Plant molecular genetics Plant nuclear genome Chromatin and




















- Slides: 51

Plant molecular genetics • • • Plant nuclear genome Chromatin and DNA methylation RNA interference Transposible elements, Viruses Genomes of plastids and mitochondria • • • Classical genetic mapping Transgenosis and reverse genetics Genomics, transcriptomics and proteomics

Components of plant genome • nuclear genome = genome sensu stricto • plastids - plastome • mitochondria - chondriome

Plant genome sizes 63 Mbp – Genlisea aurea 125 000 Mbp - Fritillaria 149 000 Mbp - Paris japonica - currently the largest genome of multicellular organisms (not only plant) http: //data. kew. org/cvalues/

Genome size = C-value • C-value = size of genome in non-replicated gamete genome size (bp) = (0. 910 x 109) x DNA content (pg) = genome size (bp) / (0. 910 x 109) 1 pg = cca 910 Mbp; MW (1 bp) = cca 660 Da What is the amount of nuclear DNA in somatic cells? - depends on cell type a cell-cycle phase - usually 2 x (G 1) and 4 x (G 2), in endosperm 3 x/6 x C-value

Plant genome sizes 10 Mbp Ostreococcus (single cell alga) 63 Mbp Genlisea aurea 135 Mbp Arabidopsis thaliana Ratio of globe volumes differing 3000 times 500 Mbp Oryza 2 500 Mbp Zea mays 5 000 Mbp Hordeum Paris japonica 17 000 Mbp Triticum 149 000 000 bp 84 000 Mbp Fritillaria (largest diploid) 149 000 Mbp Paris (oktaploid) - Angiosperms – size differences up to almost 3 000 times - Gymnosperms – genome sizes often around 10 000 Mb - Gene number differs not so much (app. 20 – 200(300) thousand)

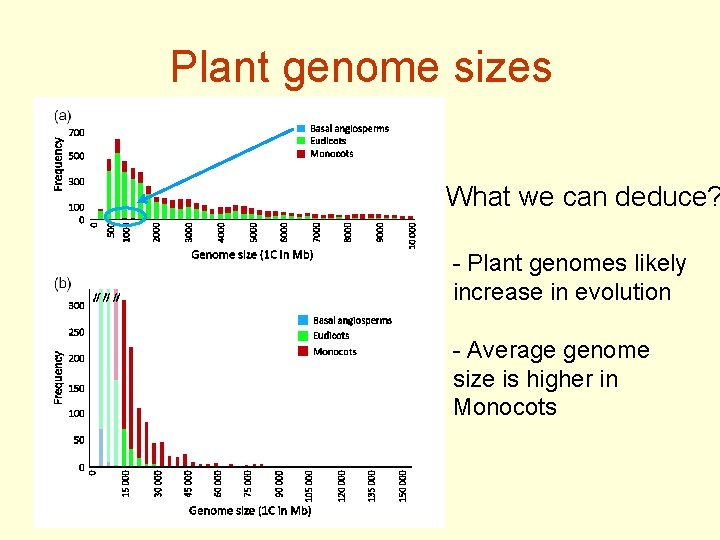
Plant genome sizes What we can deduce? - Plant genomes likely increase in evolution - Average genome size is higher in Monocots

C-value paradox - there is only weak correlation between plant complexity and the size of its genome Sources of genome size differences: - whole genomes duplications (polyploidization) - multiplication of invasive DNA (transposable elements) genomes of related organisms often strongly differ in size Rice: 500 Mbp Wheat: 17 000 Mbp

Transposible elements (TE) genomic DNA sequences that can move around to different (another) positions within the genome of a single cell • „CUT and PASTE“ cleaving out, transfer and insertion to a new site • „COPY and PASTE“ copying (repeated), insertion of a new copies into new sites (original TE remains anaffected)

C-value paradox - there is only a weak correlation between plant complexity and the size of its genome Sources of genome size differences: - whole genomes duplications (polyploidization) - replication of invasive DNA (transposable elements) It is not a one-way road to genome obesity! Genome size can be reduced via deletions mediated by homologous recombination.

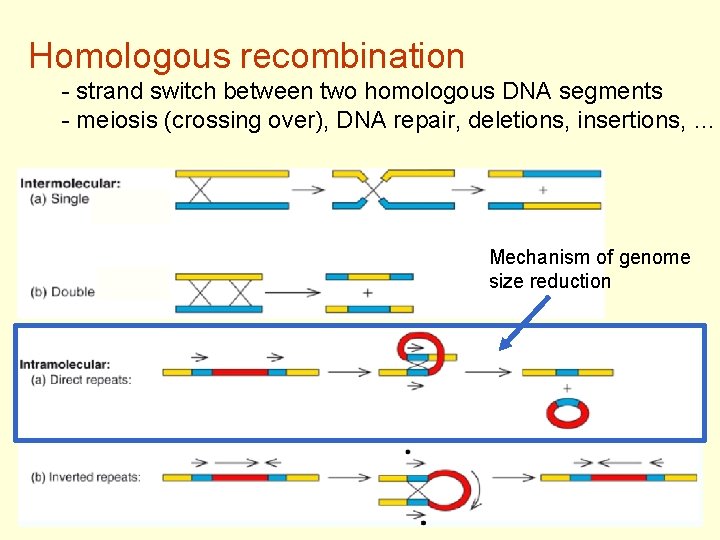
Homologous recombination - strand switch between two homologous DNA segments - meiosis (crossing over), DNA repair, deletions, insertions, … Mechanism of genome size reduction

Sequences in plant genomes Unique sequences – genes, but also non-coding regulatory sequences (!) Repetitive: • Medium repetitive DNA – Tandem repeats of r. DNA, t. RNA genes, histon genes – Gene families with multiple members (highly similar) – Transposable elements – can be also highly repetitive • Highly repetitive – Tandem arranged simple sequence repeats (SSR) – Centromers (180 bp repeat in Arabidopsis) a telomers (TTTAGGG)n – often low complexity DNA

Sequence complexity (~ the amount of information) Repetitive = low complexity AAAAAAAAAAA complexity 1 (21 x. A) ATCATCATCATC complexity 3 (7 x. ATC) (what is the complexity if it is a coding sequence? ) unique ATCGTATCGCGATTTTAACGT complexity 21 (1 x. AT…) - unique x repetitive – depends on the size of the evaluated frame (= size of analyzed DNA fragments in reassociation kinetics)

