Practical 14 Use gel electrophoresis to separate DNA




































- Slides: 44

Practical 14 Use gel electrophoresis to separate DNA fragments of different length A DNA profile is a genetic fingerprint of an organisms DNA Everyone's DNA is unique DNA profiling can be used to identify people (Criminals, Paternity) and to determine genetic relationships between animals and plants What do we need to revise first?

Learning outcomes: DNA profiling • 6. 3 Know how DNA profiling is used for identification and determining genetic relationships between organisms (plants and animals). Polymerase chain reaction • 6. 4 Know how DNA can be amplified using the polymerase chain reaction (PCR). CORE PRACTICAL 14: • Use gel electrophoresis to separate DNA fragments of different length.

Restriction enzymes and Gel electrophoresis: Links • http: //media. pearsoncmg. com/intl/snab_2009/top ic_6/interactives/6_1/topic_6_1. html PCR: • http: //media. pearsoncmg. com/intl/snab_2009/top ic_6/interactives/6_4/topic_6_4. html • Reminder of some of the linked theory. .

Introns and Exons - basic • DNA contains a lot of ‘junk’ which do not code for proteins (and therefore is theoretically useless to the organism? ) Exons Introns Exons • N. B. The introns are removed during transcription to make m. RNA • http: //uktv. co. uk/alibi/homepage/sid/6920

What’s happening in an Intron? • Introns are basically short DNA sequences often repeated many times • The sequence of repeated bases = Short Tandem Repeats (STRs) or satellites • An STR can contain 2 -50 base pairs • And can be repeated from 5 -several hundred times! • Occur at same place/loci on both chromosomes of an homologous pair • But the number of times they are repeated can be different • BUT There will be variations in the number of repeats between the maternal and paternal chromosome in an individual. As there a large number of introns with lots of STR loci – and there can be a large variation of number of repeats – it means that it is highly unlikely for two individuals to have to the same combination of STR’s

How has the discovery of microsatellites (STRs) enabled us to develop DNA profiling? 3 marks • The number of repeating STRs / microsatellites vary considerably in different people. • This has enabled DNA profiling to be developed, which identifies the natural variations found in every persons DNA. • Scientists can create a virtually unique DNA profile.

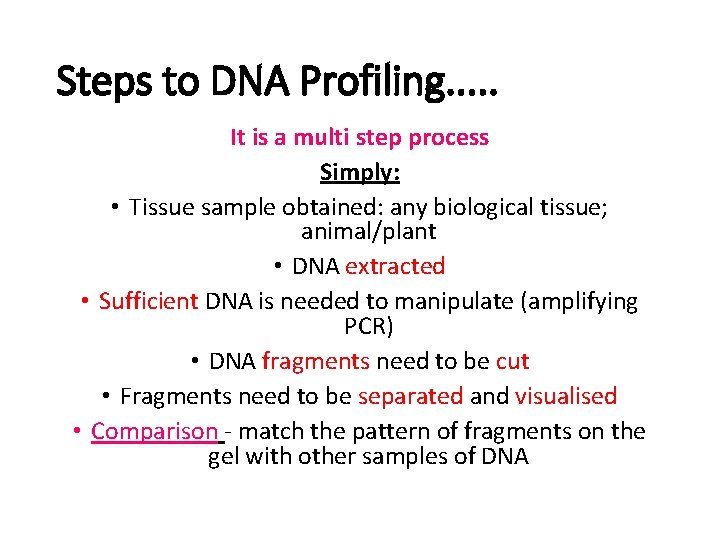
Steps to DNA Profiling. . . It is a multi step process Simply: • Tissue sample obtained: any biological tissue; animal/plant • DNA extracted • Sufficient DNA is needed to manipulate (amplifying PCR) • DNA fragments need to be cut • Fragments need to be separated and visualised • Comparison - match the pattern of fragments on the gel with other samples of DNA

Steps to DNA Profiling. . . DNA fingerprinting requires very little sample and modern techniques are improving this all the time. Once the DNA is obtained the basic steps are: • Treatment by Restriction Enzymes to create fragments of DNA • PCR: amplify genetic material • Gel Electrophoresis: to separate the fragments • Southern Blotting: Visualising • Interpretation: Comparison - match the pattern of fragments on the gel with other samples of DNA

Obtaining the DNA: • Tissue is placed in a buffer solution that contains salt and detergent, heated and is physically broken down: • The detergent causes the cell membrane to break down by dissolving the lipids and proteins of the cell and disrupting the bonds that hold the cell membrane together releasing DNA from the nucleus and mitochondria. • The detergent, combined with the heat treatment causes lipids (fatty molecules) and proteins to precipitate out of the solution, leaving the DNA. • DNA is not soluble in alcohol. When alcohol is added to the mixture, the components of the mixture, except for DNA, stay in solution while the DNA precipitates out into the alcohol layer

