PML Toward a HighLevel Formal Language for Biological

PML: Toward a High-Level Formal Language for Biological Systems Bor-Yuh Evan Chang and Manu Sridharan July 24, 2003

Why Formal Models for Biology? • Experiments have led to an enormous wealth of (detailed) knowledge but in a fragmented form – serve as a common language for sharing • modular, compositional, varying levels of abstraction • Much information described through prose or graph-like diagrams with loose semantics – make assumptions explicit /24/2003 2

Why Formal Models for Biology? • Mathematical abstraction convenient for reasoning and simulation – DNA ! string over the alphabet {A, C, G, T} • enables the use of string comparison algorithms – Cellular Pathways ! ? /24/2003 3

Previous Abstractions • Chemical kinetic models – can derive differential equations – well-studied, with considerable theoretical basis – variables do not directly correspond with biological entities – may become difficult to see how multiple equations relate to each other /24/2003 4
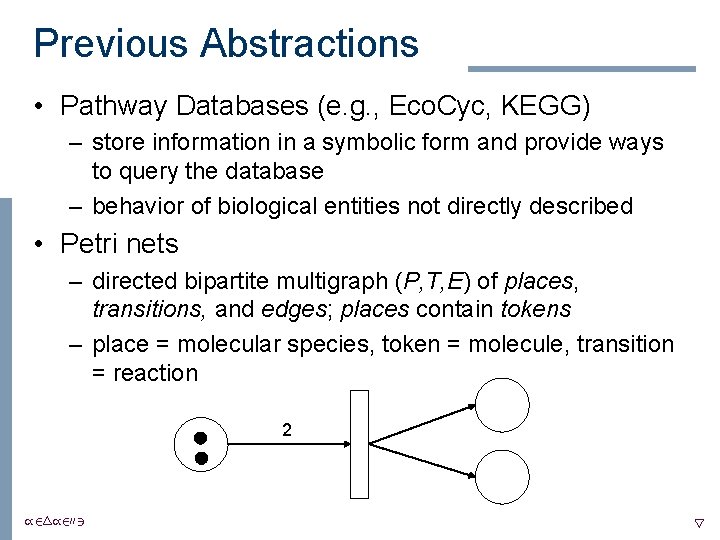
Previous Abstractions • Pathway Databases (e. g. , Eco. Cyc, KEGG) – store information in a symbolic form and provide ways to query the database – behavior of biological entities not directly described • Petri nets – directed bipartite multigraph (P, T, E) of places, transitions, and edges; places contain tokens – place = molecular species, token = molecule, transition = reaction 2 /24/2003 5

Previous Abstractions • Concurrent computational processes – each biological entity is a process that may carry some state and interacts with other processes – each process described by a “program” – prior proposals based on process algebras, such as the -calculus [Regev et al. ’ 01] – we take this view /24/2003

Computer Systems vs. Biological Processes • Similarities – elementary pieces build-up components that in turn build-up large components and so forth to create highly complex systems – all systems seem to have similar cores but exhibit great diversity • Differences! – theory of computation and computer systems are purely man-made (controlled-design) but biology is observational /24/2003

Model of Concurrent Computation • Must choose a machine model as a basis – The -calculus [Milner ’ 90 and others] • A formalism aimed at capturing the essence of concurrent computation. – focuses on communication by message passing • System composed of processes • Communication on channels – send: – receive: send message m on channel c receive message on channel c, call it x – Many variants—the stochastic -calculus /24/2003 8