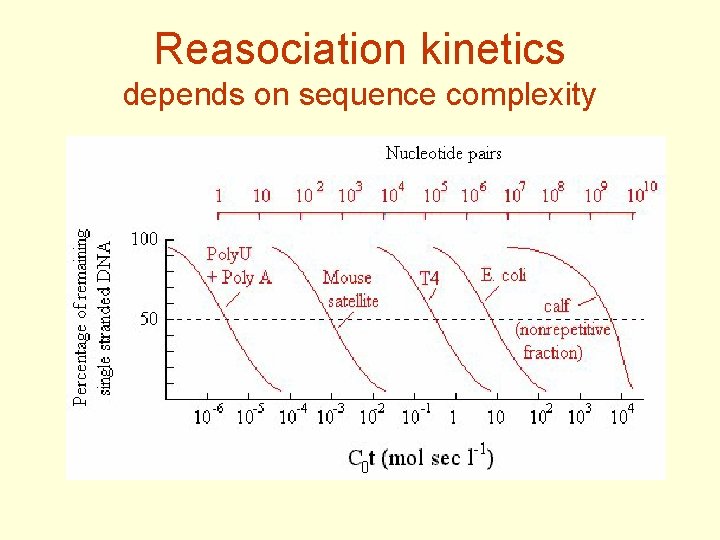
Measuring of genome complexity - reasociation kinetics • DNA fragmented to 300 - 500 bp, denatured • Monitoring of reassociation in time - separation (e. g. chromatographic) of ss and ds DNA • Analysis of kinetics (Cot curves) shows representation of various types of repetitive DNA – rare sequences reasociate more slowly that repetitive

Reasociation kinetics depends on sequence complexity

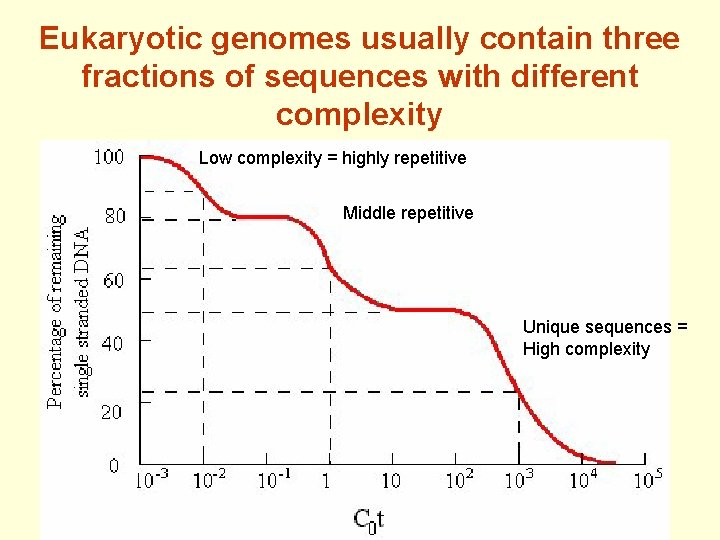
Eukaryotic genomes usually contain three fractions of sequences with different complexity Low complexity = highly repetitive Middle repetitive Unique sequences = High complexity

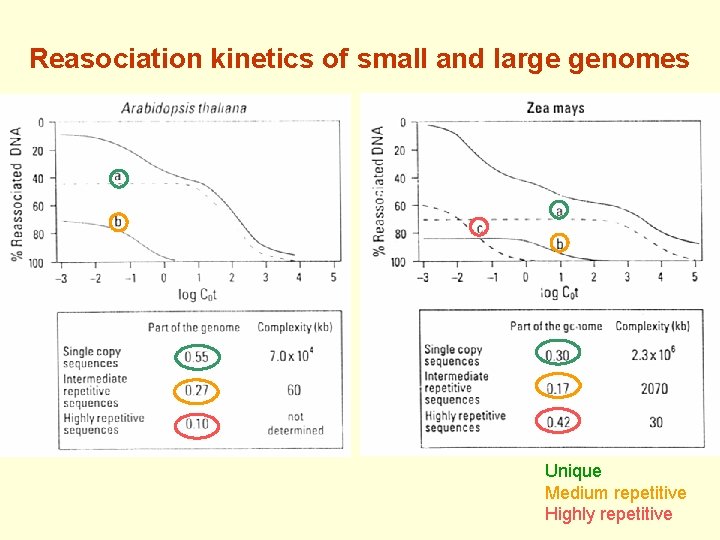
Reasociation kinetics of small and large genomes Unique Medium repetitive Highly repetitive

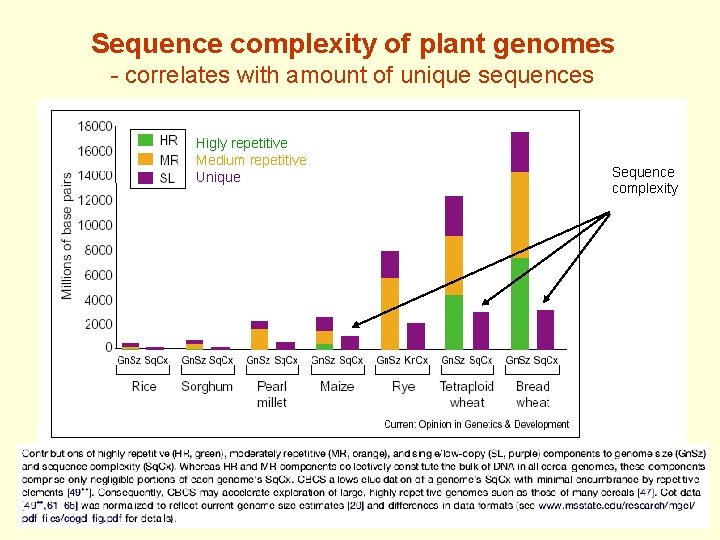
Sequence complexity of plant genomes - correlates with amount of unique sequences Higly repetitive Medium repetitive Unique Sequence complexity

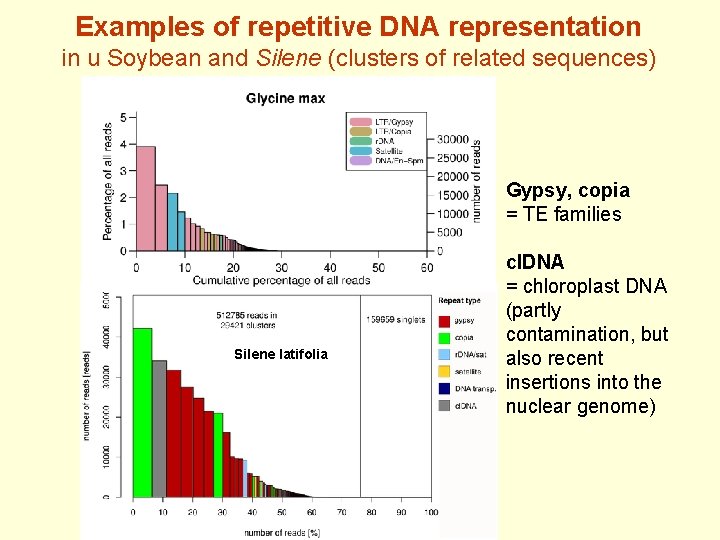
Examples of repetitive DNA representation in u Soybean and Silene (clusters of related sequences) Gypsy, copia = TE families Silene latifolia cl. DNA = chloroplast DNA (partly contamination, but also recent insertions into the nuclear genome)

