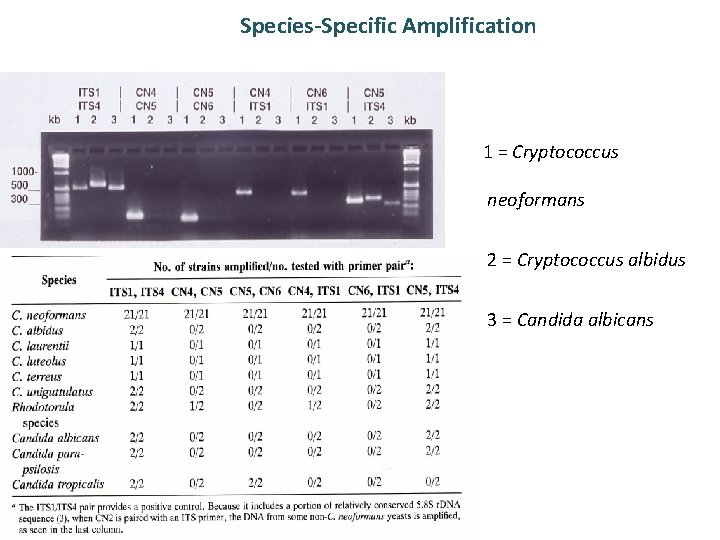
Species-Specific Amplification 1 = Cryptococcus neoformans 2 = Cryptococcus albidus 3 = Candida albicans
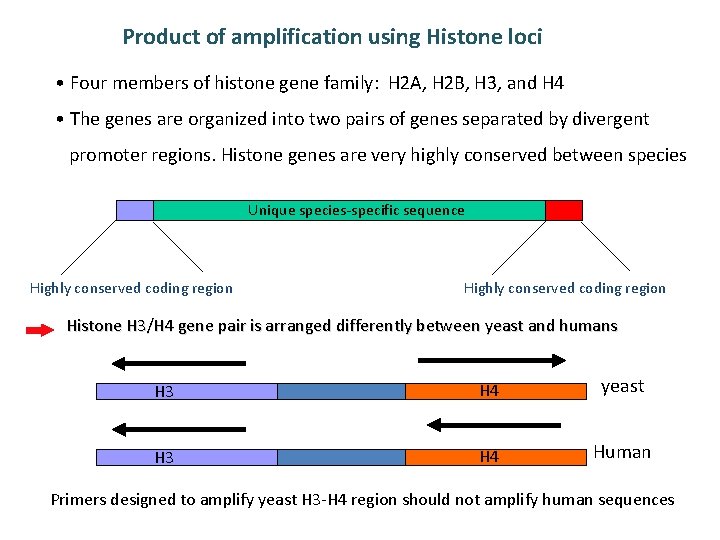
Product of amplification using Histone loci • Four members of histone gene family: H 2 A, H 2 B, H 3, and H 4 • The genes are organized into two pairs of genes separated by divergent promoter regions. Histone genes are very highly conserved between species Unique species-specific sequence Highly conserved coding region Histone H 3/H 4 gene pair is arranged differently between yeast and humans H 3 H 4 yeast H 3 H 4 Human Primers designed to amplify yeast H 3 -H 4 region should not amplify human sequences
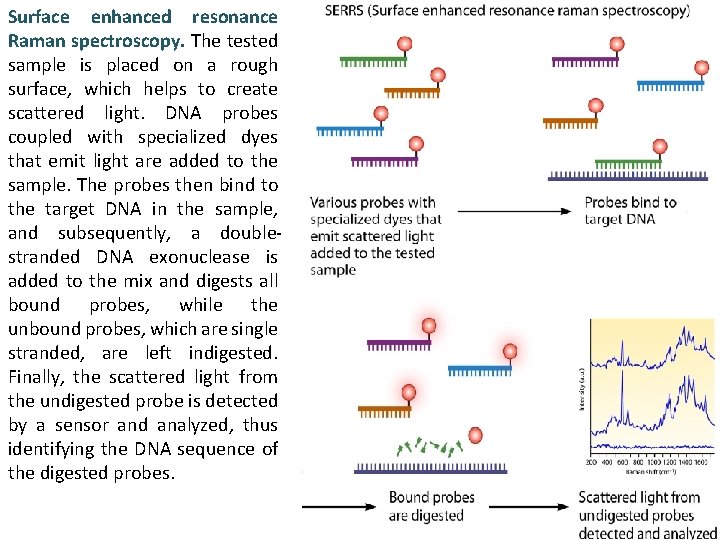
Surface enhanced resonance Raman spectroscopy. The tested sample is placed on a rough surface, which helps to create scattered light. DNA probes coupled with specialized dyes that emit light are added to the sample. The probes then bind to the target DNA in the sample, and subsequently, a doublestranded DNA exonuclease is added to the mix and digests all bound probes, while the unbound probes, which are single stranded, are left indigested. Finally, the scattered light from the undigested probe is detected by a sensor and analyzed, thus identifying the DNA sequence of the digested probes.
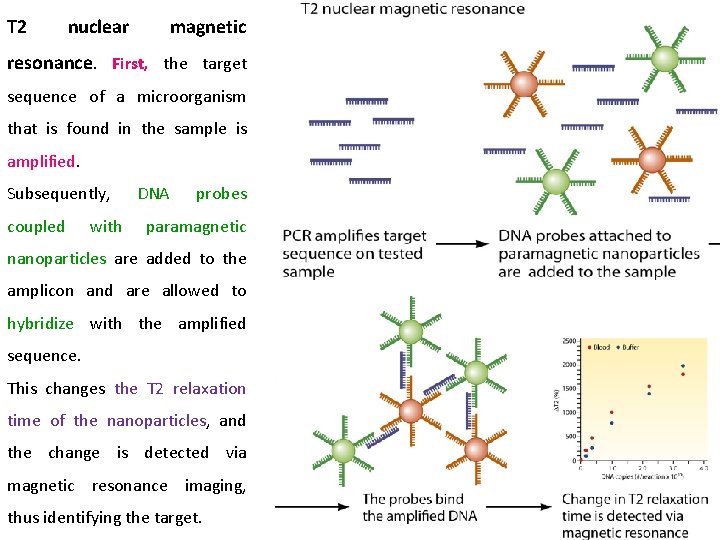
T 2 nuclear magnetic resonance. First, the target sequence of a microorganism that is found in the sample is amplified. Subsequently, coupled with DNA probes paramagnetic nanoparticles are added to the amplicon and are allowed to hybridize with the amplified sequence. This changes the T 2 relaxation time of the nanoparticles, and the change is detected via magnetic resonance imaging, thus identifying the target.

