Pneumonia Microbiology of Pneumonia J Matthew Velkey Ph
Pneumonia Microbiology of Pneumonia J. Matthew Velkey, Ph. D Department of Cell Biology Duke University School of Medicine Andrew Alspaugh, MD Department of Medicine Infectious Disease Division Duke University School of Medicine 1

Learning Objectives • Be aware of the spectrum of pathogens associated with pneumonia • Identify common bacterial causes and microbiological features of pneumonia in the following contexts: • community-acquired • unique populations: • aspiration • obstructive disease (COPD, CF) • immunocompromise • healthcare associated pneumonia 2
Common Pathogens Associated with Pneumonia Bacterial Viral Fungal 3
Bacteria Associated with Pneumonia “typical” “atypical” OTHERS: aspiration obstruction HCAP 4

Pathogens Causing Community-Acquired Pneumonia • “Typical” = easily cultured – Streptococcus pneumoniae – Haemophilus influenzae • “Atypical” = not culturable by standard methods – Mycoplasma pneumoniae – Legionella pneumophila – Chlamydophila (Chlamydia) pneumoniae 5
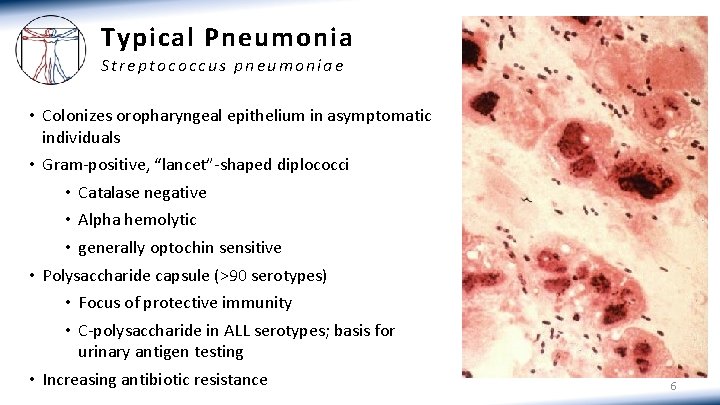
Typical Pneumonia Streptococcus pneumoniae • Colonizes oropharyngeal epithelium in asymptomatic individuals • Gram-positive, “lancet”-shaped diplococci • Catalase negative • Alpha hemolytic • generally optochin sensitive • Polysaccharide capsule (>90 serotypes) • Focus of protective immunity • C-polysaccharide in ALL serotypes; basis for urinary antigen testing • Increasing antibiotic resistance 6
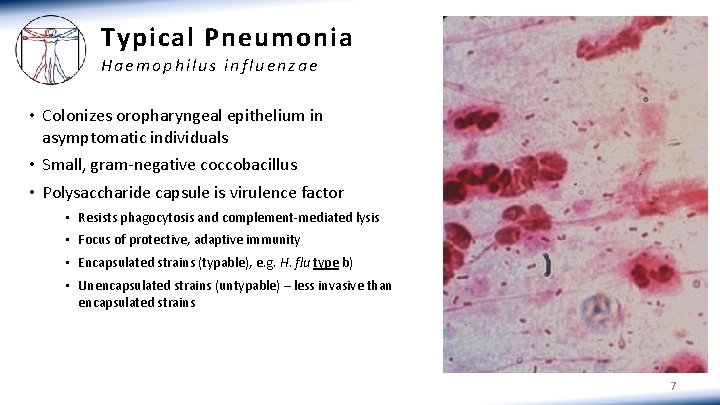
Typical Pneumonia Haemophilus influenzae • Colonizes oropharyngeal epithelium in asymptomatic individuals • Small, gram-negative coccobacillus • Polysaccharide capsule is virulence factor • Resists phagocytosis and complement-mediated lysis • Focus of protective, adaptive immunity • Encapsulated strains (typable), e. g. H. flu type b) • Unencapsulated strains (untypable) – less invasive than encapsulated strains 7

