Plant Proteomics Recent Surprises Deriving from Subcellular Proteomic

Plant Proteomics: Recent Surprises Deriving from Subcellular Proteomic Analyses Targeted to Chloroplasts and their Subcompartments 3 rd Plant Genomics Congress Europe 11 th-12 th May 2015 Laboratoire de Physiologie Cellulaire & Végétale Laboratoire de Biologie à Grande Echelle

From almost nothing to almost everything Oxygen The chloroplast Light ATP, NADPH Amino acids Fatty acids Starch Phosphate Sulfate Nitrite Sugars Hormones Pigments (chlorophylls, carotenoids) Vitamins (prenylquinones…) H 2 O CO 2 Nuleobases Galactolipids Hydrogen … The chloroplast, the energy factory of the plant cell and the place of lipids, polymers of sugars, hydrogen. . . synthesis
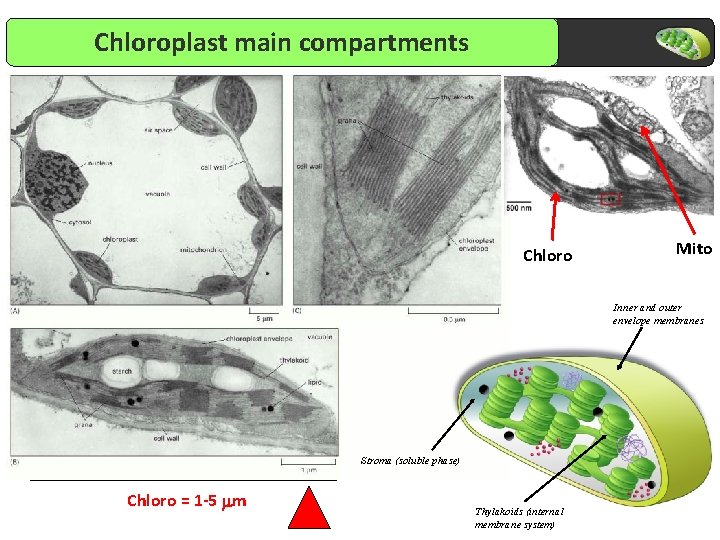
Chloroplast main compartments Chloro Mito Inner and outer envelope membranes Stroma (soluble phase) Chloro = 1 -5 mm Thylakoids (internal membrane system)
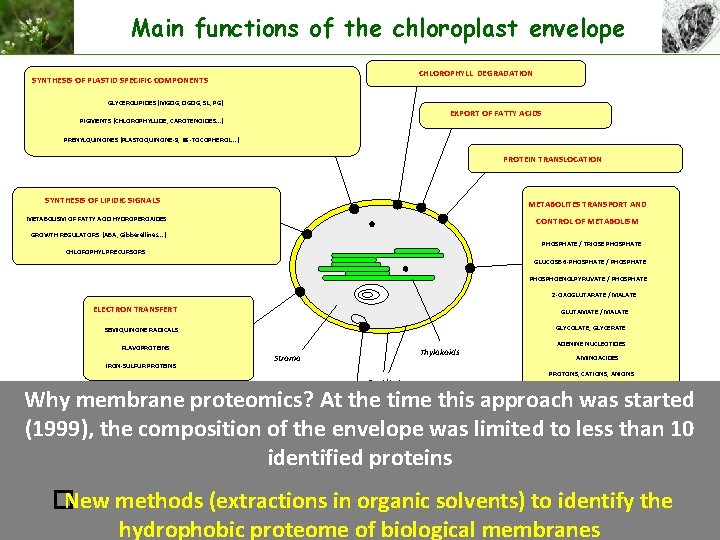
Main functions of the chloroplast envelope CHLOROPHYLL DEGRADATION SYNTHESIS OF PLASTID SPECIFIC COMPONENTS GLYCEROLIPIDES (MGDG, DGDG, SL, PG) EXPORT OF FATTY ACIDS PIGMENTS (CHLOROPHYLLIDE, CAROTENOIDES. . . ) PRENYLQUINONES (PLASTOQUINONE-9, a -TOCOPHEROL. . . ) PROTEIN TRANSLOCATION SYNTHESIS OF LIPIDIC SIGNALS METABOLITES TRANSPORT AND METABOLISM OF FATTY ACID HYDROPEROXIDES CONTROL OF METABOLISM GROWTH REGULATORS (ABA, Gibberellines. . . ) PHOSPHATE / TRIOSE PHOSPHATE CHLOROPHYL PRECURSORS GLUCOSE-6 -PHOSPHATE / PHOSPHATE PHOSPHOENOLPYRUVATE / PHOSPHATE 2 -OXOGLUTARATE / MALATE ELECTRON TRANSFERT GLUTAMATE / MALATE GLYCOLATE, GLYCERATE SEMIQUINONE RADICALS FLAVOPROTEINS IRON-SULFUR PROTEINS Thylakoids Stroma Envelope ADENINE NUCLEOTIDES AMINOACIDES PROTONS, CATIONS, ANIONS Why membrane proteomics? At the time this approach was started (1999), the composition of the envelope was limited to less than 10 identified proteins COORDINATION OF NUCLEAR AND PLASTID GENOME EXPRESSION INTERACTIONS with cp. DNA : DNA-binding protein, Topoisomerase II INTERACTIONS with TRANSLATION APPARATUS : m. RNA stabilization factors, Ribosome Recycling Factor �New methods (extractions in organic solvents) to identify the Review: Joyard et al. (1998) Plant Physiol. Block et al. , (2007) Photosynthes. Res. hydrophobic proteome of biological membranes