Creating DNA Fragments • Need to use restriction enzymes (N. B. they can also be referred to as restriction endonucleases) • Restriction enzymes are found naturally in bacteria – their job is to cut up invading viral DNA – why is this important? • They are basically a bacterial defence mechanism (like larger organisms immune systems) • They cut the foreign DNA up, (Molecular scissors ) at a specific base sequence. (protects own (bacterial) DNA!) • Different restriction enzymes cut at different base sequences. • These enzymes only cut DNA at specific base sequences e. g. before and after STRs - this is what makes them so valuable forensic scientists!! • Hundreds have been discovered (400) and type II are very useful as they cut at specific nucleotide sequences (recognition sites): • Eco RI, Hind III

Restriction endonucleases cont… Eco RI – cuts at this site (known as a recognition site) between the G and A bases in the sequence GAATTC G A A T T C C T T A A G 1 st enzyme found in R strain of E. coli Hind III – cuts at this site (known as a recognition site) A A G C T T C G A A

• Therefore these enzymes can be used before specific STR regions (at the same loci). Cuts away this fragment of DNA from the rest of the genome • On the same person you would get exactly the same STR lengths on identical DNA samples • On different samples you should get different lengths • ALLOWS IDENTIFICATION!!!!

Restriction enzymes: Task: 5 minutes individual Complete activity 6. 1 READ and answer questions 1 -2 You can have a go at animation later. . 1. What are restriction enzymes? 2. What is their significance to molecular biologists and forensic scientists?

Restriction enzymes

Answers 1. What are restriction enzymes? A group of enzymes which are able to cut a DNA molecule at particular points along its length, at recognition sites. Each restriction enzyme recognises a specific base sequence and will only cut a DNA molecule where this sequence occurs. 2. What is their significance to molecular biologists and forensic scientists? Restriction enzymes are very important tools in genetic engineering. They can be used, for example, to isolate particular genes and insert them into the DNA of another organism. Can also be used in Genetic screening and DNA profiling

Polymerase Chain Reaction • The problem is – that to compare DNA – you need a lot of it • Also many samples maybe very small samples (a drop of blood, one strand of hair) • Therefore the polymerase chain reaction is used to amplify the DNA section (STR area) for comparison What is PCR? PCR is a technique that can amplify small DNA samples into large ones for scientific analysis (Biological photocopying!) It is highly specific, easily automated and capable of amplifying minute amounts of a sample. It has had a major impact on clinical medicine, genetic disease diagnostics, forensic science and evolutionary biology.

How does PCR work? The whole process is only possible because of special heat-stable enzyme called taq polymerase, isolated from thermophilic bacteria. This enzyme can tolerate temperatures of 95°C and has an optimum temperature of 72°C Most DNA polymerases function best at 37°C, denatured at higher temperatures

How does PCR work? The target length of DNA to be copied is selected using primers. Primers: Short DNA sequences (20 -30 nucleotides) complementary to the DNA adjacent to the STR, at the start of the fragment you want The primers are marked with fluorescent tags. The PCR reaction mixture contains: 1. DNA sample, 2. Excess of appropriate primers, 3. DNA polymerase enzyme (creates new DNA strand: phosphodiester bonds) 4. Nucleotides (to build new DNA) 5. Buffer

