Polymerase Chain Reaction CONTENTSIntroduction PCR reaction components Mechanism















- Slides: 18

Polymerase Chain Reaction

CONTENTSIntroduction. PCR reaction components. Mechanism of basic PCR. Comparison of PCR with gene cloning. Limitations. Variants of PCR. Applications of PCR.

INTORDUCTION Polymerase chain reaction is a method of amplification of SPECIFIC DNA sequence by in vitro DNA synthesis. Polymerase chain reaction is useful because of automated instrumentation. AMPLIFICATION- ENLARGEMENT IN VITRO—UNDER CONTROLLED CONDITION PCR is a so powerful technique that from single copy of DNA molecule, millions of copies can be obtained with high accuracy , specificity and in very short time. Discovered by Kary mulis (1985), Noble prize in 1993. It is a important discovery for molecular Biology. By this technique we can get large quantities of a DNA fragment, without cloning. This technique is very suitable, where the quantity of biological specimen available is very low.

(1) Taq polymerase (From Thermophilus aquaticus Bacteria) Other DNA polymerase. Pflu DNA polymerase (From Pyrococcus furiosus) Vent DNA polymerase (From Thermococcus litoralis) (2) Oligonucleotide primer—Two 1. 17 -30 nucleotide length 2. G +C content 50% 3. Complementary the template strand. 4. Should not be complementary to each other. (3) (4) (5) (6) (7) Buffers- Tris buffer of p. H 8. 8 at 250 C gives p. H 7. 3 at 720 C. Mg+2 required by Taq enzyme activity Detergents (non ionic) (Triton x-100, Tween 80 final concentration of 0. 01%) Nucleotide – d. NTP (De oxy ribonucleotide tri phosphate) Template DNA-A short fragment of DNA. The concentration of the target sequence in the PCR reaction is generally in nanograms (5 -100 ng )

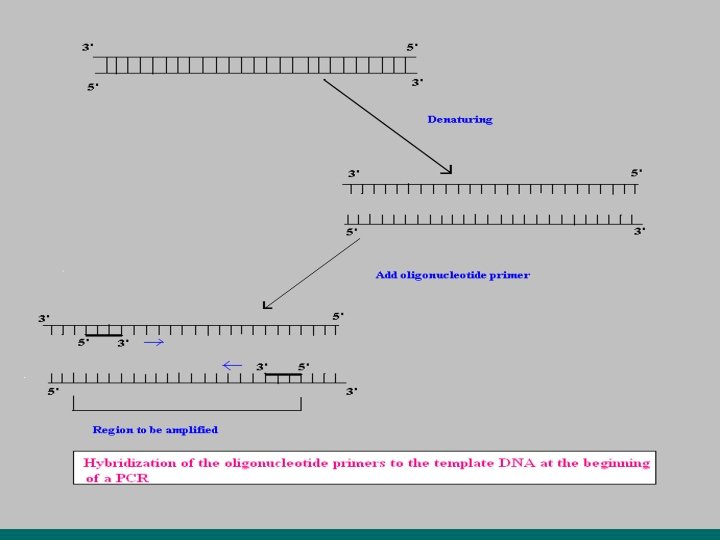
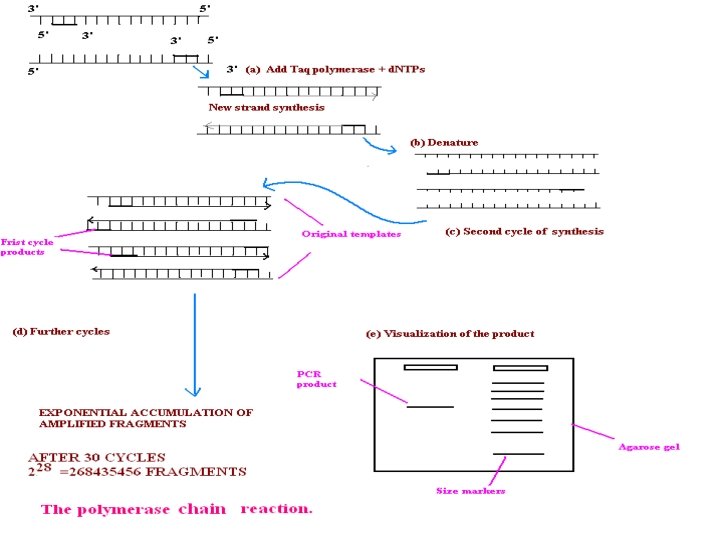
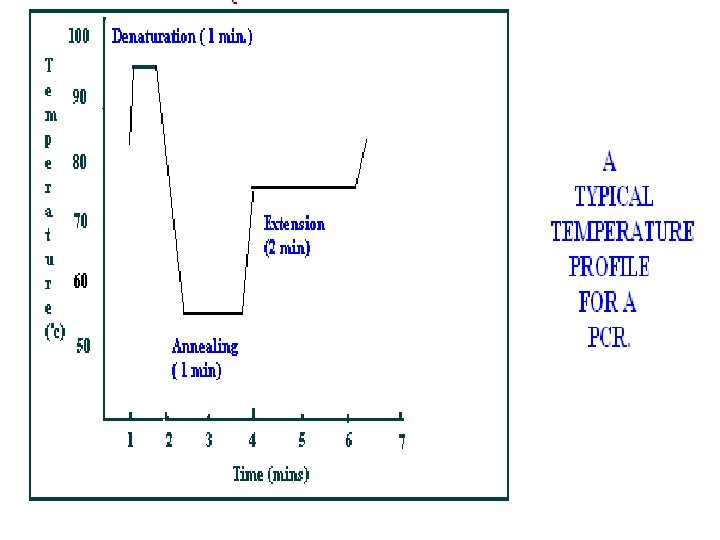
In principle PCR involves three steps to amplify a specific DNA sequence. (1) Melting /Denaturing of target DNA (denature at 940 C for 20 sec. ) (2) Anneling /Renaturing of primer (anneal at 550 C for 20 sec. ) (3) Primer extension (extend at 720 C for 30 sec. ) This three-temperature cycle is repeated 25 -50 times and the result is million fold increases of the copies of target DNA. For total 30 cycles. Overall single cycle time is 3. 75 min.