Atypical Pneumonia Mycoplasma pneumoniae • Smallest self-replicating organism • “Fried egg” colony on growth medium • culture rarely performed • emerging molecular tests • empiric treatment • Bacterium that lacks peptidoglycan • Therefore resistant to many cell wall-targeting antibiotics • Pathogenesis • • Attachment to respiratory epithelium Entry into cell Intracellular replication Host cell lysis • Association with “cold agglutinin” syndrome Mycoplasma colonies reveal a characteristic “fried egg appearance – Note that cultures are rarely used in most clinical settings 8
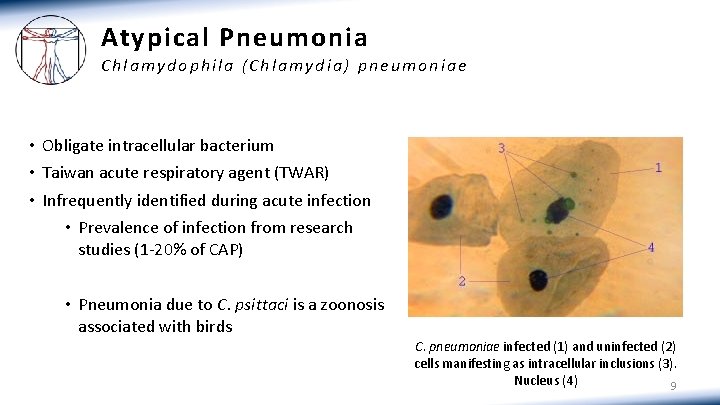
Atypical Pneumonia Chlamydophila (Chlamydia) pneumoniae • Obligate intracellular bacterium • Taiwan acute respiratory agent (TWAR) • Infrequently identified during acute infection • Prevalence of infection from research studies (1 -20% of CAP) • Pneumonia due to C. psittaci is a zoonosis associated with birds C. pneumoniae infected (1) and uninfected (2) cells manifesting as intracellular inclusions (3). Nucleus (4) 9
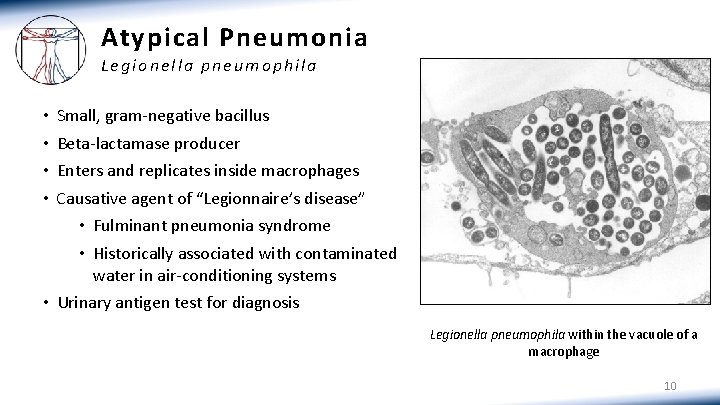
Atypical Pneumonia Legionella pneumophila • • Small, gram-negative bacillus Beta-lactamase producer Enters and replicates inside macrophages Causative agent of “Legionnaire’s disease” • Fulminant pneumonia syndrome • Historically associated with contaminated water in air-conditioning systems • Urinary antigen test for diagnosis Legionella pneumophila within the vacuole of a macrophage 10

