Posttranscriptional Modification of RNA Effect on Biology and

Post-transcriptional Modification of RNA: Effect on Biology and Virulence of Salmonella Amin A. Fadl University of Wisconsin-Madison
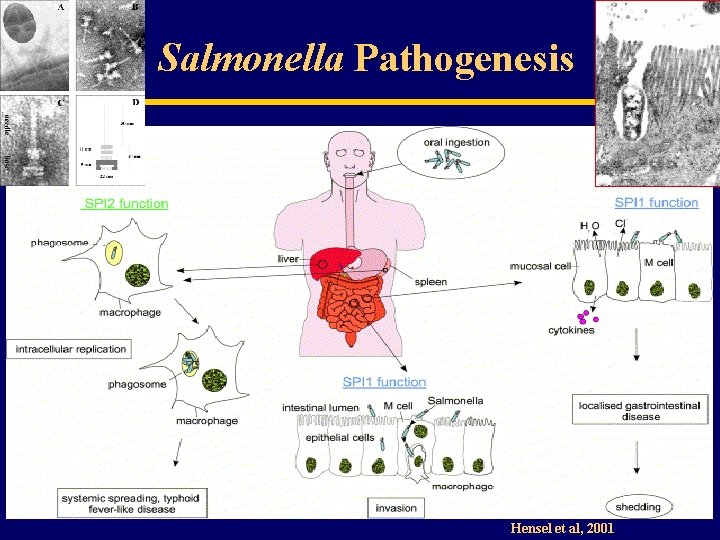
Salmonella Pathogenesis Hensel et al, 2001