PCR-RFLP (Restriction Fragment Length Polymorphism Analysis) ITS 1, 5. 8 S, ITS 2 region • Clinical specimen or direct PCR of a single colony form a primary isolation plate, tissue sample, or clean culture • Digestion with restriction enzymes • Agarose gel electrophoresis • ID via comparison with the data base • Species Level

DNA Array Technology • Small glass or silicon matrices on which potentially thousands of short oligonucleotides can be immobilized e. g. - species-specific DNA probes - antifungal resistance genes - virulence genes C. g. C. p. C. l. C. k. C. n. C. li. C. m. C. t. C. u. Macroarray C. glabrata C. parapsilopsis C. lipolytica C. krusei C. norvegensis C. lusitaniae C. multigemmis C. tropicalis C. utilis Microarray

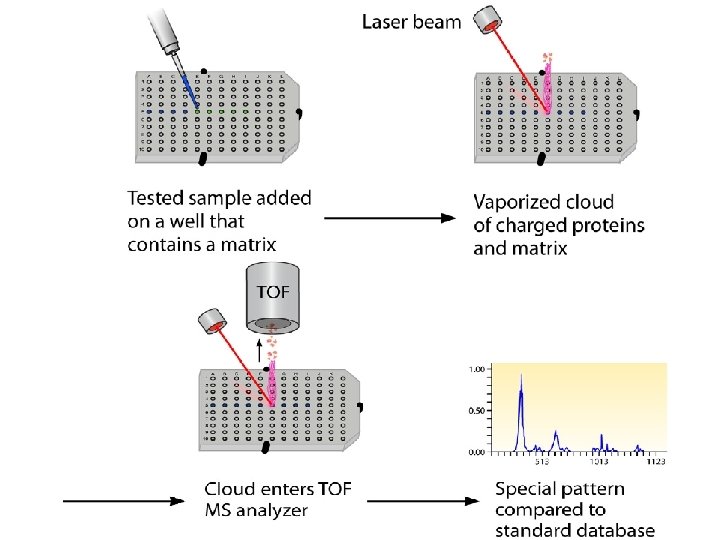
Matrix-assisted laser desorption ionization– time of flight mass spectrometry (MALDI-TOF) • The tested sample is added to a well that contains matrix material which has the ability to absorb UV light and transform it into heat. • A laser beam is targeted to the mix. The laser beam is absorbed by the matrix, and part of the analyte-matrix mix is vaporized and ionized, creating a cloud of ionized proteins and matrix. • This cloud subsequently is subjected to an electric field, which leads the particles to accelerate toward a detector. The mass and charge of each particle determine the time needed to reach the detector. • This allows the mass spectrometer to determine the characteristics of the particles within the tested sample. Comparison of the produced spectral pattern against a standard database allows for identification of the microorganism in the sample.
- Slides: 33