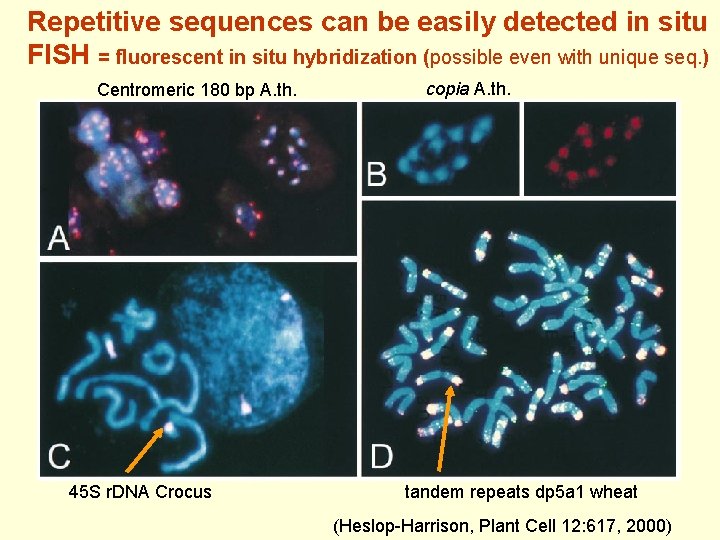
Repetitive sequences can be easily detected in situ FISH = fluorescent in situ hybridization (possible even with unique seq. ) Centromeric 180 bp A. th. 45 S r. DNA Crocus copia A. th. tandem repeats dp 5 a 1 wheat (Heslop-Harrison, Plant Cell 12: 617, 2000)

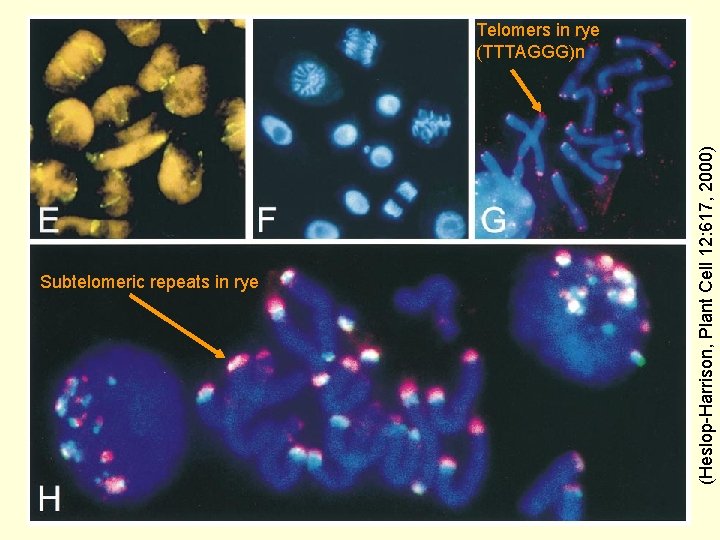
Subtelomeric repeats in rye (Heslop-Harrison, Plant Cell 12: 617, 2000) Telomers in rye (TTTAGGG)n

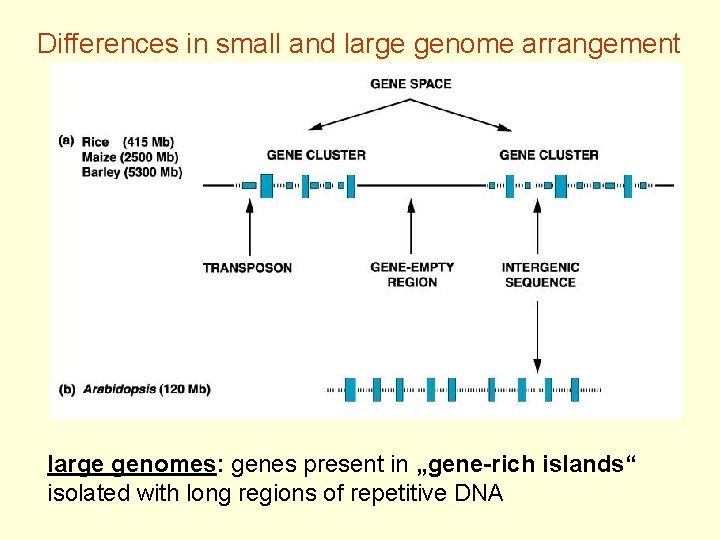
Differences in small and large genome arrangement large genomes: genes present in „gene-rich islands“ isolated with long regions of repetitive DNA

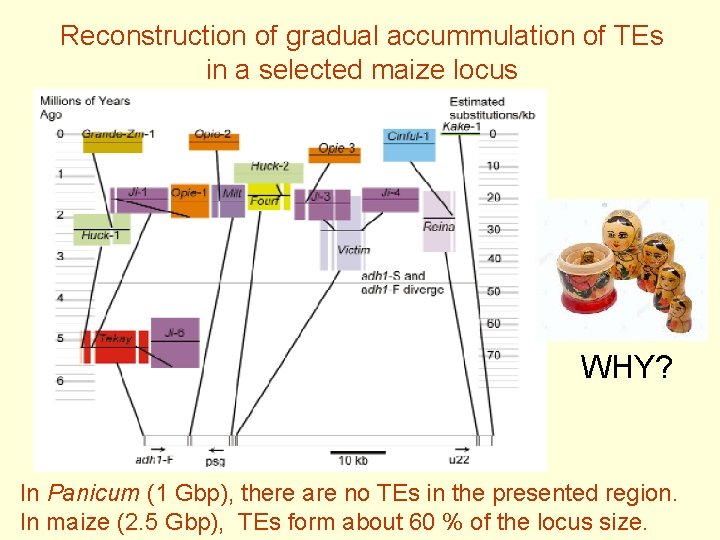
Reconstruction of gradual accummulation of TEs in a selected maize locus WHY? In Panicum (1 Gbp), there are no TEs in the presented region. In maize (2. 5 Gbp), TEs form about 60 % of the locus size.