t. RNA Modification Enzymes v t. RNAs are key molecules of translational machinery that ensure decoding of successive codons in m. RNA inside the ribosome. v Post-transcriptional t. RNA modification is found in all organisms and is required for t. RNA functions, control gene expression v Glucose-inhibited division gene (Gid. A, Mnm. G), methylaminomethyl (Mnm. E), Ribosomal Small Subunit Methyltransferase (Rsm. G, Gid. B)
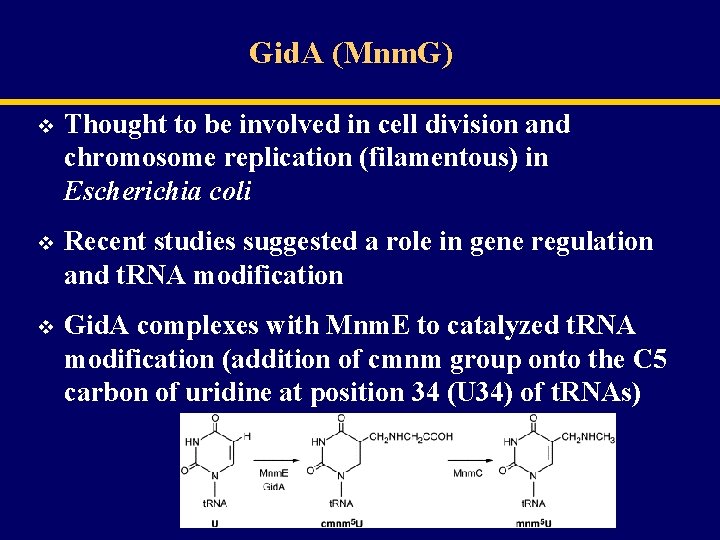
Gid. A (Mnm. G) v Thought to be involved in cell division and chromosome replication (filamentous) in Escherichia coli v Recent studies suggested a role in gene regulation and t. RNA modification v Gid. A complexes with Mnm. E to catalyzed t. RNA modification (addition of cmnm group onto the C 5 carbon of uridine at position 34 (U 34) of t. RNAs)
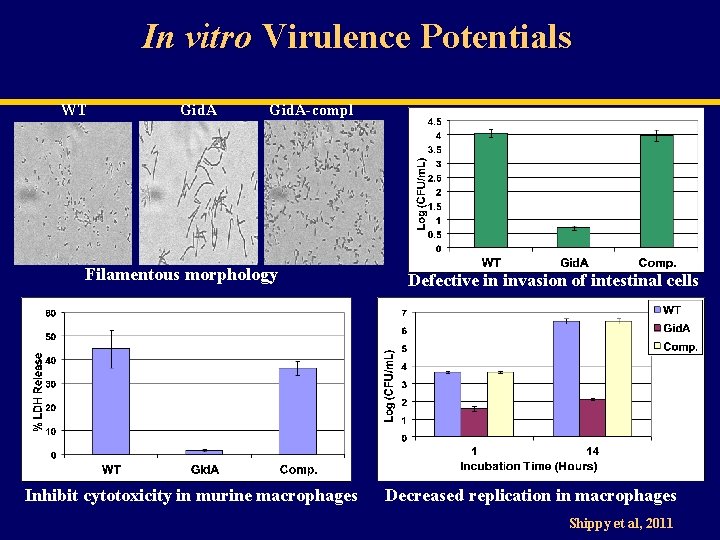
In vitro Virulence Potentials WT Gid. A-compl Filamentous morphology Inhibit cytotoxicity in murine macrophages Defective in invasion of intestinal cells Decreased replication in macrophages Shippy et al, 2011
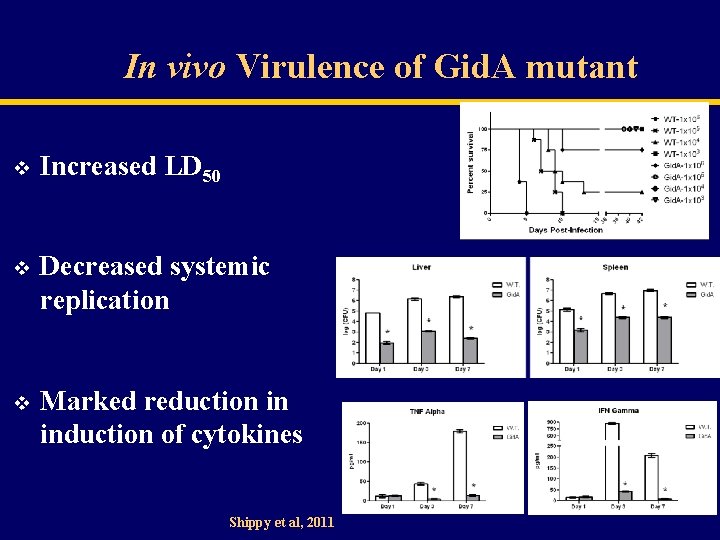
In vivo Virulence of Gid. A mutant v Increased LD 50 v Decreased systemic replication v Marked reduction in induction of cytokines Shippy et al, 2011