The -calculus • Syntax • Operational Semantics /24/2003 9

The -calculus • Congruence /24/2003 10

Modeling in the -calculus • The -calculus is concise and compact, yet powerful – not clear if another machine model would be particularly better or worse • However, it is far too low-level for direct modeling (ad-hoc structuring) /24/2003 11
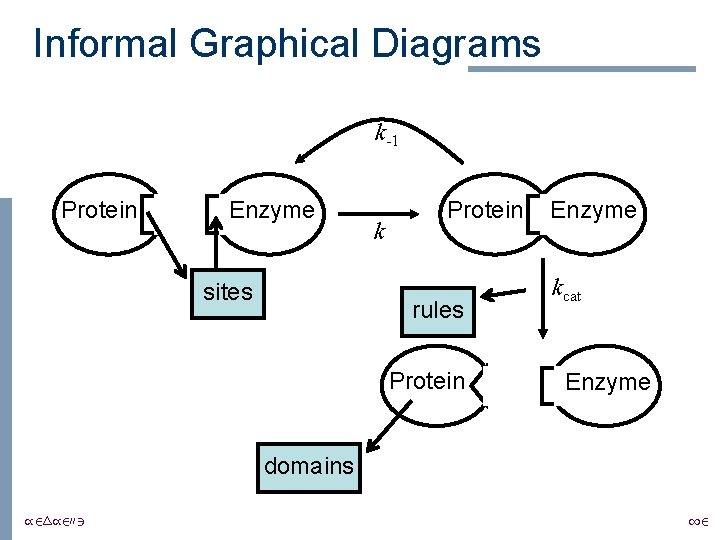
Informal Graphical Diagrams k-1 Protein Enzyme sites k Protein rules Protein Enzyme kcat Enzyme domains /24/2003 12
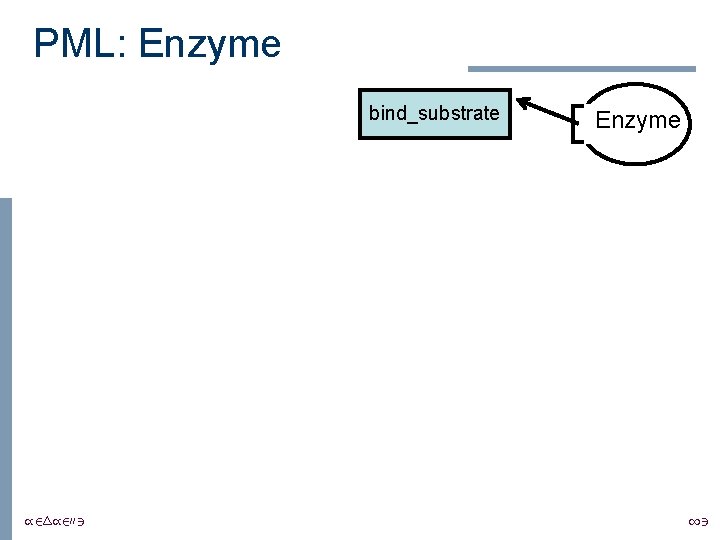
PML: Enzyme bind_substrate /24/2003 Enzyme 13
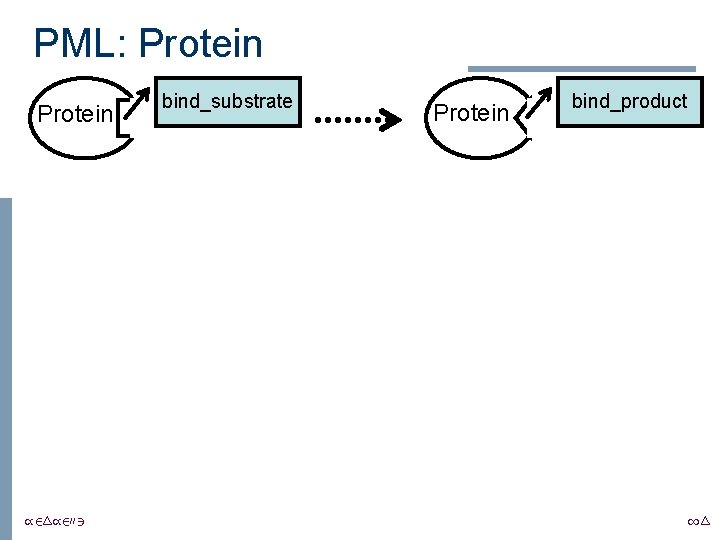
PML: Protein /24/2003 bind_substrate Protein bind_product 14