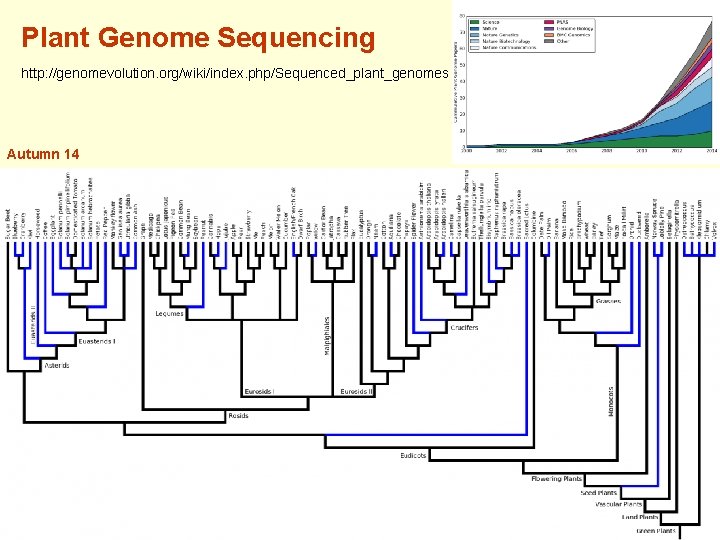
Plant Genome Sequencing http: //genomevolution. org/wiki/index. php/Sequenced_plant_genomes Autumn 14

Arabidopsis thaliana the most important model of plant biology 1 week 4 weeks 3 weeks 6 weeks

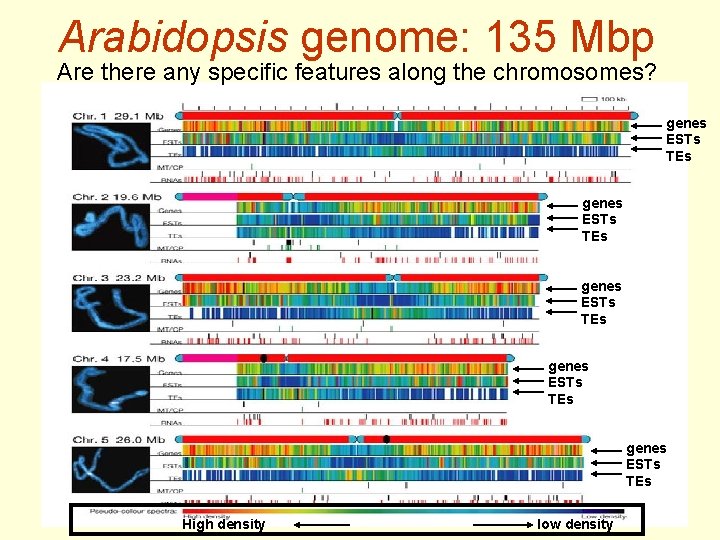
Arabidopsis genome: 135 Mbp Are there any specific features along the chromosomes? genes ESTs TEs genes ESTs TEs High density low density

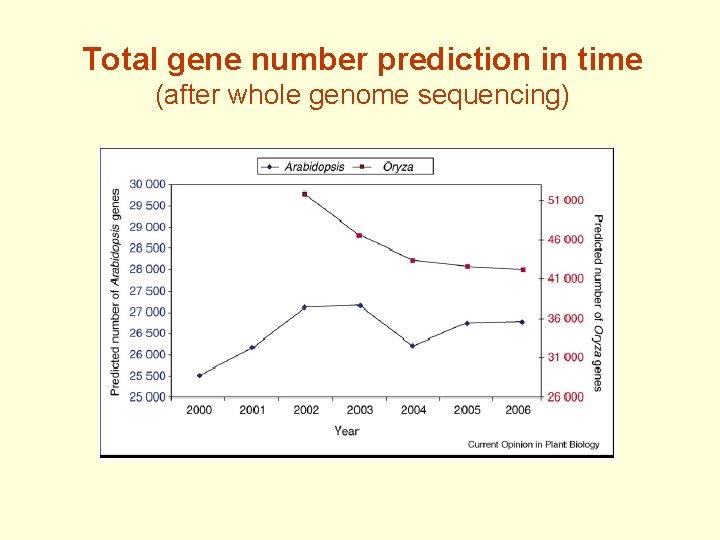
Total gene number prediction in time (after whole genome sequencing)

Genome of Arabidopsis - statistics Value Feature DNA molecule Chr. 1 Length (bp) Top arm (bp) Bottom arm (bp) 29, 105, 111 14, 449, 213 14, 655, 898 Base composition (%GC) Overall Coding Non-coding Number of genes Gene density (kb per gene ) Average gene Length (bp) Average peptide Length (bp) Exons Number Total length (bp) Average per gene Average size (bp) Number of genes With ESTs (%) Number of ESTs Chr. 2 Chr. 3 Chr. 4 Chr. 5 SUM 19, 646, 945 3, 607, 091 16, 039, 854 23, 172, 617 13, 590, 268 9, 582, 349 17, 549, 867 3, 052, 108 14, 497, 759 25, 53, 409 11, 132, 192 14, 803, 217 115, 409, 949 33. 4 44. 0 32. 4 35. 5 44. 0 32. 9 35. 4 44. 3 33. 0 35. 5 44. 1 32. 8 34. 5 44. 1 32. 5 6, 543 4. 0 4, 036 4. 9 5, 220 4. 5 3, 825 4. 6 5, 874 4. 4 approx. 27, 000 2, 078 1, 949 1, 925 2, 138 1, 974 protein coding genes 446 421 424 448 429 35, 482 8, 772, 559 5. 4 247 19, 631 5, 100, 288 4. 9 259 26, 570 6, 654, 507 5. 1 250 20, 073 5, 150, 883 5. 2 256 31, 226 7, 571, 013 5. 3 242 60. 8 56. 9 59. 8 61. 4 30, 522 14, 989 20, 732 16, 605 22, 885 132, 982 33, 249, 250 105, 773 + hundreds of MIR genes - role in regulation of gene expression

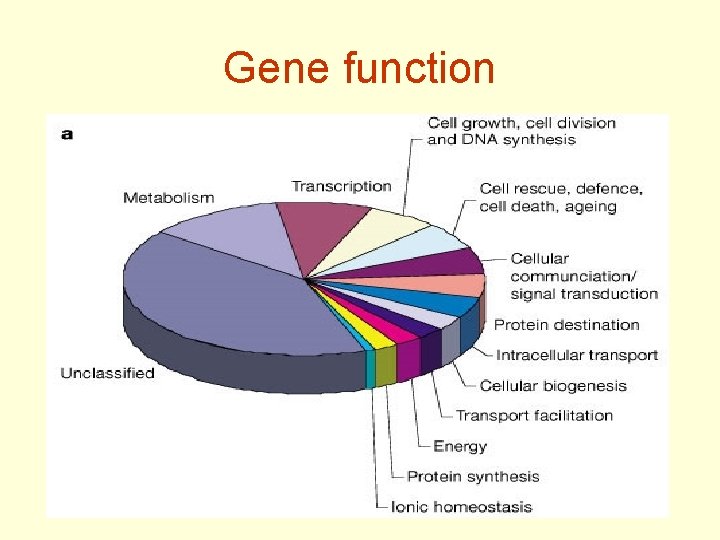
Gene function

The majority of plant genes form gene families Number of paraloques • duplications of long chromosomal regions (remnants of ancient polyploidy) • gene families are often in tandem arrangement, but also spread in the genome • tandem repeats are composed of near, but also far paralogues (recombinations)