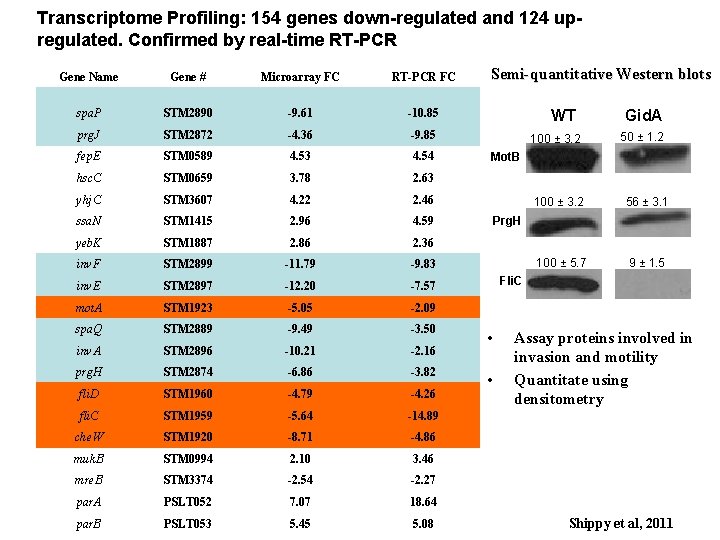
Transcriptome Profiling: 154 genes down-regulated and 124 upregulated. Confirmed by real-time RT-PCR Gene Name Gene # Microarray FC RT-PCR FC spa. P STM 2890 -9. 61 -10. 85 prg. J STM 2872 -4. 36 -9. 85 fep. E STM 0589 4. 53 4. 54 hsc. C STM 0659 3. 78 2. 63 yhj. C STM 3607 4. 22 2. 46 ssa. N STM 1415 2. 96 4. 59 yeb. K STM 1887 2. 86 2. 36 inv. F STM 2899 -11. 79 -9. 83 inv. E STM 2897 -12. 20 -7. 57 mot. A STM 1923 -5. 05 -2. 09 spa. Q STM 2889 -9. 49 -3. 50 inv. A STM 2896 -10. 21 -2. 16 prg. H STM 2874 -6. 86 -3. 82 fli. D STM 1960 -4. 79 -4. 26 fli. C STM 1959 -5. 64 -14. 89 che. W STM 1920 -8. 71 -4. 86 muk. B STM 0994 2. 10 3. 46 mre. B STM 3374 -2. 54 -2. 27 par. A PSLT 052 7. 07 18. 64 par. B PSLT 053 5. 45 5. 08 Semi-quantitative Western blots WT 100 ± 3. 2 Gid. A 50 ± 1. 2 Mot. B 100 ± 3. 2 56 ± 3. 1 100 ± 5. 7 9 ± 1. 5 Prg. H Fli. C • • Assay proteins involved in invasion and motility Quantitate using densitometry Shippy et al, 2011
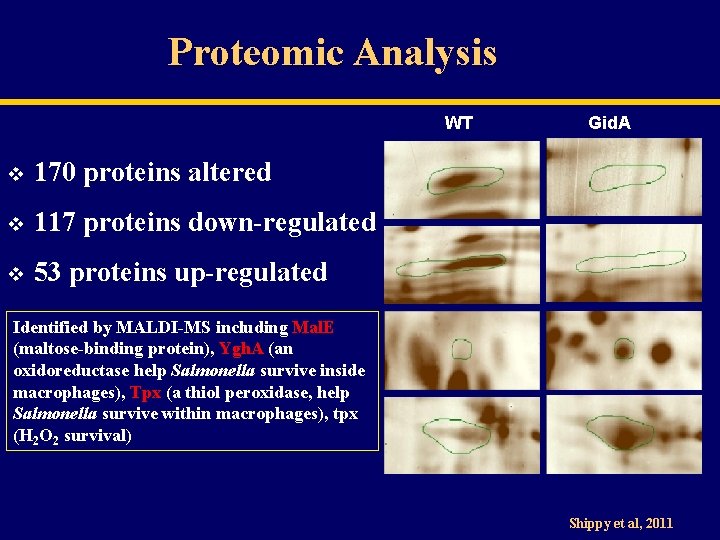