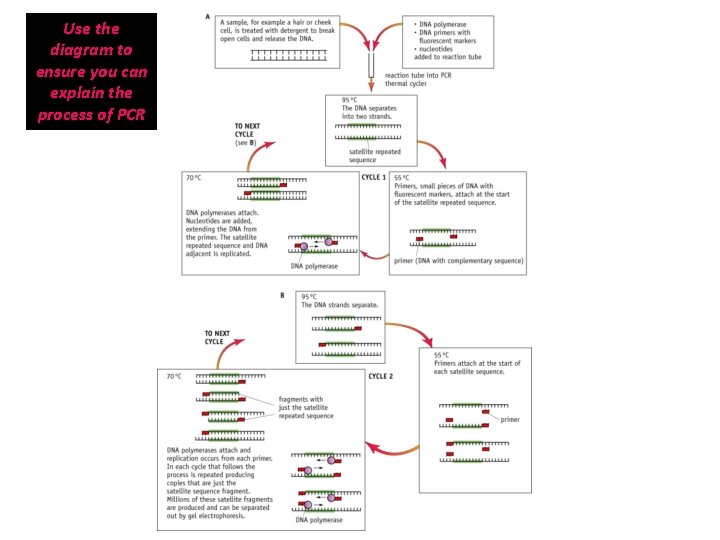
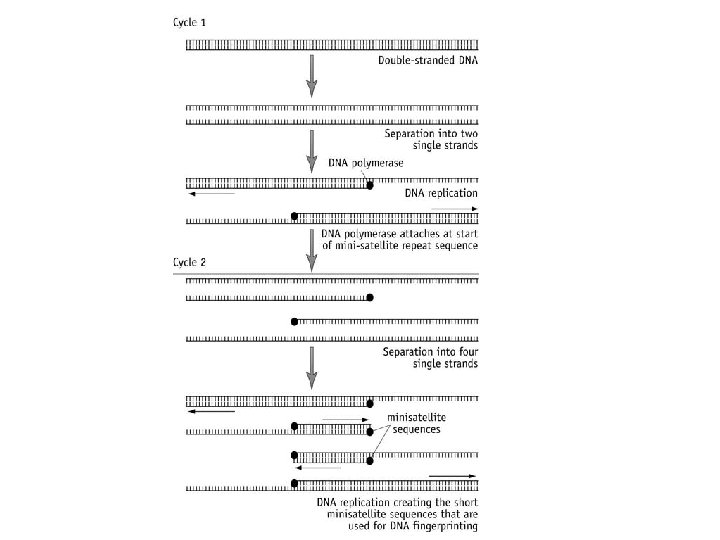
How does PCR cycle work? The forensic sample is placed in the tube with the PCR reaction mixture In the PCR machine this tube undergoes a series of temperature changes: Stage 1: • DNA strands are denatured by heating to 95°C (20 secs) • Hydrogen bonds break and DNA strands become separated. • (How is DNA normally separated? ) Enzyme helicase Stage 2: • Cool between 55 -65°C (20 seconds) • Allows primers (with fluorescent markers) to join/anneal at the start of the STR repeat sequence). • Using excess of primers ensures that some bind with the DNA, rather than the original strands joining again, thus extending the DNA

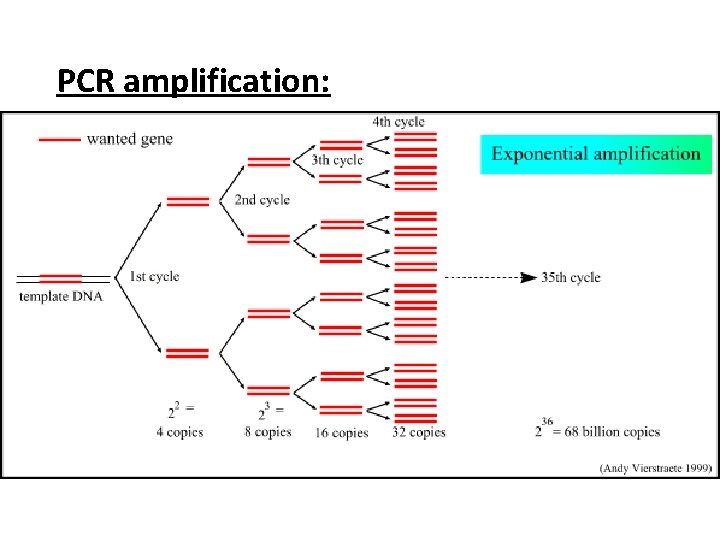
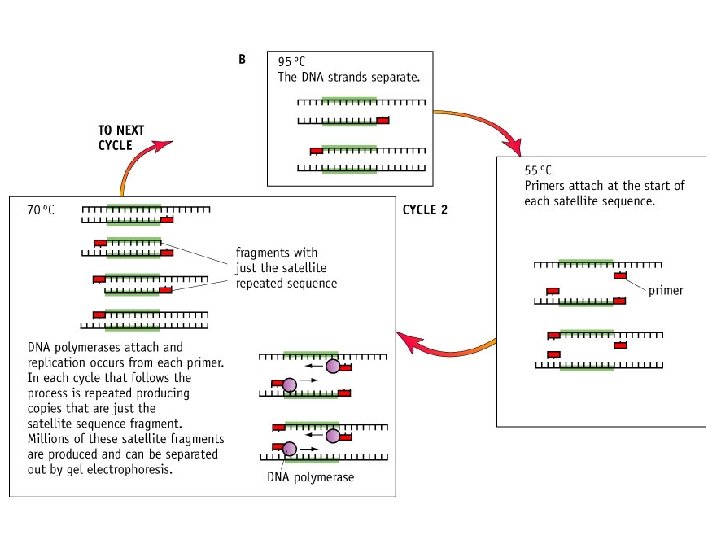
Stage 3: • Heat to 70° C (30 secs) • DNA polymerases attach (optimum temperature) • Complimentary DNA nucleotides are added to the separated template strands, by complementary base pairing; hydrogen bonds; DNA polymerase catalyses the joining of nucleotides; Phosphodiester bonds. • Producing a 2 new copies of DNA strand complementary to the DNA template strand The new strands are then denatured and the process repeated until large enough samples have been produced. The cycle is repeated many times and each time the number is doubled.