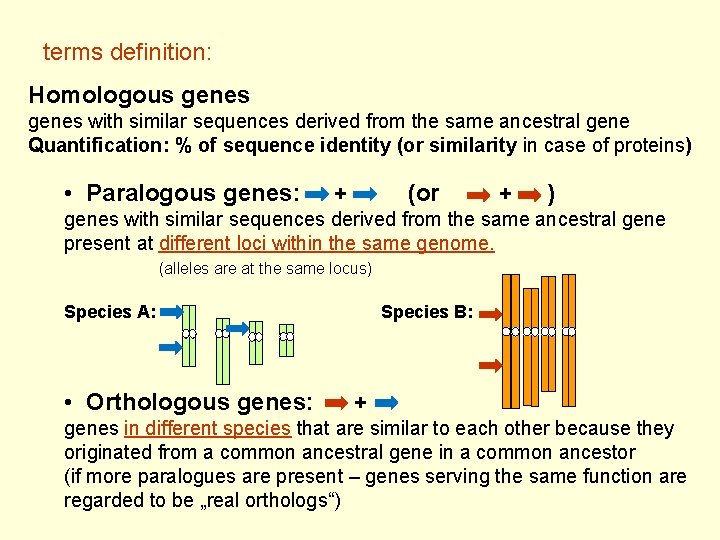
terms definition: Homologous genes with similar sequences derived from the same ancestral gene Quantification: % of sequence identity (or similarity in case of proteins) • Paralogous genes: + (or + ) genes with similar sequences derived from the same ancestral gene present at different loci within the same genome. (alleles are at the same locus) Species A: • Orthologous genes: Species B: + genes in different species that are similar to each other because they originated from a common ancestral gene in a common ancestor (if more paralogues are present – genes serving the same function are regarded to be „real orthologs“)

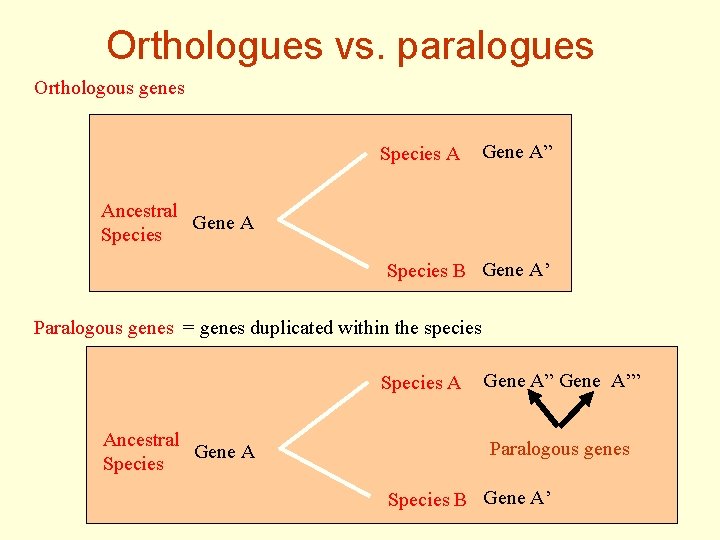
Orthologues vs. paralogues Orthologous genes Species A Gene A” Ancestral Gene A Species B Gene A’ Paralogous genes = genes duplicated within the species Species A Ancestral Gene A Species Gene A” Gene A’” Paralogous genes Species B Gene A’

Mechanisms of gene duplications = increase in paralogue number (within a genome) • tandem duplication (recombination) • transpozition • segmental duplications • whole genome duplications

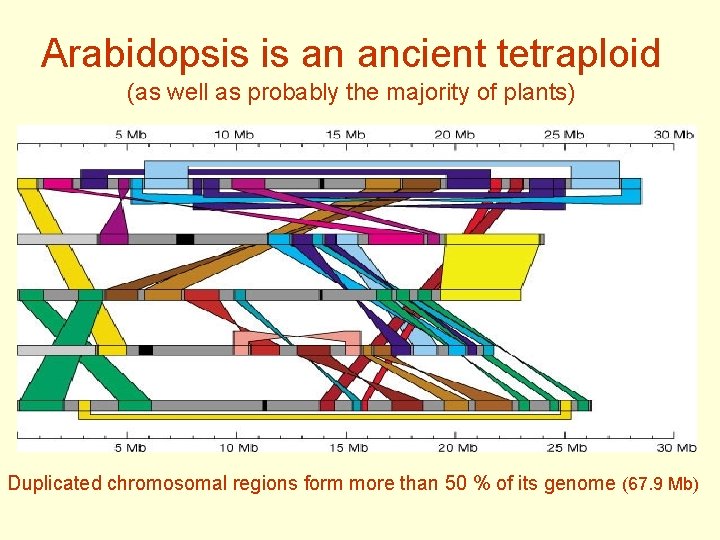
Arabidopsis is an ancient tetraploid (as well as probably the majority of plants) Duplicated chromosomal regions form more than 50 % of its genome (67. 9 Mb)

Polyploidy Who? autopolyploidy x allopolyploidy (the „same“ or different genomes) When? paleopolyploidy x neopolyploidy (paleo- followed by rearrangements/reductions resulting in diploidization) • Polyploidy significantly increases genome (and organism) plasticity • Polyploidy played very important role in plant (genome) evolution; Examples of neofunctionalization allowed by polypoidy: formation of nitrogen fixing nodules (Fabaceae) formation of juicy fruits (Solanaceae)