Proteomic Analysis WT v 170 proteins altered v 117 proteins down-regulated v 53 proteins up-regulated Gid. A Identified by MALDI-MS including Mal. E (maltose-binding protein), Ygh. A (an oxidoreductase help Salmonella survive inside macrophages), Tpx (a thiol peroxidase, help Salmonella survive within macrophages), tpx (H 2 O 2 survival) Shippy et al, 2011
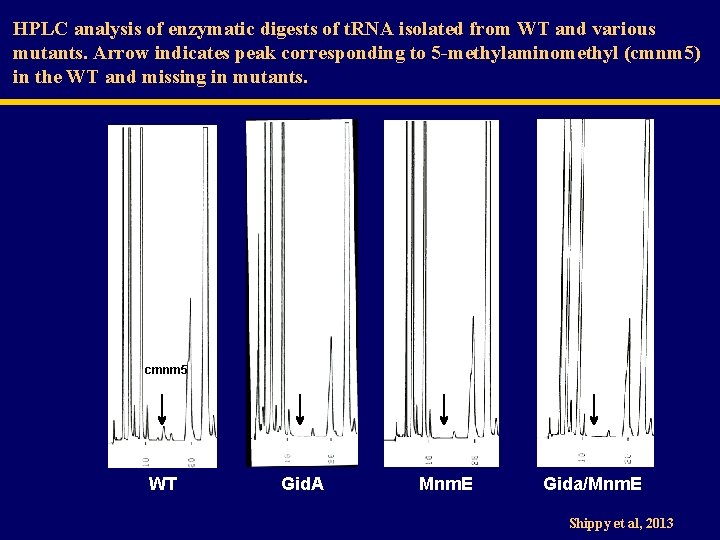
HPLC analysis of enzymatic digests of t. RNA isolated from WT and various mutants. Arrow indicates peak corresponding to 5 -methylaminomethyl (cmnm 5) in the WT and missing in mutants. cmnm 5 WT Gid. A Mnm. E Gida/Mnm. E Shippy et al, 2013
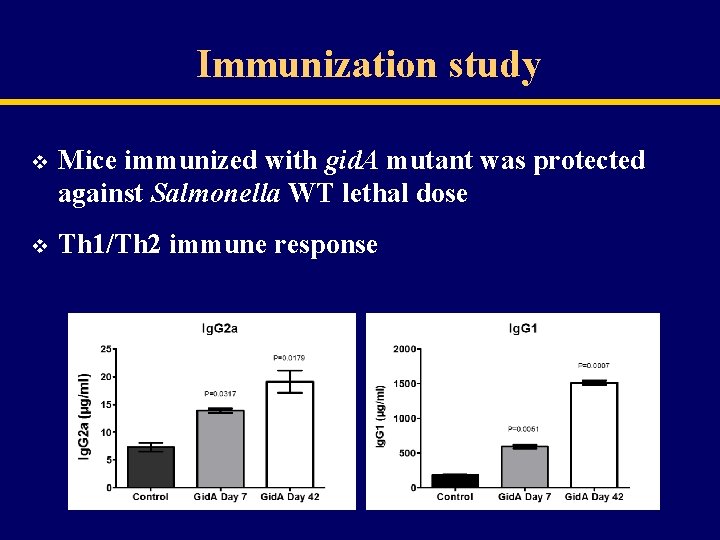
Immunization study v Mice immunized with gid. A mutant was protected against Salmonella WT lethal dose v Th 1/Th 2 immune response
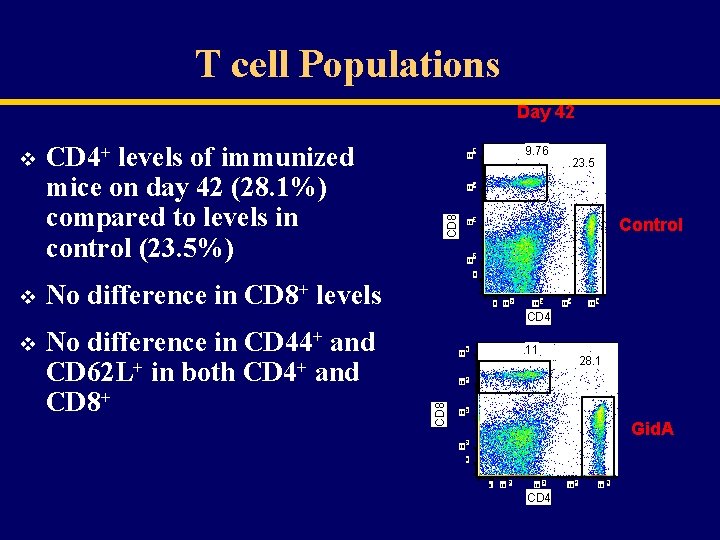
T cell Populations Day 42 CD 4+ levels of immunized mice on day 42 (28. 1%) compared to levels in control (23. 5%) 10 CD 8 v 10 4 10 3 10 9. 76 5 23. 5 Control 2 0 v No difference in CD 8+ levels 0 10 2 10 3 10 4 10 5 CD 4 No difference in CD 44+ and CD 62 L+ in both CD 4+ and CD 8+ 10 CD 8 v 11 5 10 4 10 3 28. 1 Gid. A 10 2 0 0 10 2 10 3 CD 4 10 5
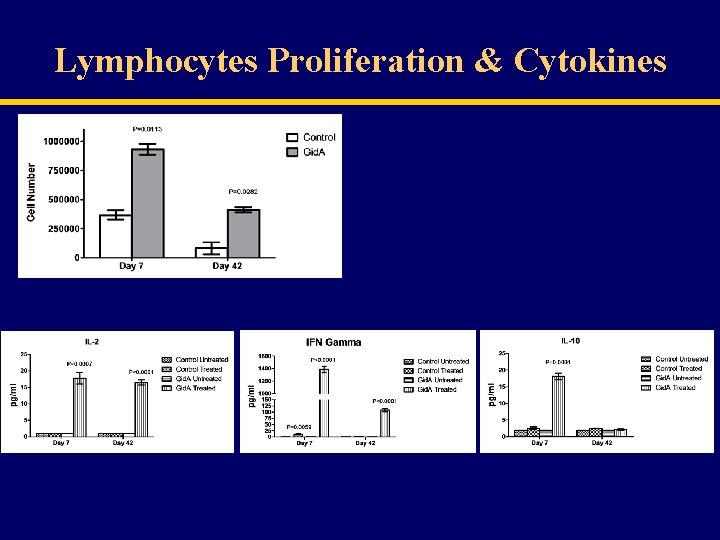
Lymphocytes Proliferation & Cytokines
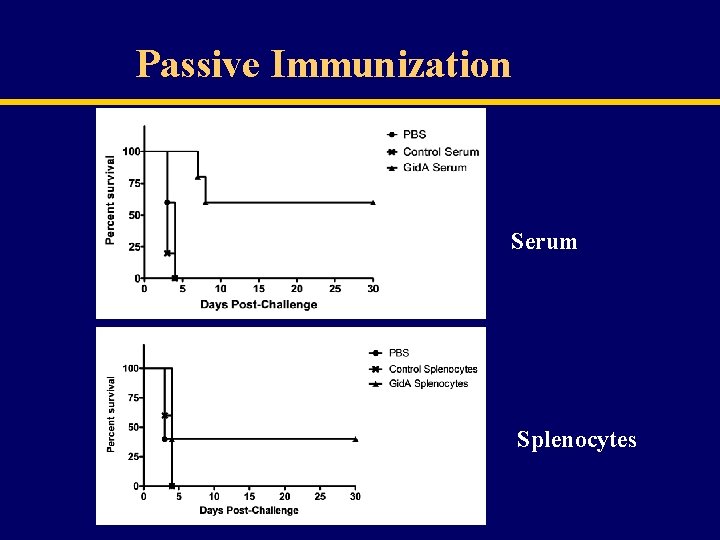
Passive Immunization Serum Splenocytes