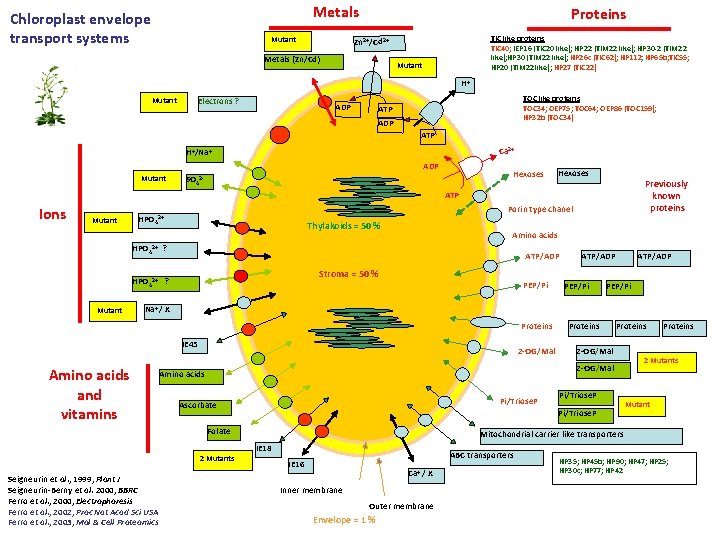
Metals Chloroplast envelope transport systems Mutant Proteins TIC like proteins TIC 40; IEP 16 (TIC 20 like); HP 22 (TIM 22 like); HP 30 -2 (TIM 22 like); HP 30 (TIM 22 like); HP 26 c (TIC 62); HP 112; HP 65 b; TIC 55; HP 20 (TIM 22 like); HP 27 (TIC 22) Zn 2+/Cd 2+ Metals (Zn/Cd) Mutant H+ Mutant Electrons ? ADP TOC like proteins TOC 34; OEP 75; TOC 64; OEP 86 (TOC 159); HP 32 b (TOC 34) ATP ADP ATP Ca 2+ H+/Na+ ADP Mutant Hexoses SO 42 - Hexoses Previously known proteins ATP Ions Mutant Porin type chanel HPO 42+ Thylakoids = 50 % Amino acids HPO 42+ ? ATP/ADP Stroma = 50 % HPO 42+ ? Mutant PEP/Pi ATP/ADP PEP/Pi Na+ / X Proteins IE 45 Amino acids and vitamins 2 -OG/Mal Proteins 2 -OG/Mal Amino acids Pi/Triose. P Ascorbate Proteins 2 Mutants Mutant Mitochondrial carrier like transporters IE 18 2 Mutants Pi/Triose. P Folate Seigneurin et al. , 1999, Plant J Seigneurin-Berny et al. 2000, BBRC Ferro et al. , 2000, Electrophoresis Ferro et al. , 2002, Proc Nat Acad Sci USA Ferro et al. , 2003, Mol & Cell Proteomics Proteins ABC transporters IE 16 Ca+ / X Inner membrane Outer membrane Envelope = 1 % HP 35; HP 45 b; HP 90; HP 47; HP 25; HP 30 c; HP 77; HP 42

Back to large-scale proteomics Independent studies targeted to chloroplast subcompartments (envelope, thylakoids and stroma) have revealed lots of redundancy: cross contaminations or multiple localizations? Project: Revisiting the chloroplast proteome through subplastidial localization of more than 1000 proteins Aim: chloroplast envelope proteome
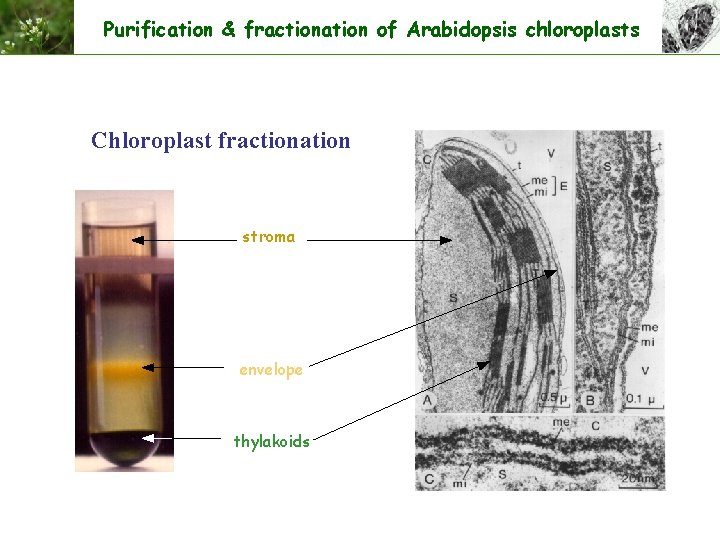
Purification & fractionation of Arabidopsis chloroplasts Chloroplast fractionation stroma envelope thylakoids
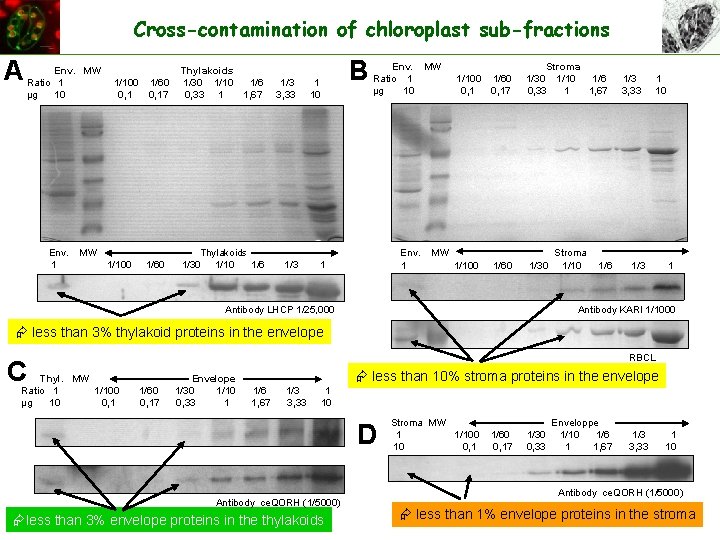
Cross-contamination of chloroplast sub-fractions A Env. MW Thylakoids Ratio 1 1/100 1/60 1/30 1/10 1/6 1/3 1 µg 10 0, 17 0, 33 1 1, 67 3, 33 10 B Env. MW Stroma Ratio 1 1/100 1/60 1/30 1/10 1/6 1/3 1 µg 10 0, 17 0, 33 1 1, 67 3, 33 10 Env. MW Stroma Env. MW Thylakoids 1 1/100 1/60 1/30 1/10 1/6 1/3 1 Antibody KARI 1/1000 Antibody LHCP 1/25, 000 less than 3% thylakoid proteins in the envelope RBCL C Thyl. MW Envelope Ratio 1 1/100 1/60 1/30 1/10 1/6 1/3 1 µg 10 0, 17 0, 33 1 1, 67 3, 33 10 less than 10% stroma proteins in the envelope D Stroma MW Enveloppe 1 1/100 1/60 1/30 1/10 1/6 1/3 1 10 0, 17 0, 33 1 1, 67 3, 33 10 Antibody HMA 1 (1/1000) Antibody ce. QORH (1/5000) less than 3% envelope proteins in the thylakoids Antibody ce. QORH (1/5000) less than 1% envelope proteins in the stroma