PML: A Simple System /24/2003 15

Compartments • Critical part of biological pathways – prevents interactions that would otherwise occur • Description of the behavior of a molecule should not depend on the compartment • Regev et al. use “private” channels in the calculus for both complexing and compartmentalization /24/2003 1
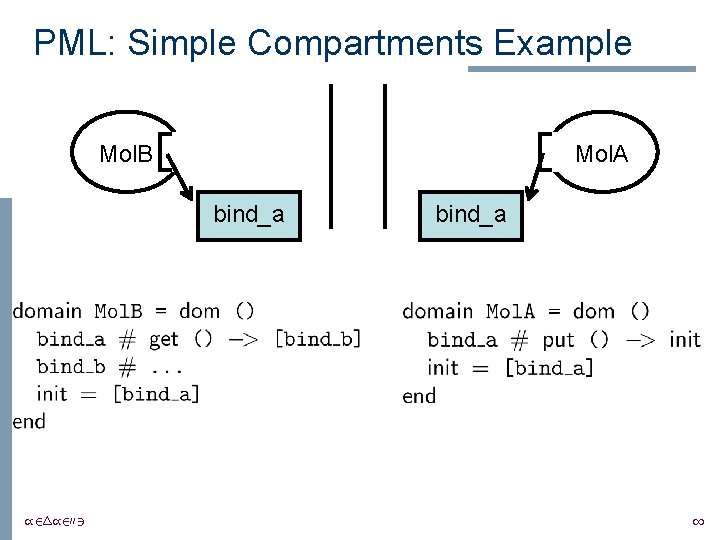
PML: Simple Compartments Example Mol. B Mol. A bind_a /24/2003 bind_a 1
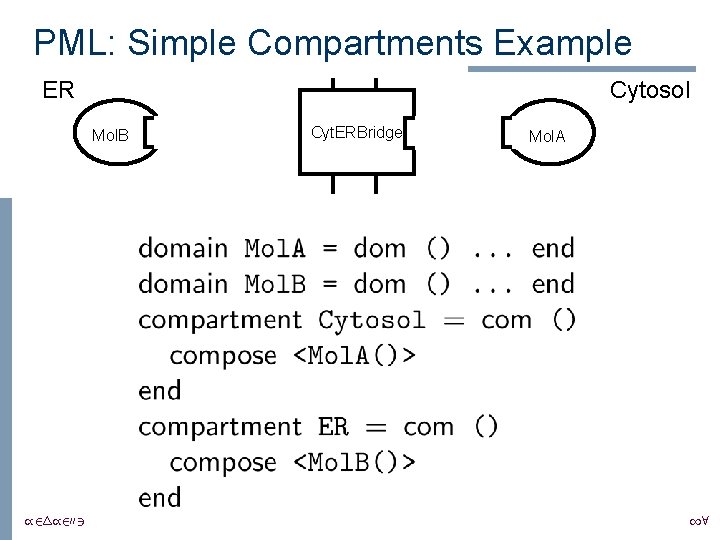
PML: Simple Compartments Example ER Cytosol Mol. B /24/2003 Cyt. ERBridge Mol. A 18

PML: Simple Compartments Example ER Cytosol Mol. B /24/2003 Cyt. ERBridge Mol. A 19

Semantics of PML • Defined in terms of the -calculus via two translations – from PML to Core. PML • “flattens” compartments, removes bridges, explicit rule names /24/2003 20

Semantics of PML – from Core. PML to the -calculus /24/2003 21

Larger Models • Modeled a general description of ER cotranslational-translocation – unclearly or incompletely specified aspects became apparent • e. g. , can the signal sequence and translocon bind without SRP? Yes [Herskovits and Bibi ’ 00] • Extended to model targeting ER membrane with minor modifications /24/2003 22

Benefits of PML • Easier to write and understand because of more consistent biological metaphor (binding sites) • Block structure for controlling namespace and modularity • Special syntax for compartments – separate complexing from compartmentalization /24/2003 23

Future Work • Naming? • Proximity of molecules • Integrating quantitative information (reaction rates, etc. ) – start from work by Priami et al. • Type systems • Graphical and simulation tools /24/2003 24


Example: Cotranslational Translocation • Ribosome translates m. RNA exposing a signal sequence • Signal sequence attracts SRP stopping translation • SRP receptor (on ER membrane) attracts SRP • Signal sequence interacts with translocon, SRP disassociates resuming translation • Signal peptidase cleaves the signal sequence in the ER lumen, Hsc 70 chaperones aid in protein folding /24/2003 2

Example: Cotranslational Translocation /24/2003 2

Example: Cotranslational Translocation /24/2003 28

Example: Cotranslational Translocation /24/2003 29

Example: Cotranslational Translocation /24/2003 30

Example: Cotranslational Translocation /24/2003 31

Example: Cotranslational Translocation /24/2003 32

Example: Cotranslational Translocation /24/2003 33
- Slides: 33