Summary 1 v Deletion of gid. A severely affected the morphology and the virulence of Salmonella v Increase in Th 1 and Th 2 with marked level of Th 2 in the sera of immunized mice v Lymphocytes from immunized mice showed a strong response to Salmonella antigen v Passive immunization with lymphocytes and sera provided protection against lethal dose challenge

Regulation of gid. AB Operon v Increased filamentous morphology of gid. A mutant in growth media supplemented with glucose v gid. A thought to be modulated by the Asn. C v Bioinformatic analysis indicated two promoters

Transcriptional analysis of gid. AB v Real-time RT-PCR to detect gid. A and gid. B expression under different environmental conditions v gid. A and asn. C promoter activity under different conditions using lac. Z fusion assay v Effect of asn. C deletion on Gid. A expression
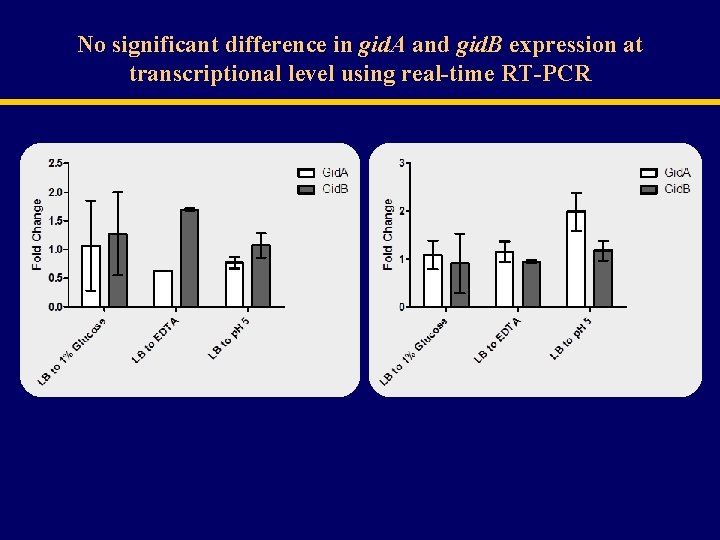
No significant difference in gid. A and gid. B expression at transcriptional level using real-time RT-PCR
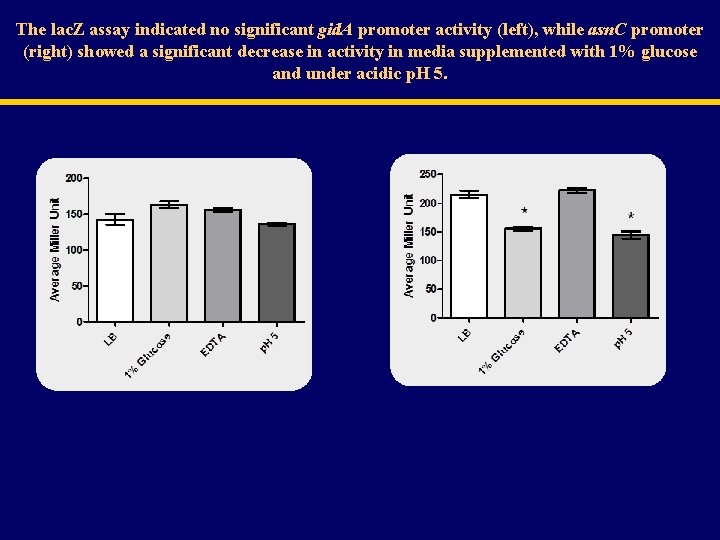
The lac. Z assay indicated no significant gid. A promoter activity (left), while asn. C promoter (right) showed a significant decrease in activity in media supplemented with 1% glucose and under acidic p. H 5.

Gid. A Expression in Salmonella Grown Under Different Conditions WT Salmonella 100 ± 1. 0 LB LB + 1% glucose 179 ± 3. 8 138 ± 5. 4 LB + 100 µM EDTA 201 ± 15. 7 LB p. H 5 Deletion of Salmonella asn. C increased Gid. A Expression
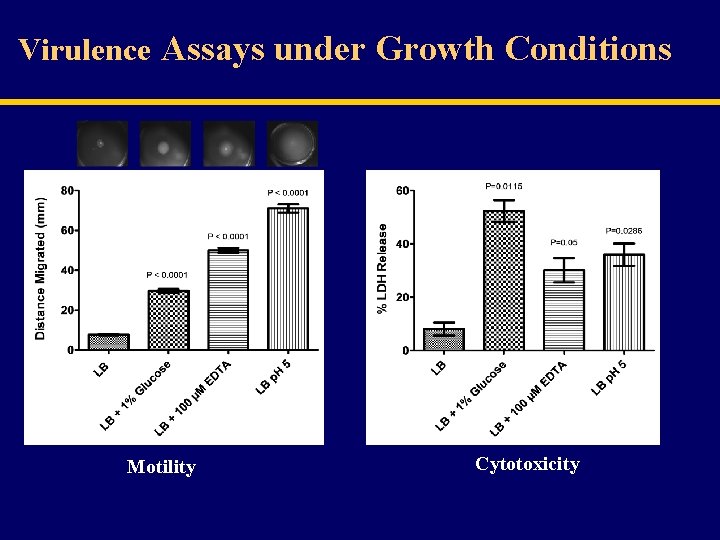
Virulence Assays under Growth Conditions Motility Cytotoxicity

Summary 2 v Transcriptional analysis, using real-time RT-PCR, indicated no significant difference in gid. A expression under various conditions. v No gid. A promoter activity, asn. C promoter showed decreased activity under glucose and acidic p. H. v Significance increase in Gid. A protein expression under different conditions and when asn. C deleted. v Suggested that Gid. A expression is modulated by environmental condition and by the Asn. C mostly at post-transcriptional level

Acknowledgements Dan Katie Nick Megan Jackie ? Dareen

Acknowledgements (Collaborators) v Ralph Albrecht, Ph. D. , Animal Sciences v Mark Cook, Ph. D. , Animal Sciences v Philip Bochsler, DVM, Ph. D. , WVDL v Ogi Okwumabua, Ph. D. , WVDL v Richard Gourse, Ph. D. , Bacteriology v Charles Lauhon, Ph. D. , School of Pharmacy v Ashok Chopra, Ph. D. , Microbiology & Immunology, UTMB

Thank you . Question…comment?