Subplastidial proteomics (collaborations LPCV & EDy. P) AT_CHLORO database V 1 http: //www. grenoble. prabi. fr/at_chloro ENV STR § 5 years of work § 228 samples analyzed by HPLC-MS/MS (most of them 2 to 4 times): 494 analysis THY � creation of a database of 12441 distinct peptides corresponding to 1325 validated proteins (i. e. 2 peptides with Mascot score > 30) - >700 proteins in envelope fractions (460> & 300 only). ENV 318 - Experimental data for subplastidial localization of 1300 proteins 94 235 69 13 134 THY 133 - Curated annotation of the chloroplast envelope components - Revisiting the plastidial localisation of metabolic pathways - Identification of new proteins with “non-canonical” targeting peptides STR Joyard et al. , Mol Plant 2009 Ferro et al. , Mol Cell Proteomics 2010 Joyard et al. , Prog Lipid Res 2010 Joshi et al. , Plant Physiol 2011 Agrawal et al. , Mass Spectrom Rev 2011 Bruley et al. , Frontiers Plant Sci 2012 Rolland et al. , Annual Rev Genet 2012 Dell’Aglio et al. , Mol Bio. Syst 2013 Axe 2 Labex GRAL 2012 -2015 ANR Génoplante 2007 -2010

Application of chloroplast proteomics 1 -Revisiting the plastidial localisation of metabolic pathways
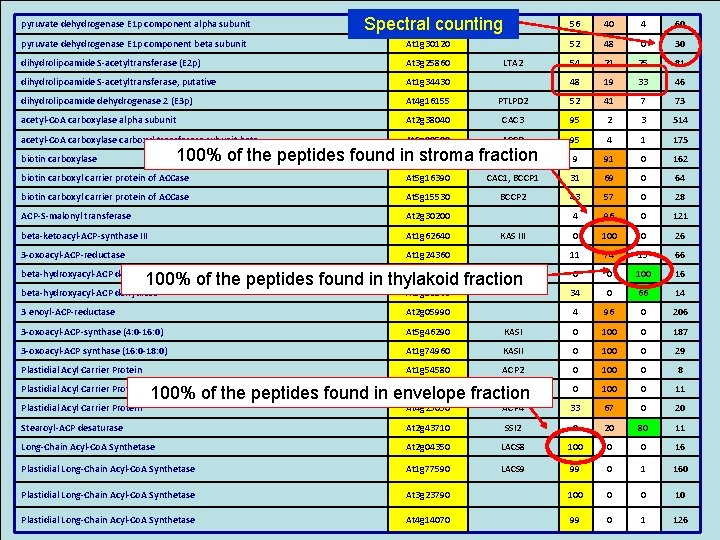
pyruvate dehydrogenase E 1 p component alpha subunit At 1 g 01090 Spectral counting 56 40 4 60 pyruvate dehydrogenase E 1 p component beta subunit At 1 g 30120 52 48 0 30 dihydrolipoamide S-acetyltransferase (E 2 p) At 3 g 25860 LTA 2 54 21 25 81 dihydrolipoamide S-acetyltransferase, putative At 1 g 34430 48 19 33 46 dihydrolipoamide dehydrogenase 2 (E 3 p) At 4 g 16155 PTLPD 2 52 41 7 73 acetyl-Co. A carboxylase alpha subunit At 2 g 38040 CAC 3 95 2 3 514 acetyl-Co. A carboxylase carboxyl transferase subunit beta At. Cg 00500 ACCD 95 4 1 175 9 91 0 162 100% of the peptides found in stroma fraction At 5 g 35360 CAC 2 biotin carboxylase biotin carboxyl carrier protein of ACCase At 5 g 16390 CAC 1, BCCP 1 31 69 0 64 biotin carboxyl carrier protein of ACCase At 5 g 15530 BCCP 2 43 57 0 28 ACP-S-malonyl transferase At 2 g 30200 4 96 0 121 beta-ketoacyl-ACP-synthase III At 1 g 62640 KAS III 0 