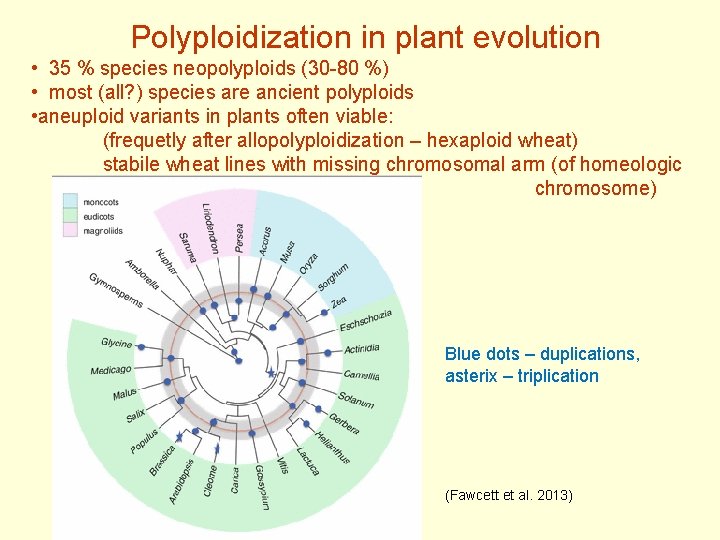
Polyploidization in plant evolution • 35 % species neopolyploids (30 -80 %) • most (all? ) species are ancient polyploids • aneuploid variants in plants often viable: (frequetly after allopolyploidization – hexaploid wheat) stabile wheat lines with missing chromosomal arm (of homeologic chromosome) Blue dots – duplications, asterix – triplication (Fawcett et al. 2013)

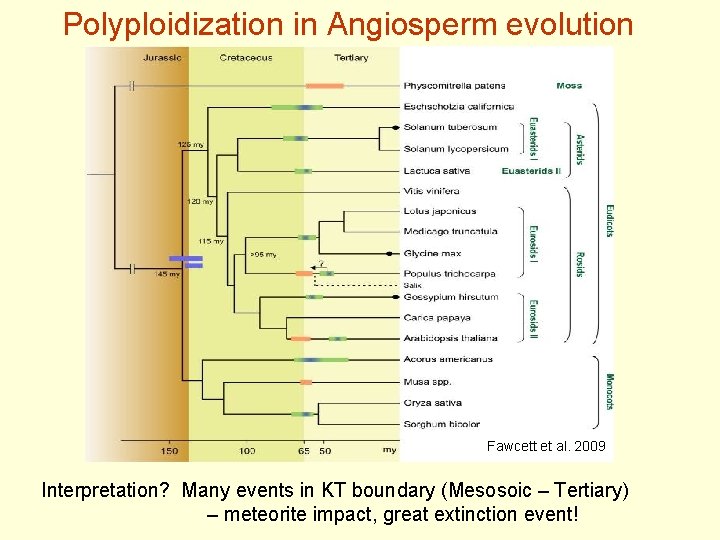
Polyploidization in Angiosperm evolution Fawcett et al. 2009 Interpretation? Many events in KT boundary (Mesosoic – Tertiary) – meteorite impact, great extinction event!

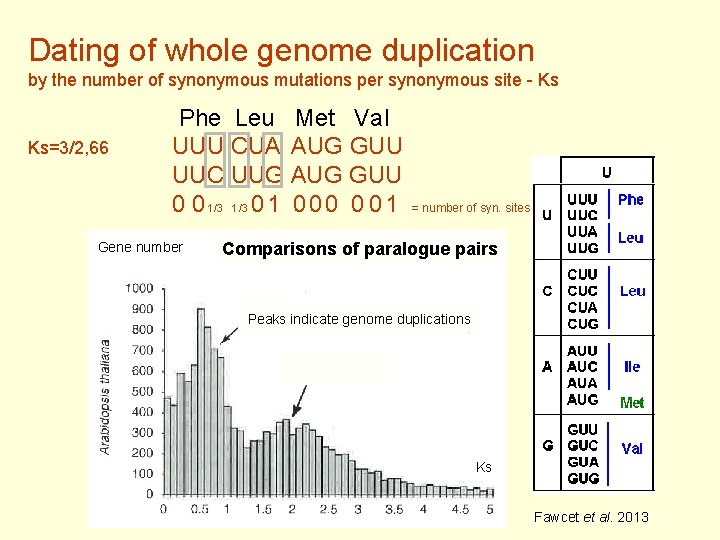
Dating of whole genome duplication by the number of synonymous mutations per synonymous site - Ks Ks=3/2, 66 Phe Leu Met Val UUU CUA AUG GUU UUC UUG AUG GUU 0 0 1/3 0 1 0 0 0 1 = number of syn. sites Gene number Comparisons of paralogue pairs Peaks indicate genome duplications Ks Fawcet et al. 2013

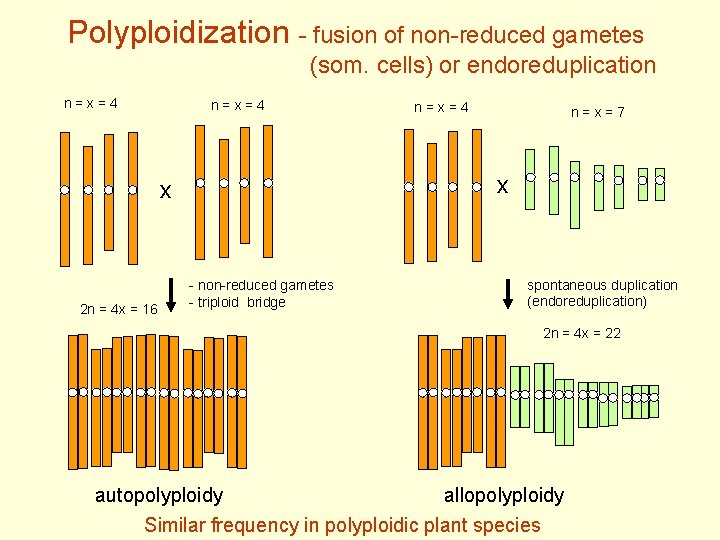
Polyploidization - fusion of non-reduced gametes (som. cells) or endoreduplication n = x = 4 n = x = 7 x x 2 n = 4 x = 16 n = x = 4 - non-reduced gametes - triploid bridge spontaneous duplication (endoreduplication) 2 n = 4 x = 22 autopolyploidy allopolyploidy Similar frequency in polyploidic plant species

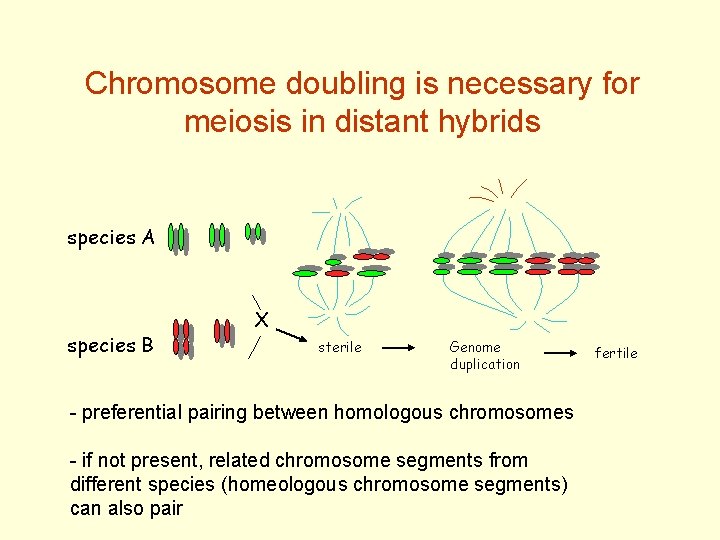
Chromosome doubling is necessary for meiosis in distant hybrids species A species B X sterile Genome duplication - preferential pairing between homologous chromosomes - if not present, related chromosome segments from different species (homeologous chromosome segments) can also pair fertile