Post-transcriptional Modification of RNA: Effect on Biology and Virulence of Salmonella Amin A. Fadl University of Wisconsin-Madison
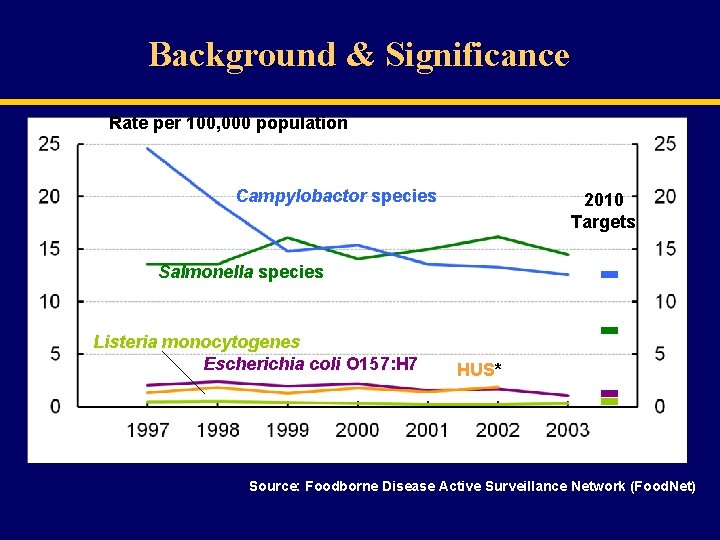
Background & Significance Rate per 100, 000 population Campylobactor species 2010 Targets Salmonella species Listeria monocytogenes Escherichia coli O 157: H 7 HUS* Source: Foodborne Disease Active Surveillance Network (Food. Net)

Background & Significance Cases Overall foodborne illness % 76, 000 Bacterial foodborne illness 4, 200, 000 5. 5 Foodborne salmonellosis 1, 400, 000 1. 7 salmonellosis from SE 194, 408 0. 25 Egg association: 40 to 80% 77, 000155, 000 < 0. 20 Out of 1. 4 million cases of salmonellosis, 95% (1. 3 million) associated with food; 20% (234, 000) from SE (about 75% associated with eggs). * Cost $23 billion (Salmonella $2. 65 billion) CDC, 2002
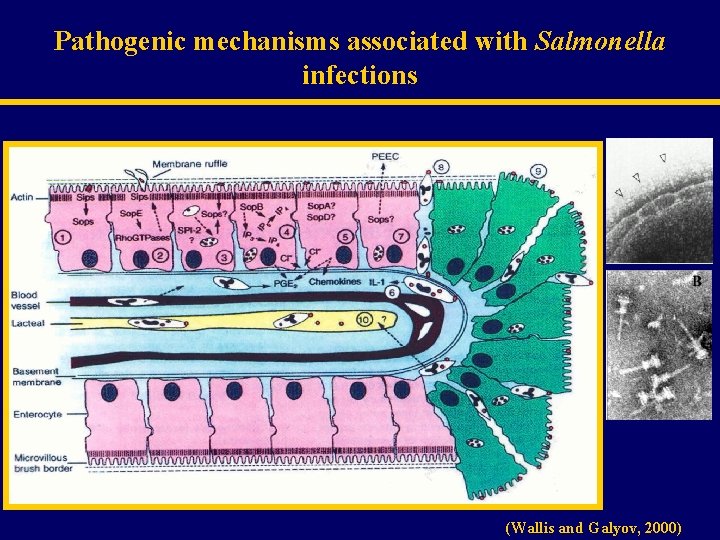
Pathogenic mechanisms associated with Salmonella infections (Wallis and Galyov, 2000)
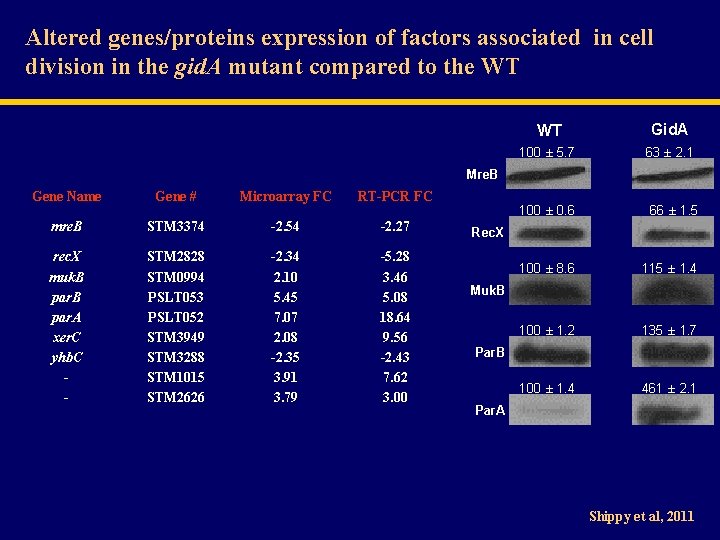
Altered genes/proteins expression of factors associated in cell division in the gid. A mutant compared to the WT WT Gid. A 100 ± 5. 7 63 ± 2. 1 Mre. B Gene Name Gene # Microarray FC RT-PCR FC mre. B STM 3374 -2. 54 -2. 27 rec. X muk. B par. A xer. C yhb. C - STM 2828 STM 0994 PSLT 053 PSLT 052 STM 3949 STM 3288 STM 1015 STM 2626 -2. 34 2. 10 5. 45 7. 07 2. 08 -2. 35 3. 91 3. 79 -5. 28 3. 46 5. 08 18. 64 9. 56 -2. 43 7. 62 3. 00 100 ± 0. 6 66 ± 1. 5 100 ± 8. 6 115 ± 1. 4 100 ± 1. 2 135 ± 1. 7 100 ± 1. 4 461 ± 2. 1 Rec. X Muk. B Par. A Shippy et al, 2011
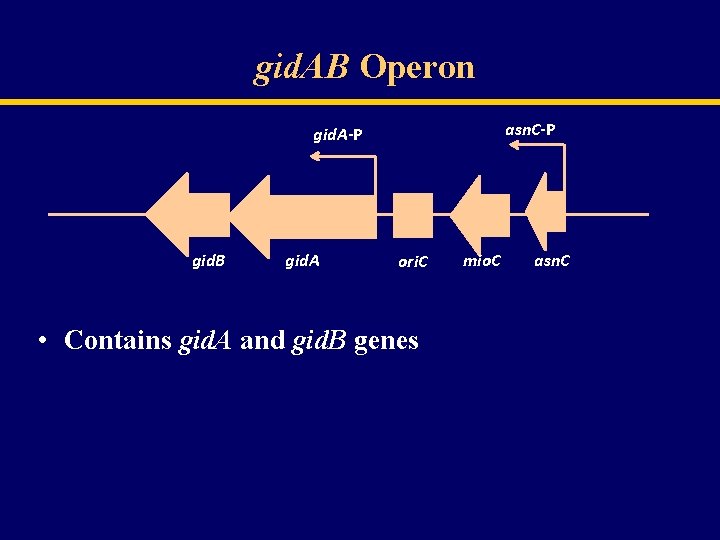
gid. AB Operon asn. C-P gid. A-P gid. B gid. A ori. C • Contains gid. A and gid. B genes mio. C asn. C