100 0 26 3 -oxoacyl-ACP-reductase At 1 g 24360 11 74 15 66 0 0 100 16 beta-hydroxyacyl-ACP dehydrase At 2 g 22230 100% of the peptides found in thylakoid fraction beta-hydroxyacyl-ACP dehydrase At 5 g 10160 34 0 66 14 3 enoyl-ACP-reductase At 2 g 05990 4 96 0 206 3 -oxoacyl-ACP-synthase (4: 0 -16: 0) At 5 g 46290 KASI 0 100 0 187 3 -oxoacyl-ACP synthase (16: 0 -18: 0) At 1 g 74960 KASII 0 100 0 29 Plastidial Acyl Carrier Protein At 1 g 54580 ACP 2 0 100 0 8 0 100 0 11 Plastidial Acyl Carrier Protein At 1 g 54630 ACP 3 100% of the peptides found in envelope fraction At 4 g 25050 ACP 4 33 67 0 20 Stearoyl-ACP desaturase At 2 g 43710 SSI 2 0 20 80 11 Long-Chain Acyl-Co. A Synthetase At 2 g 04350 LACS 8 100 0 0 16 Plastidial Long-Chain Acyl-Co. A Synthetase At 1 g 77590 LACS 9 99 0 1 160 Plastidial Long-Chain Acyl-Co. A Synthetase At 3 g 23790 100 0 0 10 Plastidial Long-Chain Acyl-Co. A Synthetase At 4 g 14070 99 0 1 126
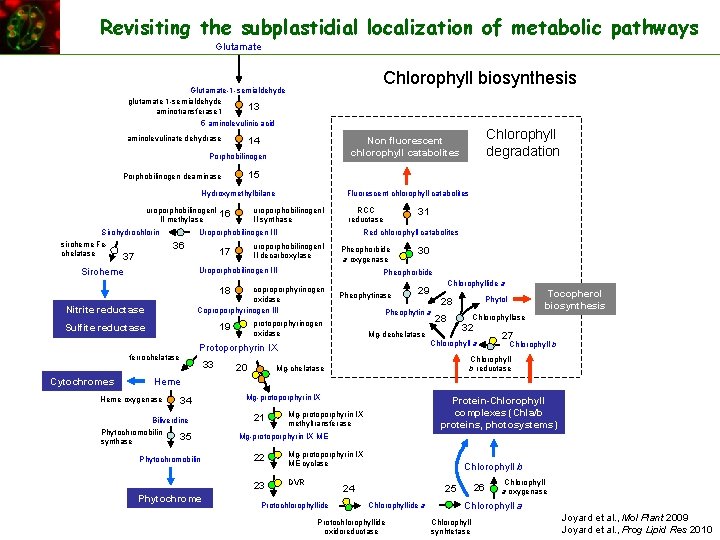
Revisiting the subplastidial localization of metabolic pathways Glutamate Chlorophyll biosynthesis Glutamate-1 -semialdehyde glutamate 1 -semialdehyde 13 aminotransferase 1 5 aminolevulinic acid aminolevulinate dehydrase Porphobilinogen 15 Porphobilinogen deaminase Hydroxymethylbilane uroporphobilinogen. I II methylase Sirohydrochlorin siroheme Fechelatase 37 Fluorescent chlorophyll catabolites uroporphobilinogen. I II synthase 16 RCC reductase Siroheme uroporphobilinogen. I II decarboxylase 17 Pheophorbide a oxygenase Uroporphobilinogen III 30 Pheophorbide Nitrite reductase coproporphyrinogen oxidase Coproporphyrinogen III Sulfite reductase protoporphyrinogen oxidase 18 19 31 Red chlorophyll catabolites Uroporphobilinogen III 36 Pheophytinase Cytochromes 33 20 Chlorophyllide a 29 Pheophytin a 28 Tocopherol biosynthesis Phytol 28 Mg-dechelatase Chlorophyllase 32 Chlorophyll a Protoporphyrin IX ferrochelatase Chlorophyll degradation Non fluorescent chlorophyll catabolites 14 27 Chlorophyll b reductase Mg-chelatase Heme oxygenase 34 Biliverdine Phytochromobilin synthase 35 Phytochromobilin Phytochrome Mg-protoporphyrin IX 21 Protein-Chlorophyll complexes (Chla/b proteins, photosystems) Mg-protoporphyrin IX methyltransferase Mg-protoporphyrin IX ME 22 Mg-protoporphyrin IX ME cyclase 23 DVR Chlorophyll b 24 Protochlorophyllide 26 25 Chlorophyllide a Protochlorophyllide oxidoreductase Chlorophyll a oxygenase Chlorophyll a Chlorophyll synhtetase Joyard et al. , Mol Plant 2009 Joyard et al. , Prog Lipid Res 2010