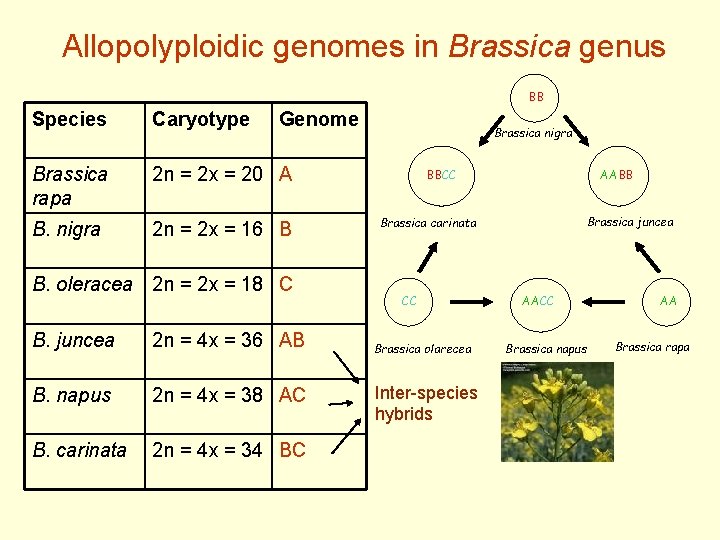
Allopolyploidic genomes in Brassica genus BB Species Caryotype Genome Brassica rapa 2 n = 2 x = 20 A B. nigra 2 n = 2 x = 16 B B. oleracea 2 n = 2 x = 18 C B. juncea 2 n = 4 x = 36 AB B. napus 2 n = 4 x = 38 AC B. carinata 2 n = 4 x = 34 BC Brassica nigra BBCC AABB Brassica juncea Brassica carinata CC Brassica olarecea Inter-species hybrids AACC Brassica napus AA Brassica rapa

Fade of duplicated genes differ: “connected genes“ encoding interacting proteins (components of signal pathways, complex subunits, …) - easily preserve in the genome after duplication - loss or partial duplication of one component results in gene inbalance decreasing fitness (gene dosage theory) - duplication of genes for the whole complex allows its specialization for a new function = increased complexity “single genes“ - can be more easily lost after genome duplication, but can be preserved after individual duplication

Fade of duplicated genes: neofunctionalization x subfunctionalization Escape from adaptive conflict (EAC model) - secondary functions likely present already in the ancester = adaptive conflict (cannot serve two functions perfectly) - duplication allows adaptive evolution of both functions without selection constrains = escape

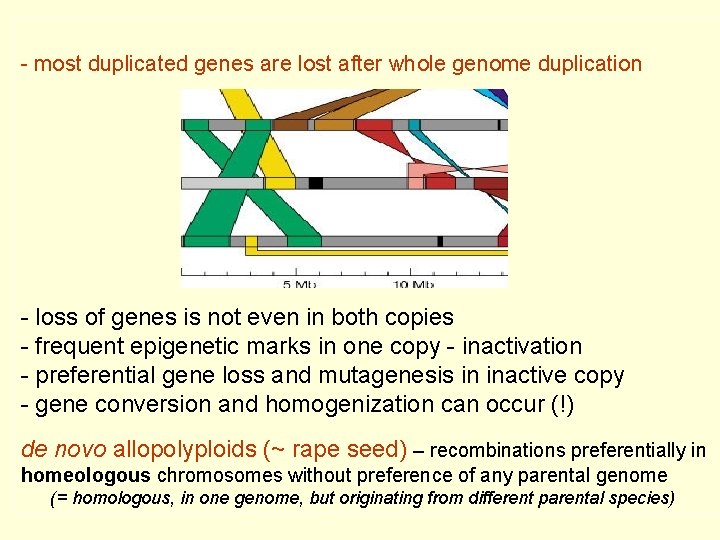
- most duplicated genes are lost after whole genome duplication - loss of genes is not even in both copies - frequent epigenetic marks in one copy - inactivation - preferential gene loss and mutagenesis in inactive copy - gene conversion and homogenization can occur (!) de novo allopolyploids (~ rape seed) – recombinations preferentially in homeologous chromosomes without preference of any parental genome (= homologous, in one genome, but originating from different parental species)

Changes in newly formed allopolyploid genome: - losses of parts or whole chromosomes (aneuploidy = decreased fertility) - frequent activation of TE - expression of homeologous genes is not usually additive - transcriptome usually more reduced than genome - different regulation of gene expression - often different organ specific expression of genes from each parent, - new sites of expression, new regulation - „divergent resolution“ – potential mechanism of speciation (different gene loss in individuals - lethality in F 2, - absence of essential gene = reproduction barrier

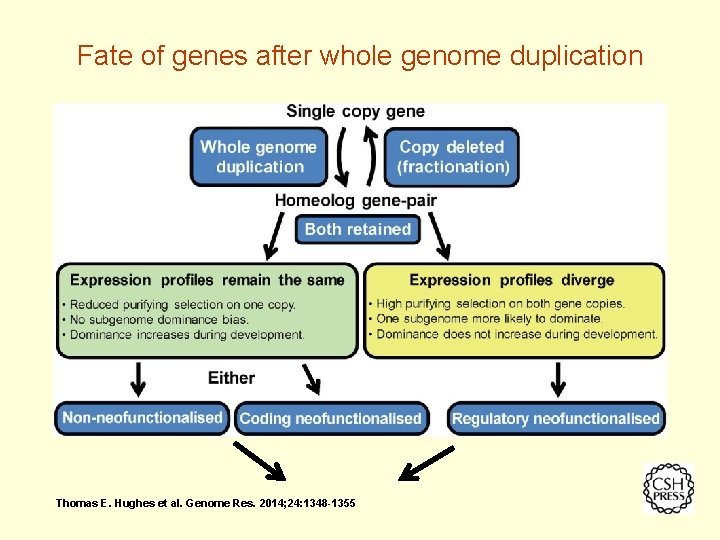
Fate of genes after whole genome duplication Thomas E. Hughes et al. Genome Res. 2014; 24: 1348 -1355