Regulation of gid. AB operon gid. B gid. A mio. C asn. C • gid. A thought to be modulated by the Asn. C
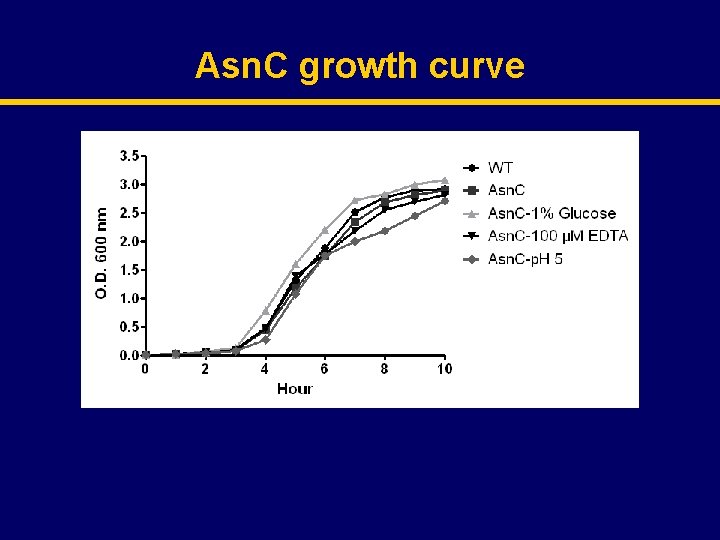
Asn. C growth curve
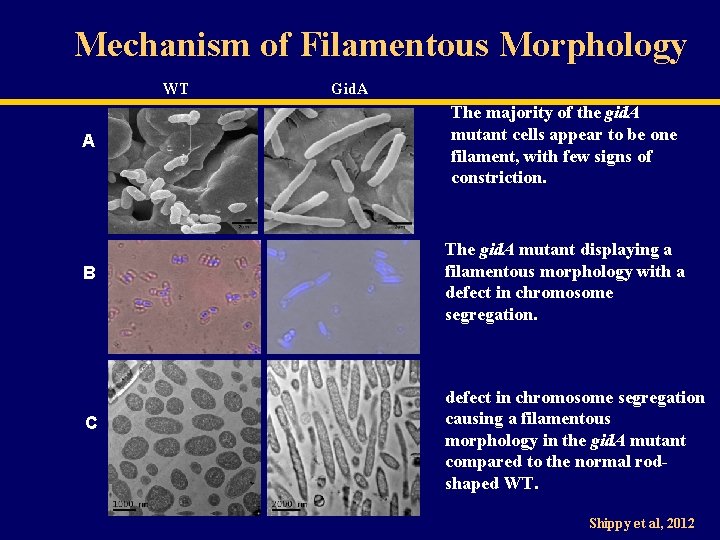
Mechanism of Filamentous Morphology WT Gid. A A The majority of the gid. A mutant cells appear to be one filament, with few signs of constriction. B The gid. A mutant displaying a filamentous morphology with a defect in chromosome segregation. C defect in chromosome segregation causing a filamentous morphology in the gid. A mutant compared to the normal rodshaped WT. Shippy et al, 2012