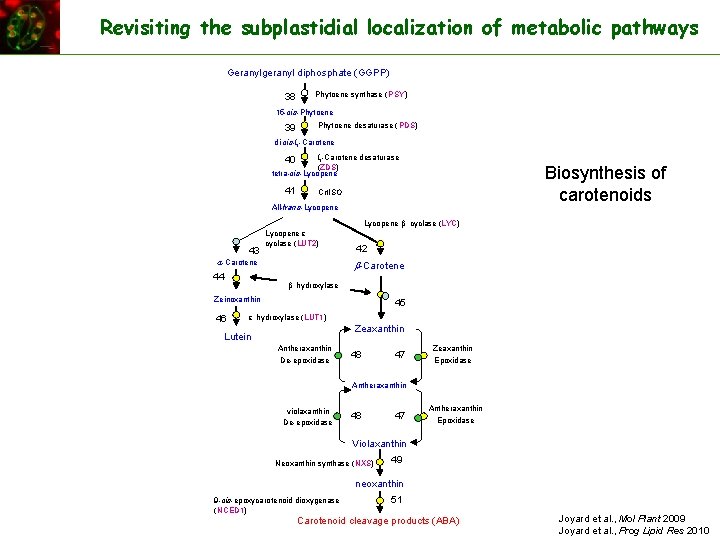
Revisiting the subplastidial localization of metabolic pathways Geranylgeranyl diphosphate (GGPP) 38 Phytoene synthase (PSY) 15 -cis-Phytoene 39 Phytoene desaturase (PDS) di-cis-z-Carotene desaturase (ZDS) tetra-cis-Lycopene 40 41 Biosynthesis of carotenoids Crt. ISO All-trans-Lycopene b cyclase (LYC) 43 Lycopene e cyclase (LUT 2) a-Carotene 44 42 b-Carotene b hydroxylase Zeinoxanthin 46 45 e hydroxylase (LUT 1) Zeaxanthin Lutein Antheraxanthin De-epoxidase 48 47 Zeaxanthin Epoxidase Antheraxanthin violaxanthin De-epoxidase 48 47 Antheraxanthin Epoxidase Violaxanthin Neoxanthin synthase (NXS) 49 neoxanthin 9 -cis-epoxycarotenoid dioxygenase (NCED 1) 51 Carotenoid cleavage products (ABA) Joyard et al. , Mol Plant 2009 Joyard et al. , Prog Lipid Res 2010
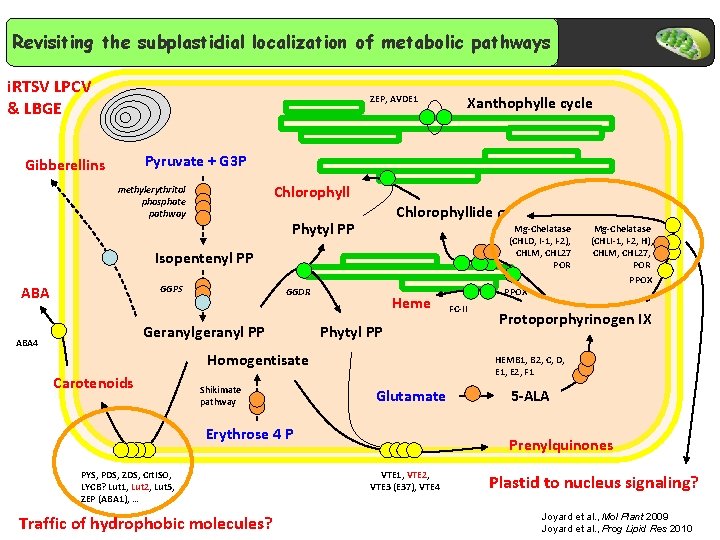
Revisiting the subplastidial localization of metabolic pathways i. RTSV LPCV & LBGE ZEP, AVDE 1 Xanthophylle cycle Pyruvate + G 3 P Gibberellins Chlorophyll methylerythritol phosphate pathway Chlorophyllide a Phytyl PP Mg-Chelatase (CHLD, I-1, I-2), CHLM, CHL 27 POR Isopentenyl PP ABA GGPS GGDR Geranylgeranyl PP ABA 4 Heme Phytyl PP Homogentisate Carotenoids Shikimate pathway Traffic of hydrophobic molecules? PPOX FC-II Protoporphyrinogen IX HEMB 1, B 2, C, D, E 1, E 2, F 1 Glutamate Erythrose 4 P PYS, PDS, ZDS, Crt. ISO, LYCB? Lut 1, Lut 2, Lut 5, ZEP (ABA 1), … Mg-Chelatase (CHLI-1, I-2, H), CHLM, CHL 27, POR 5 -ALA Prenylquinones VTE 1, VTE 2, VTE 3 (E 37), VTE 4 Plastid to nucleus signaling? Joyard et al. , Mol Plant 2009 Joyard et al. , Prog Lipid Res 2010

Application of chloroplast proteomics 2 -Characterization of previously unidentified chloroplast envelope transporters
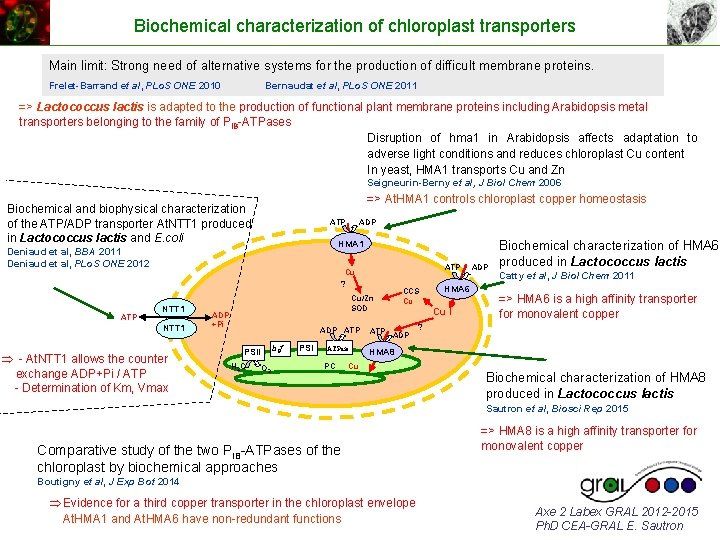
Biochemical characterization of chloroplast transporters Main limit: Strong need of alternative systems for the production of difficult membrane proteins. Frelet-Barrand et al, PLo. S ONE 2010 Bernaudat et al, PLo. S ONE 2011 => Lactococcus lactis is adapted to the production of functional plant membrane proteins including Arabidopsis metal transporters belonging to the family of PIB-ATPases