DNA preparation which have desire gene (to amplify) + + both primer in excess quantity Mg. Cl 2 + d. ATP d. TTP d. CTP d. GTP (nucleotides) + DNA polymerase Mixture in Thermocycler (perkin Elmer Cetus instrument) (A) Melting/denaturing of target DNA It is the first step to producing single strand DNA by heating up to 90 -98 0 C. (B) Anneling temperature 40 -60 0 C anneals two complementary primers to ends of separated single strands of target DNA. Primer form’s hydrogen bonds between nucleotides. (C) Primer extension Temperature 72 0 C allows the presence of thermostable “Taq DNA polymerase” enzyme and all the four essential nucleotide triphosphates allow synthesis of complementary strands in the usual manner.



It has been modified in a variety of ways to suit specific situations and applications. Some important variations of PCR are as follows. Inverse PCR- This PCR can be use to amplify the sequences flanking a segment the border sequences of which are known. This is done by using as primers complementary to the 5’ ends of the segment in question. This orients the free 3’OH of the primers outward of the sequence, as a result, the newly synthesized chain grows away from the border of the concerned segment.

Anchored PCR- When sequence of only one end of the desired segment is known, the primer complementary to the 3’strand or this end is used to produce several copies of only one strand of the gene. Now a poly-G tail is added to the 3’ends of the single-strand DNA copies produced by PCR. This allows the use of complementary poly-C, to be used as primer for copying the DNA single strands generated by PCR, and gives rise to the complete DNA duplex that can be amplified.

INTERSEQUENCE – SPECIFIC (ISSR) PCR- A PCR method for DNA fingerprinting that amplifies regions between some simple sequence repeats to produce a unique fingerprint of amplified fragment lengths

Asymmetric PCRThis PCR can be used to generate single-strand copies of a DNA sequence. • Quantities of the two primers for the 3’end of the segment in the reaction mixture (100: 1 ratio) • Use for DNA sequencing.


(1) Contamination of the reaction mixture of PCR even by a single gene can result in to impure product of PCR reaction. (2) PCR is most useful for the amplification of DNA segments less than 2 kb in length. (3) Base substitution errors by Taq polymerase are 1 per 9000 bp and frame shifts occur 1 per 40000. These are significant if PCR extends over 30 cycles when error occurs in every 300 bp of product. High error rate of Taq is because it lacks 3’-5’ exonuclease activity.

Comparison of PCR and gene cloning. Parameter Final result PCR Gene cloning Selective amplification of specific sequence. In vitro & in vivo Nanogram (ng) Microgram DNA Restriction enzyme, ligase, vector, bacterial cell Automation Yes No Labour intensive No Yes Error probability Less More Applications More Less Cost Less More Time for a typical experiment Four hours Days, weeks or even months Manipulation Concentration of starting material Biological reagents required The amplified DNA product is detected by agarose gel electrophoresis, followed by staining with Ethidium bromide. and sequencing of amplified DNA. Insertions of amplified DNA fragment in to traditional cloning vector if expression is desired.

(1. ) Diagnosis of genetic diseases. (2. ) DNA fingerprinting. (3. ) For research purpose. (4. ) For molecular mapping. (5. ) For preparation of molecular markers. (6. ) Study of DNA polymorphism. (7. ) Determine the sex of embryos. (Using Y chromosome-specific primers) (8. ) PCR can be used to detect the presence of a gene transferred in to an organism (transgene) by using the end sequences of the transgene for amplification of DNA from the transgenic organism.

REFERENCES(1) (2) (3) (4) (5) (6) Winnacker, Ernst-L –Introduction to Gene Technology. T. A. BROWN – Gene cloning and DNA analysis. H. S. Chawla –Plant biotechnology. B. D. Singh – Biotechnology. WWW. GOOGLE. COM. Bartlett & Stirling(2003)- A short History of the PCR In : Methods Mol. Biol. (7) Chemical Synthesis, Sequencing. &Amplification of DNA. Arizona state university Retrieved on 29 october 2007.

Thank s