PCR amplification:

Use the diagram to ensure you can explain the process of PCR


TASK 1: PCR animation and work sheet. 10 minutes • http: //media. pearsoncmg. com/intl/snab_2009/top ic_6/interactives/6_4/topic_6_4. html • Look at animation 1(later/together) I can forward? • Activity 6. 4 Questions 1 -5 • LO: To understand how the polymerase chain reaction amplifies DNA


A 1 B 2 A 2 Plenary What can you remember? B 1 D C E Lots of good animations: WATCH Watch

UK Forensic Science Service (FSS) • In the UK – DNA is generally analysed at 10 short tandem repeat sequences: micro-satellites • Each STR is only 4 bases in length + an additional primer is used to determine sex, on the sex chromosome • This is enough to guarantee that the odds of someone else sharing same result are unlikely (one in a billion!) • To match loci • To match a sample of DNA all ten loci must be of the same length! • If even one is different there is no match So we now have the amplified sample of DNA What’s next?

Separating the fragments of DNA: Gel Electrophoresis • The DNA fragments produced by restriction enzymes or PCR can be separated • Gel Electrophoresis is a technique used to separate molecules by their size and by their charge • To allow visualisation and analysis!

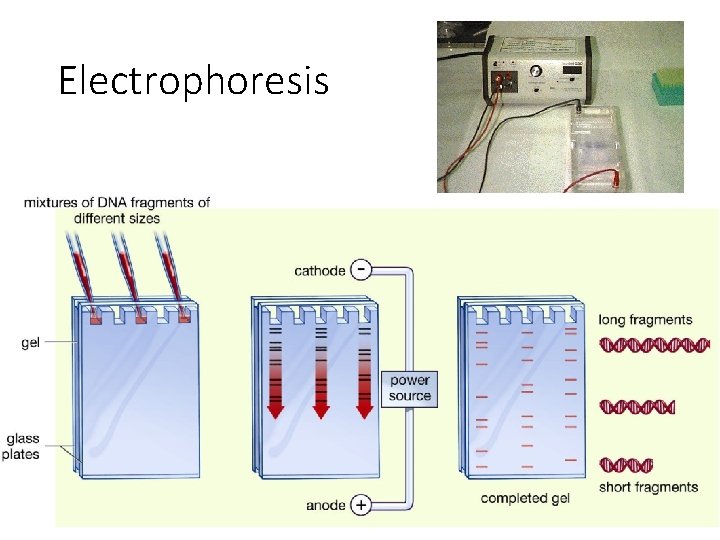
How it works • A gel is made from a chemical called agarose (made from seaweed!) or polyacrylamide • Provides a stable medium through which fragments can move. • Samples of unknown DNA and DNA of known lengths are loaded at one end of the gel • Gel is put into buffer solution, connected to electrodes that produce a pd across the gel • An electrical current is passed through the gel • As DNA is mainly negative it is attracted to the positive electrode (anode) • The smaller the DNA fragment, with smaller numbers of repeat sequences; the quicker/faster and further it travels along the gel. So smaller samples are closet to cathode (+ve) • So DNA fragments are separated according to length (measured in number of base pairs)

Making the gel

Adding samples

Electrophoresis

Task • • • Activity 6. 1 Restriction enzymes and Gel electrophoresis http: //media. pearsoncmg. com/intl/snab_2009/topic_6 /interactives/6_1/topic_6_1. html Watch other animation Watch you tube film https: //www. youtube. com/watch? v=2 a 4 d. Suaiks. Q 4 minutes Worksheet to go with Activity 6. 1 q 1 -7 • So we now our gels prepared. . . how do we visualise them?