Plants can survive also with a haploid genome! - reprogramming of male or female gametophyte development in vitro – no gamete formation, but developmental program resembling embryogenesis - usually from immature microspores = androgenesis - female gametophyte = gynogenesis - haploid plants are often sterile - through endoreduplication (by colchicine, oryzalin or spontaneously) - completely homozygous plants – dihaploids Androgenesis in rape seed (pollen embryogenesis)

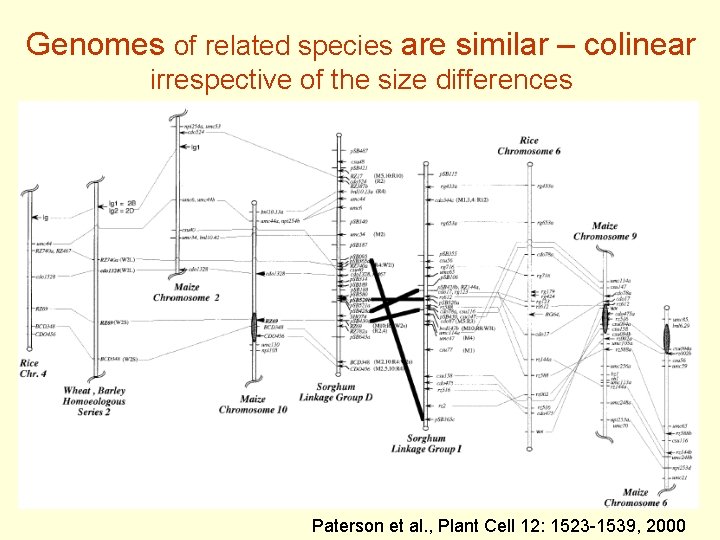
Genomes of related species are similar – colinear irrespective of the size differences Paterson et al. , Plant Cell 12: 1523 -1539, 2000

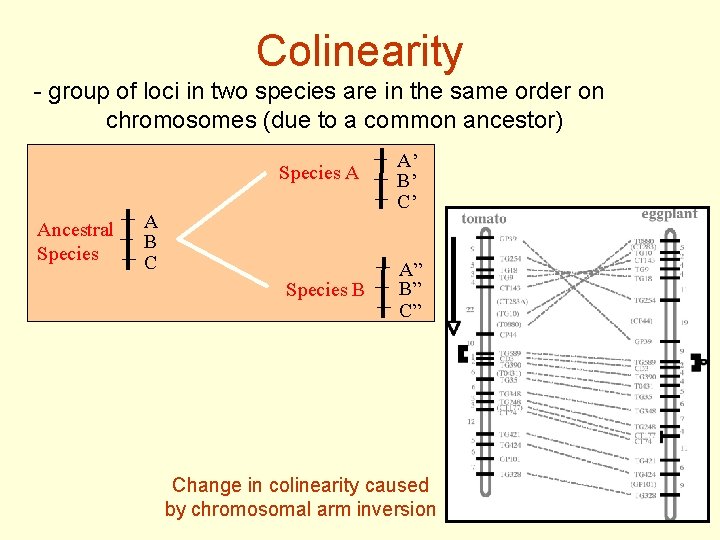
Colinearity - group of loci in two species are in the same order on chromosomes (due to a common ancestor) Ancestral Species A A’ B’ C’ Species B A” B” C” A B C Change in colinearity caused by chromosomal arm inversion

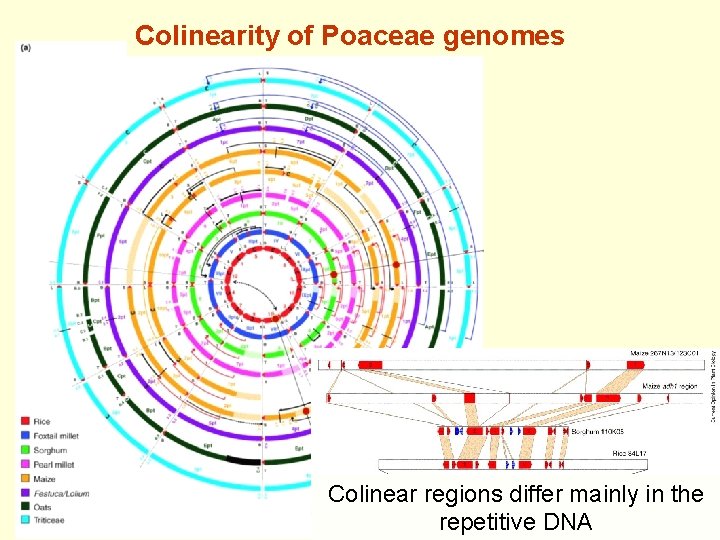
Colinearity of Poaceae genomes Colinear regions differ mainly in the repetitive DNA

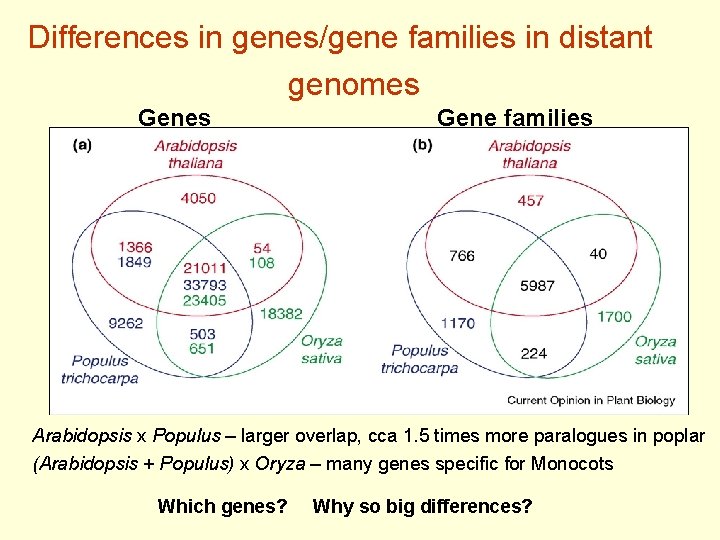
Differences in genes/gene families in distant genomes Gene families Arabidopsis x Populus – larger overlap, cca 1. 5 times more paralogues in poplar (Arabidopsis + Populus) x Oryza – many genes specific for Monocots Which genes? Why so big differences?

Summary: • Current plant genomes result from repeated cycles of partial and complete duplications, followed by reductions and modifications of duplicated sequences. • There are no genomes without redundancy. • Plant genomes are still very dynamic. • High portion of plant genomes consists of repetitive DNA (TEs).