Disruption of hma 1 in Arabidopsis affects adaptation to adverse light conditions and reduces chloroplast Cu content In yeast, HMA 1 transports Cu and Zn Seigneurin-Berny et al, J Biol Chem 2006 => At. HMA 1 controls chloroplast copper homeostasis Biochemical and biophysical characterization of the ATP/ADP transporter At. NTT 1 produced in Lactococcus lactis and E. coli ATP ADP HMA 1 Deniaud et al, BBA 2011 Deniaud et al, PLo. S ONE 2012 ATP Cu ? ATP NTT 1 - At. NTT 1 allows the counter exchange ADP+Pi / ATP - Determination of Km, Vmax CCS Cu Cu/Zn SOD ADP +Pi ADP ATP b 6 f PSII H 2 O O 2 PSI ATPase PC ATP HMA 6 Cu I ADP Biochemical characterization of HMA 6 produced in Lactococcus lactis Catty et al, J Biol Chem 2011 => HMA 6 is a high affinity transporter for monovalent copper ? HMA 8 Cu Biochemical characterization of HMA 8 produced in Lactococcus lactis Sautron et al, Biosci Rep 2015 Comparative study of the two PIB-ATPases of the chloroplast by biochemical approaches => HMA 8 is a high affinity transporter for monovalent copper Boutigny et al, J Exp Bot 2014 Evidence for a third copper transporter in the chloroplast envelope At. HMA 1 and At. HMA 6 have non-redundant functions Axe 2 Labex GRAL 2012 -2015 Ph. D CEA-GRAL E. Sautron
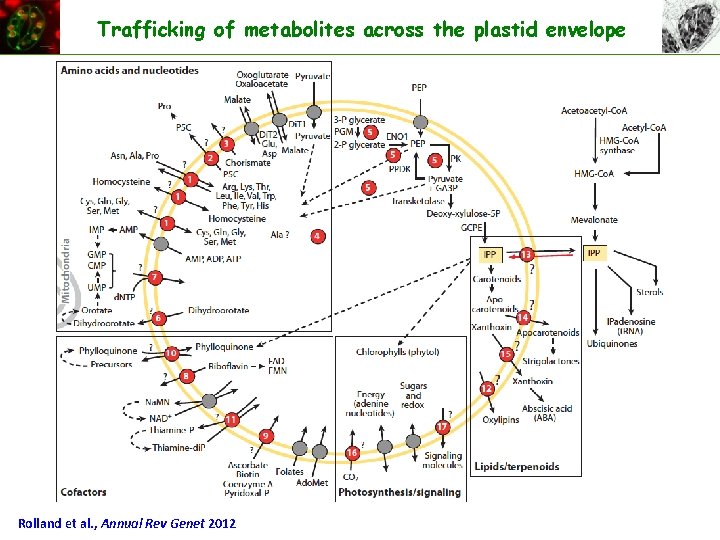
Trafficking of metabolites across the plastid envelope Rolland et al. , Annual Rev Genet 2012
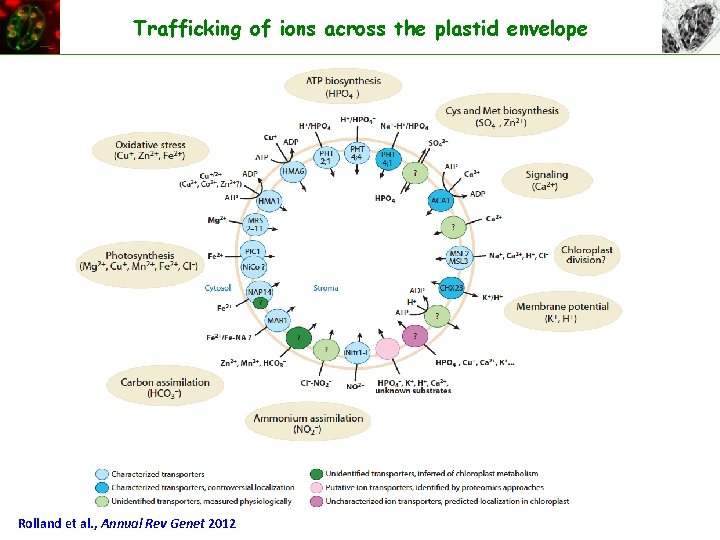
Trafficking of ions across the plastid envelope Rolland et al. , Annual Rev Genet 2012

Application of chloroplast proteomics 3 -Identification of previously unidentified chloroplast targeting pathways
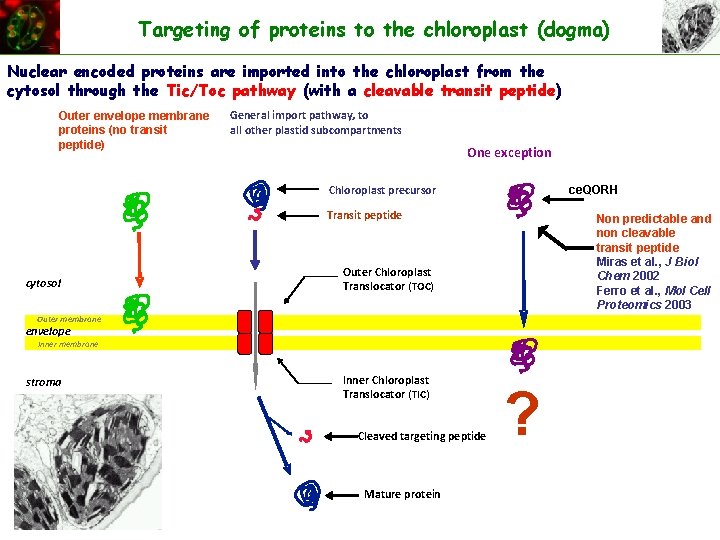
Targeting of proteins to the chloroplast (dogma) Nuclear encoded proteins are imported into the chloroplast from the cytosol through the Tic/Toc pathway (with a cleavable transit peptide) Outer envelope membrane proteins (no transit peptide) General import pathway, to all other plastid subcompartments One exception Chloroplast precursor ce. QORH Transit peptide Non predictable and non cleavable transit peptide Miras et al. , J Biol Chem 2002 Ferro et al. , Mol Cell Proteomics 2003 cytosol Outer Chloroplast Translocator (TOC) Outer membrane envelope Inner membrane stroma Inner Chloroplast Translocator (TIC) Cleaved targeting peptide Mature protein ?