Summary v The gid. A mutant displayed the following phenotypes: -filamentous morphology -impaired motility, invasion of T 84 intestinal epithelial cells, cytotoxicity, limited replication in macrophages v Transcriptome and proteome analyses showed significant alterations in genes/proteins encoding for factors involved in Salmonella pathogenesis, indicating regulatory role v The gid. A mutant was attenuated in mice and animals immunized with gid. A mutant protected from lethal dose of WT Salmonella
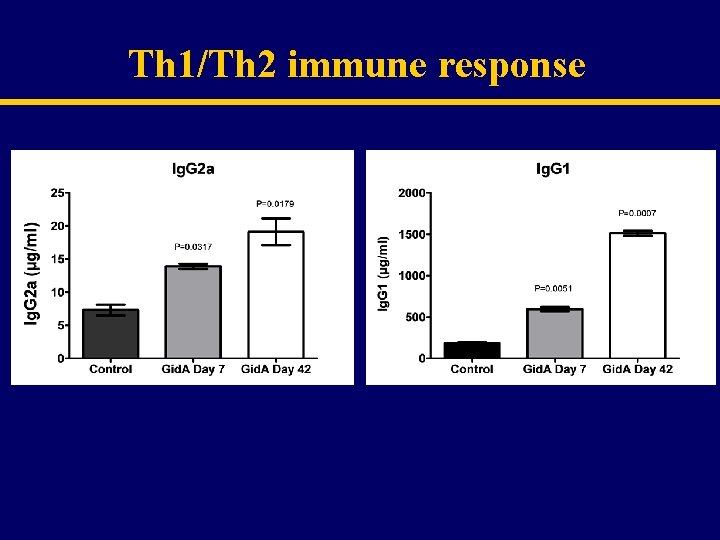
Th 1/Th 2 immune response
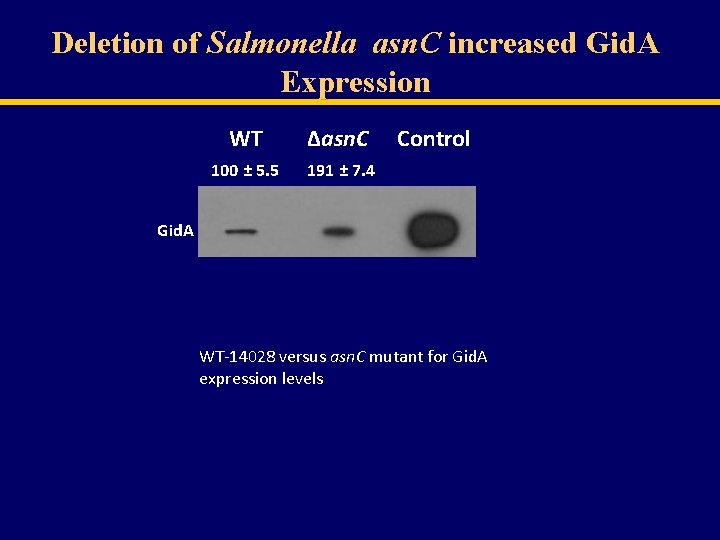
Deletion of Salmonella asn. C increased Gid. A Expression WT ∆asn. C 100 ± 5. 5 191 ± 7. 4 Control Gid. A WT-14028 versus asn. C mutant for Gid. A expression levels

Background & Significance v Model organism to study bacterial genetics and virulence. v Major cause of food-borne diseases (poultry, meat, dairy products), use as an indicator of how safe a country’s food supplies are v Multiple antibiotic-resistance strains: use in animal feed v FDA report: half of livestock and poultry feed meals and 16% of complete feeds contaminated with Salmonella
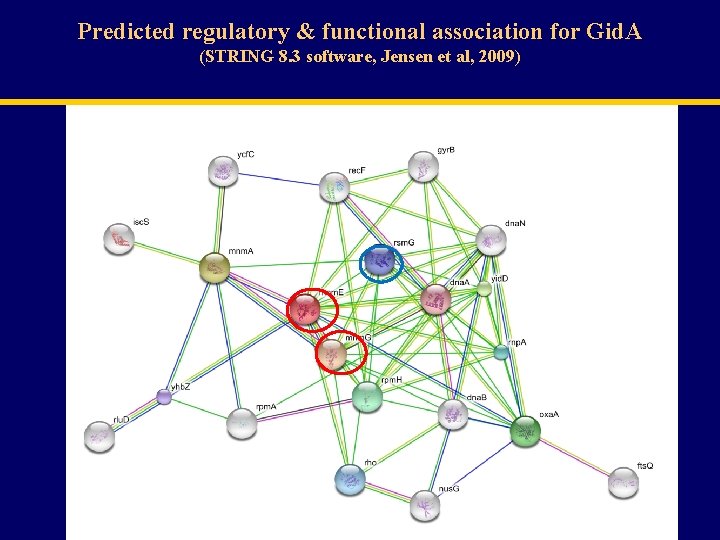
Predicted regulatory & functional association for Gid. A (STRING 8. 3 software, Jensen et al, 2009)

Effects on Biology & Virulence v Morphological changes in Aeromonas and E. coli (filamentous), Proteus mirabilis & Myxococcus (colonial). v Identified to modulate potent virulence factors: Aeromonas cytotoxic enterotoxin (Act), quorum sensing (Rhl. R expression ) in Pseudomonas, inhibition of Spe. B protease expression in S. pygenes, Shigella flexneri altered transcription regulator Vir. F
- Slides: 39