Pipelines Keyboard File Pipe input Program output Screen

Pipelines

-Keyboard -File -Pipe input Program output -Screen -File -Pipe
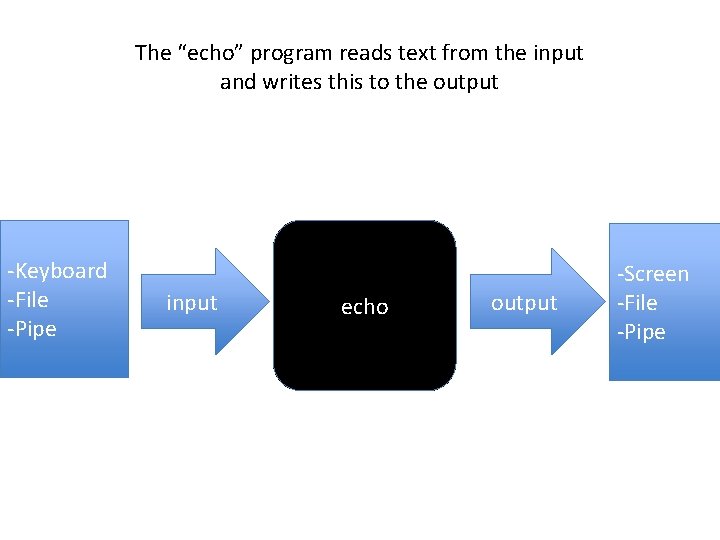
The “echo” program reads text from the input and writes this to the output -Keyboard -File -Pipe input echo output -Screen -File -Pipe
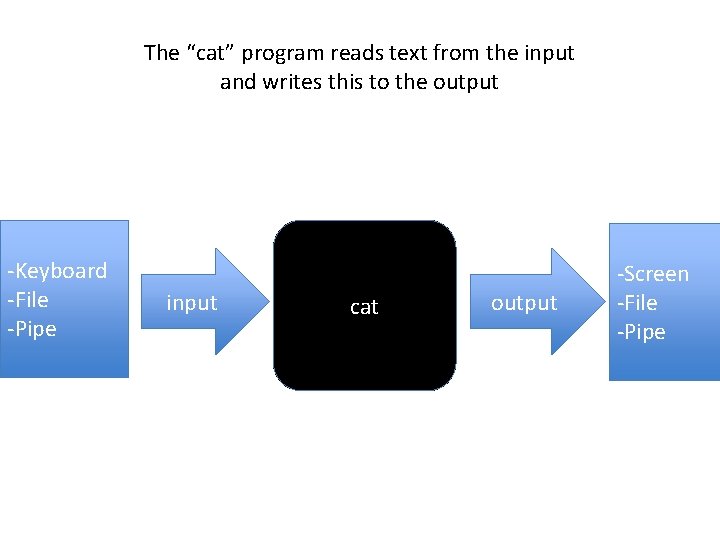
The “cat” program reads text from the input and writes this to the output -Keyboard -File -Pipe input cat output -Screen -File -Pipe