Pneumonia in predisposing conditions Aspiration Aerobic Gram-negative rods: Klebsiella Actinetobacter Enterobacter Pseudomonas Oro-pharyngeal flora: Staph aureus E. coli Streptococcus spp. Obstruction (COPD, CF) Aerobic Gram-negative rods, especially Pseudomonas Immunocompromised host • Typical and Atypical CAP pathogens and enterics, esp. Pseudomonas • Fungi: Pneumocystis, Aspergillus, Cryptococcus, Histoplasma • Viral: respiratory viruses (RSV, adeno-, metapneumo- influenza) and CMV • Helminth: Strongyloides 11
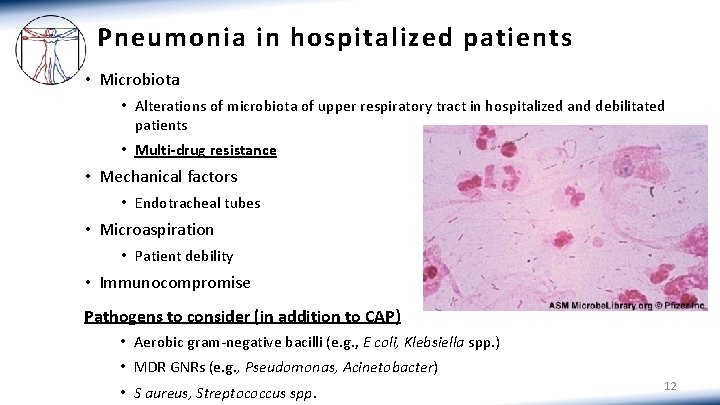
Pneumonia in hospitalized patients • Microbiota • Alterations of microbiota of upper respiratory tract in hospitalized and debilitated patients • Multi-drug resistance • Mechanical factors • Endotracheal tubes • Microaspiration • Patient debility • Immunocompromise Pathogens to consider (in addition to CAP) • Aerobic gram-negative bacilli (e. g. , E coli, Klebsiella spp. ) • MDR GNRs (e. g. , Pseudomonas, Acinetobacter) • S aureus, Streptococcus spp. 12
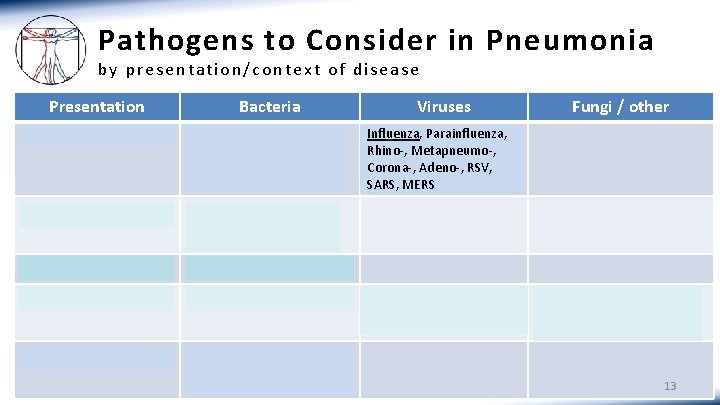
Pathogens to Consider in Pneumonia by presentation/context of disease Presentation Bacteria Community-acquired S. pneumo, H. influenzae, Mycoplasma, Chlamydophila, Legionella Aspiration Enterics (Gram neg rods), upper airway flora (Staph and other Strep spp. ) Obstruction Enterics, esp. Pseudomonas Immunocompromise CAP pathogens and enterics Healthcare-associated CAP pathogens and enterics, esp. Pseudomonas; Staph; Strep spp. Viruses Fungi / other Influenza, Parainfluenza, Rhino-, Metapneumo-, Corona-, Adeno-, RSV, SARS, MERS Respiratory viruses above and CMV Pneumocystis, Aspergillus, Cryptococcus, Histoplasma, Strongyloides (helminth) 13

Pathogens to Consider in Pneumonia by age Age Bacteria Viruses Neonates Group B streptococci Escherichia coli RSV Infants Chlamydia trachomatis Streptococcus pneumoniae RSV parainfluenza Children S. pneumoniae Haemophilus influenzae RSV parainfluenza Young adults Mycoplasma pneumoniae Chlamydophila pneumoniae S. pneumoniae Various respiratory viruses (e. g. Adenovirus) Older adults S. pneumoniae H. influenzae Legionella pneumophila Influenza 15

C r e d i t s : M i c r o b i o l o g y o f Pneumonia Location of item (slide #3 & 4): Pathogen map. RWJF Re-imagining Medical Education Project. Location of item (slide #6): Gram Stain of a film of sputum from a case of lobar pneumonia. CDC Location of item (slide #7): Haemophilus influenzae sputum sample. The American Society for Microbiology http: //www. microbelibrary. org/images/buxton/03 spt neg diplo/haemophilus influenzae fig 1. jpg Location of item (slide #8): L Young et al. “Detection of Mycoplasma in cell cultures”. Nature Protocols 5, 929 - 934 (2010) Location of item (slide #9): en. wikipedia. org/”Micrograph of. Chlamydophila(Chlamydia) pneumoniae in an epithelial cell in acute bronchitis”: 1 – infected epitheliocyte, 2 – uninfected epitheliocytes, 3 – chlamydial inclusion bodies in cell, 4 – cell nuclei Location of item (slide #10): “Legionella pneumophila”: commons. wikimedia. org Location of item (slide #12): Pseudomonas sputum sample. The American Society for Microbiology http: //www. microbelibrary. org/images/buxton/04 spt neg bacilli/pseudomonas aeruginosa fig 3. jpg. 16
- Slides: 15