Visualising the fragments: • Once the fragments have been separated, we can examine • • • the gel and see what sizes of bands are found on it. When a gel is stained with a DNA-binding dye and placed under UV light, the DNA fragments will glow, allowing us to see the DNA present at different locations along the length of the gel A well-defined “line” of DNA on a gel is called a band. Each band contains a large number of DNA fragments of the same size that have all travelled as a group to the same position. A single DNA fragment (or even a small group of DNA fragments) would not be visible by itself on a gel. The bp next to each number in the ladder indicates how many base pairs long the DNA fragment is By comparing the bands in a sample to the DNA ladder, (A solution of doublestranded DNA fragments whose molecule weights (and number of base pairs) are known and standardised. The ladder is used to calibrate electrophoresis gels so that samples of unknown DNA that have been introduced into the gel can be measured and their approximate sizes can be determined. The gel is fragile so a technique was developed to transfer the fragments onto a more resilient nylon/ nitrocellulose membrane

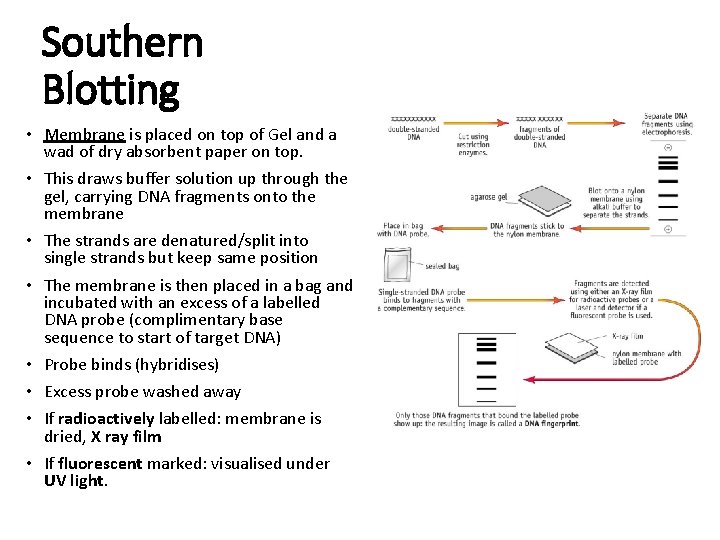
Southern Blotting • Membrane is placed on top of Gel and a wad of dry absorbent paper on top. • This draws buffer solution up through the gel, carrying DNA fragments onto the membrane • The strands are denatured/split into single strands but keep same position • The membrane is then placed in a bag and incubated with an excess of a labelled DNA probe (complimentary base sequence to start of target DNA) • Probe binds (hybridises) • Excess probe washed away • If radioactively labelled: membrane is dried, X ray film • If fluorescent marked: visualised under UV light.

A single band on profile= maternal+ paternal chromosomes have same number STR repeats at a loci 2 bands= chromosomes have different number of STR repeats

Crime Scene Investigation • http: //www. dnalc. org/resources/animations/pcr. ht ml - animation on PCR • http: //www. dnaftb. org/ - this is an excellent site for all areas of DNA and genetics

Test yourself: Exam question 1. 5 marks Describe the techniques 2. 6 marks Researchers carried out a study Left to look at? The uses of DNA Fingerprinting

Potential suspects of a crime

Or to a potential father (obviously not a mother) in a paternity case.

DNA fingerprinting has a variety of uses: Matching unknown biological samples to known samples. This is used in: 1. Forensic Science 2. Identifying a family relationship (mother/father/siblings etc) 3. Paternity disputes 4. Identifying a species e. g. Tiger DNA from a pelt, useful in conservation 5. Identifying stolen animals 6. Used in classification to identify common ancestor 7. Identify body parts 8. Industrial sabotage and contamination 9. It can also be used for screening of genetic conditions (e. g. Cystic Fibrosis) 10. Determining evolutionary relationships (animals more closely related will share patterns in their DNA e. g. Chimpanzees share around 99% of human DNA patterns the next closest relative the gorilla shares around 97%) 11. Identifying animal products made from endangered species illegally.

Environmental conservation: DNA profiling Has been used to protect indigenous populations of mammals such as the woylie (Bettongia penicillata) and birds, such as the white-tailed black cockatoo (Calyptorhynchus baudinii) and red-tailed black cockatoo (Calyptorhynchus banksii). These birds are valued by breeders and are often taken illegally from nests. DNA profiles can be used as evidence against breeders who claim birds taken from the wild have been bred in captivity

DNA profiling used on cannabis plants • FORENSIC scientists in the US are applying DNA fingerprinting methods to the cannabis plant. • They say the technique, which is being used to create a database of DNA profiles of different marijuana plants, will help them to trace the source of any sample

Is DNA profiling infallible? • Many stages, so artefacts/contamination can arise at any stage • Only a few sequences are analysed so. . . • There is the possibility of 2 identical profiles from 2 unrelated individuals 91 in a billion!) • Identical twins/close relatives may show same profile for STRs chosen. • The more satellites used: more reliable/unique