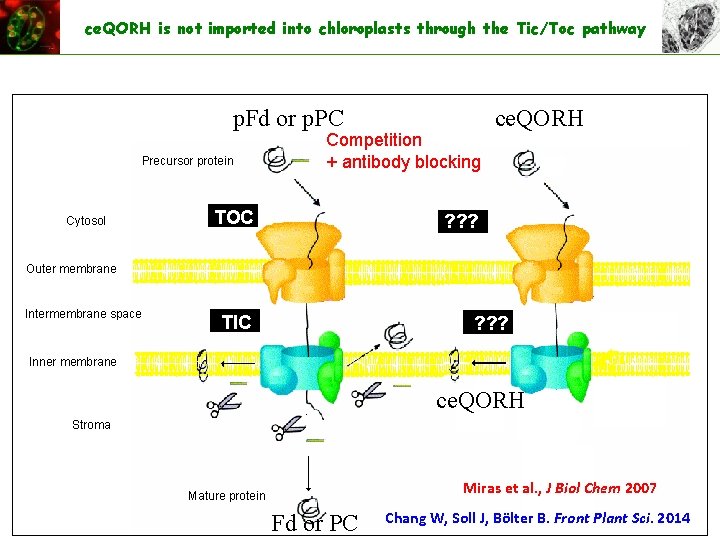
ce. QORH is not imported into chloroplasts through the Tic/Toc pathway p. Fd or p. PC Precursor protein Cytosol Competition + antibody blocking TOC ce. QORH ? ? ? Outer membrane Intermembrane space TIC ? ? ? Inner membrane ce. QORH Stroma Miras et al. , J Biol Chem 2007 Mature protein Fd or PC Chang W, Soll J, Bölter B. Front Plant Sci. 2014
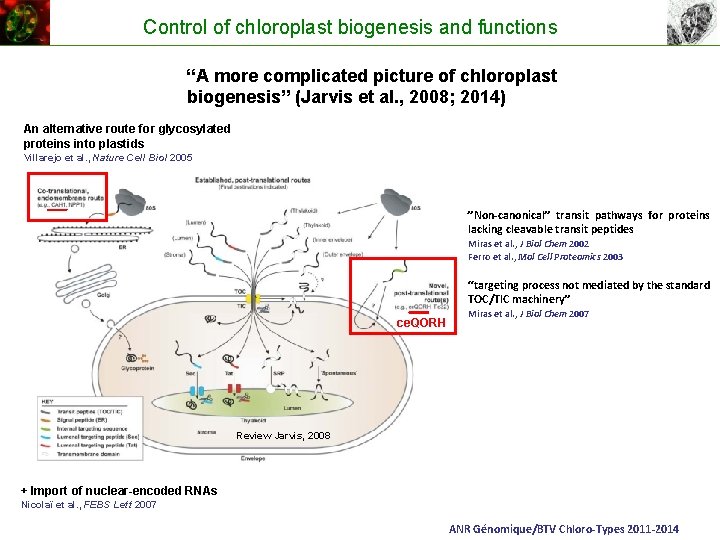
Control of chloroplast biogenesis and functions “A more complicated picture of chloroplast biogenesis” (Jarvis et al. , 2008; 2014) An alternative route for glycosylated proteins into plastids Villarejo et al. , Nature Cell Biol 2005 ”Non-canonical” transit pathways for proteins lacking cleavable transit peptides Miras et al. , J Biol Chem 2002 Ferro et al. , Mol Cell Proteomics 2003 “targeting process not mediated by the standard TOC/TIC machinery” ce. QORH Miras et al. , J Biol Chem 2007 Review Jarvis, 2008 + Import of nuclear-encoded RNAs Nicolaï et al. , FEBS Lett 2007 ANR Génomique/BTV Chloro-Types 2011 -2014

Application of chloroplast proteomics 4 -Dynamics of thylakoid membranes
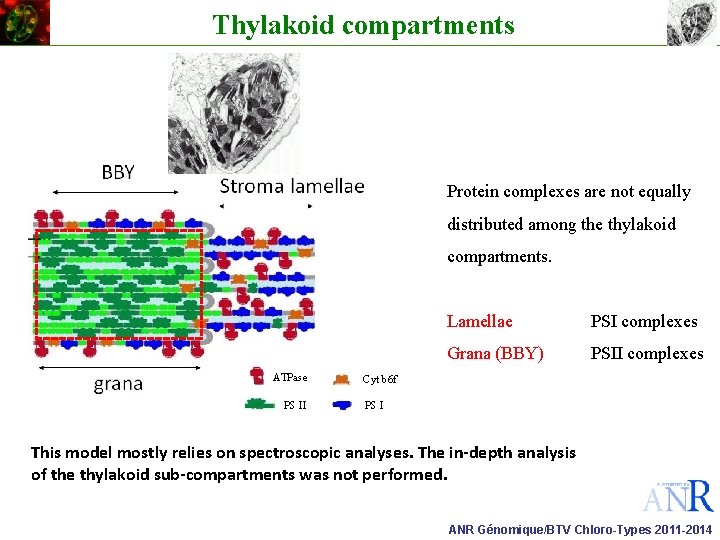
Thylakoid compartments Protein complexes are not equally distributed among the thylakoid compartments. ATPase PS II Lamellae PSI complexes Grana (BBY) PSII complexes Cyt b 6 f PS I This model mostly relies on spectroscopic analyses. The in-depth analysis of the thylakoid sub-compartments was not performed. ANR Génomique/BTV Chloro-Types 2011 -2014
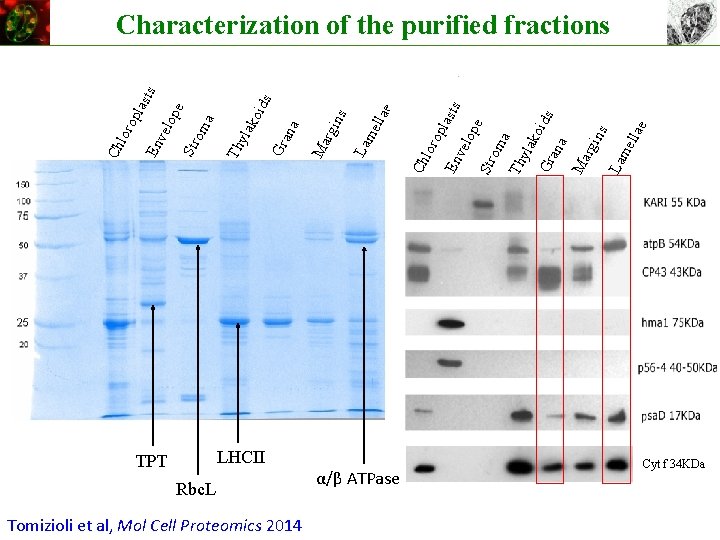
LHCII TPT Rbc. L Tomizioli et al, Mol Cell Proteomics 2014 α/β ATPase llae La me ns rgi Ma ana ko Gr yla a Th Str om op e ids sts En vel op la lor Ch ella La m ns rgi Ma Gr ana e ids ko yla Th ma pe Str o elo En v Ch lor op la sts Characterization of the purified fractions Cyt f 34 KDa

Revisiting the localization of photosynthetic complexes HIGH Grana Tomizioli et al, Mol Cell Proteomics 2014 HIGH LAM Perspectives: dynamics of the system
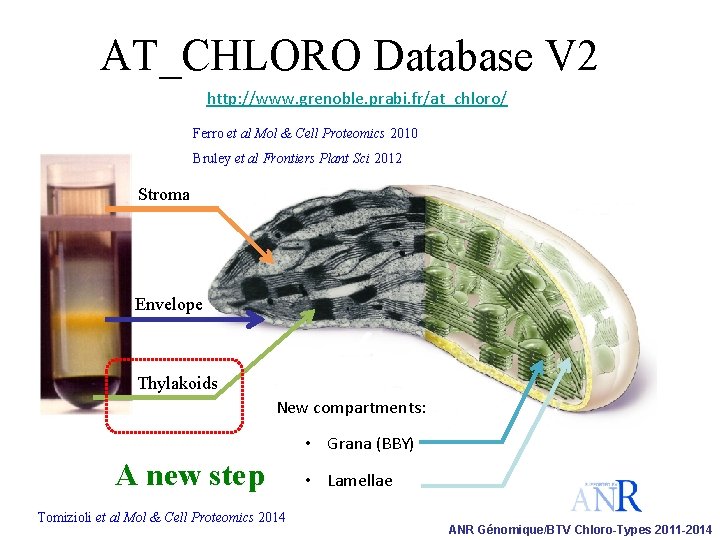
AT_CHLORO Database V 2 http: //www. grenoble. prabi. fr/at_chloro/ Ferro et al Mol & Cell Proteomics 2010 Bruley et al Frontiers Plant Sci 2012 Stroma Envelope Thylakoids New compartments: • Grana (BBY) A new step Tomizioli et al Mol & Cell Proteomics 2014 • Lamellae ANR Génomique/BTV Chloro-Types 2011 -2014
Subcellular and subplastidial proteomics ? Recent data strongly suggest that the dynamics of the plastid proteome may not only result from variations in protein abundance or post-translational modifications but might also result from dynamic changes in the subcellular or even subplastidial localization of specific plastid proteins. => Hypothesis The dynamics of subcellular (and sub-organellar) proteomes is essential for optimal adjustment of the chloroplast functioning and thus of plant adaptation to environmental constraints. => Perspectives 1) Deciphering molecular mechanisms controlling the targeting of plastid proteins shared by several cell compartments and imported into plastids through alternative pathways, 2) Exploring the dynamics of thylakoid sub-compartments, as a major process regulating photosynthetic electron transfer.

Contributors Laboratoire de Physiologie Cellulaire & Végétale CNRS/Univ. Grenoble Alpes/CEA Grenoble/INRA Sylvain Boutigny Laboratoire de Biologie à Grande Echelle Cécile Giustini CEA Grenoble/Univ. Grenoble Alpes/INSERM Gilles Curien LPCV, Team 1 Giovanni Finazzi Annie Frelet Chloroplast Jacques Joyard experts Marcel Kuntz Sophie Le Gall Michel Matringe Stéphane Miras Lucas Moyet Norbert Rolland Daniel Salvi Daphné Seigneurin-Berny Emeline Sautron Martino Tomizioli Funding: ANR, CNRS, CEA, INSERM, INRA Claire Adam Sabine Brugière Thomas Burger BGE, Team EDy. P Christophe Bruley Florence Combes MS, statistics and Magali Court bioinformatics Yohann Couté Véronique Dupierris Myriam Ferro Jérôme Garin Anne-Marie Hesse Cosmin Lazar Sylvie Kieffer-Jaquinod Christophe Masselon Mourad Mellal Claire Ramus Yves Vandenbrouck
- Slides: 29