echo uniprot_sprot_plants. fasta

cat uniprot_sprot_plants. fasta >sp|Q 43495|108_SOLLC Protein 108 OS=Solanum lycopersicum PE=2 SV=1 MASVKSSSSSFISLLLLILLVIVLQSQVIECQPQQSCTASLTGLNVCAPFLVPGSP TASTECCNAVQSINHDCMCNTMRIAAQIPAQCNLPPLSCSAN >sp|Q 9 XHP 0|11 S 2_SESIN 11 S globulin seed storage protein 2 OS=Sesamum indicum PE=2 SV=1 MVAFKFLLALSLSLLVSAAIAQTREPRLTQGQQCRFQRISGAQPSLRIQSEGGTTELWDE RQEQFQCAGIVAMRSTIRPNGLSLPNYHPSPRLVYIERGQGLISIMVPGCAETYQVHRSQ RTMERTEASEQQDRGSVRDLHQKVHRLRQGDIVAIPSGAAHWCYNDGSEDLVAVSINDVN HLSNQLDQKFRAFYLAGGVPRSGEQEQQARQTFHNIFRAFDAELLSEAFNVPQETIRRMQ SEEEERGLIVMARERMTFVRPDEEEGEQEHRGRQLDNGLEETFCTMKFRTNVESRREADI FSRQAGRVHVVDRNKLPILKYMDLSAEKGNLYSNALVSPDWSMTGHTIVYVTRGDAQVQV VDHNGQALMNDRVNQGEMFVVPQYYTSTARAGNNGFEWVAFKTTGSPMRSPLAGYTSVIR AMPLQVITNSYQISPNQAQALKMNRGSQSFLLSPGGRRS >sp|P 19084|11 S 3_HELAN 11 S globulin seed storage protein G 3 OS=Helianthus annuus GN=HAG 3 PE=3 S MASKATLLLAFTLLFATCIARHQQRQQQQNQCQLQNIEALEPIEVIQAEAGVTEIWDAYD QQFQCAWSILFDTGFNLVAFSCLPTSTPLFWPSSREGVILPGCRRTYEYSQEQQFSGEGG RRGGGEGTFRTVIRKLENLKEGDVVAIPTGTAHWLHNDGNTELVVVFLDTQNHENQLDEN QRRFFLAGNPQAQAQSQQQQQRQPRQQSPQRQRQGQGQNAGNIFNGFTPELIAQSF NVDQETAQKLQGQNDQRGHIVNVGQDLQIVRPPQDRRSPRQQQEQATSPRQQQEQQQGRR GGWSNGVEETICSMKFKVNIDNPSQADFVNPQAGSIANLNSFKFPILEHLRLSVERGELR PNAIQSPHWTINAHNLLYVTEGALRVQIVDNQGNSVFDNELREGQVVVIPQNFAVIKRAN
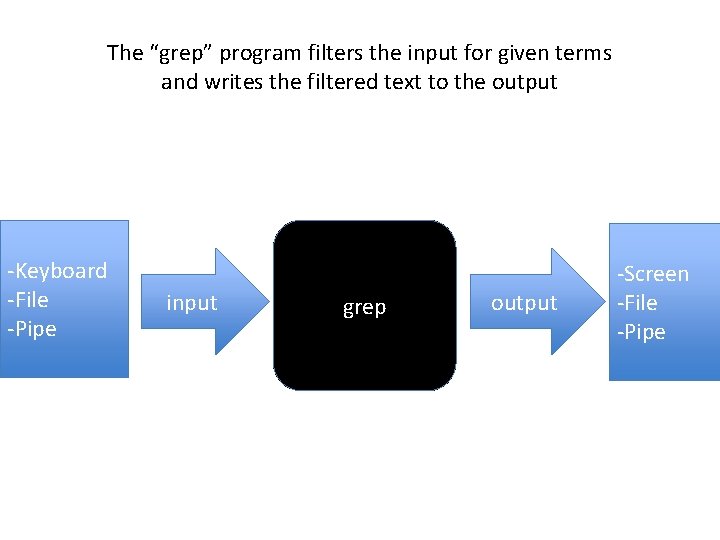
The “grep” program filters the input for given terms and writes the filtered text to the output -Keyboard -File -Pipe input grep output -Screen -File -Pipe
![grep --help Usage: grep [OPTION]. . . PATTERN [FILE]. . . Search for PATTERN grep --help Usage: grep [OPTION]. . . PATTERN [FILE]. . . Search for PATTERN](http://slidetodoc.com/presentation_image_h/db4a6ff4ebd55cb1c0215503b238c34f/image-8.jpg)
grep --help Usage: grep [OPTION]. . . PATTERN [FILE]. . . Search for PATTERN in each FILE or standard input. Example: grep -i 'hello world' menu. h main. c Regexp selection and interpretation: -E, --extended-regexp PATTERN is an extended regular expression -F, --fixed-strings PATTERN is a set of newline-separated strings -G, --basic-regexp PATTERN is a basic regular expression -P, --perl-regexp PATTERN is a Perl regular expression -e, --regexp=PATTERN use PATTERN as a regular expression -f, --file=FILE obtain PATTERN from FILE -i, --ignore-case ignore case distinctions -w, --word-regexp force PATTERN to match only whole words -x, --line-regexp force PATTERN to match only whole lines -z, --null-data a data line ends in 0 byte, not newline

grep sp uniprot_sprot_plants. fasta >sp|Q 43495|108_SOLLC Protein 108 OS=Solanum lycopersicum PE=2 SV=1 >sp|Q 9 XHP 0|11 S 2_SESIN 11 S globulin seed storage protein 2 OS=Sesamum indicum PE=2 >sp|P 19084|11 S 3_HELAN 11 S globulin seed storage protein G 3 OS=Helianthus annuus G >sp|P 13744|11 SB_CUCMA 11 S globulin subunit beta OS=Cucurbita maxima PE=1 SV=1 >sp|Q 05349|12 KD_FRAAN Auxin-repressed 12. 5 k. Da protein OS=Fragaria ananassa PE=2 >sp|O 23878|13 S 1_FAGES 13 S globulin seed storage protein 1 OS=Fagopyrum esculentum >sp|O 23880|13 S 2_FAGES 13 S globulin seed storage protein 2 OS=Fagopyrum esculentum >sp|Q 9 XFM 4|13 S 3_FAGES 13 S globulin seed storage protein 3 OS=Fagopyrum esculentum >sp|P 83004|13 SB_FAGES 13 S globulin basic chain OS=Fagopyrum esculentum PE=1 SV=1 >sp|P 48347|14310_ARATH 14 -3 -3 -like protein GF 14 epsilon OS=Arabidopsis thaliana G >sp|P 93207|14310_SOLLC 14 -3 -3 protein 10 OS=Solanum lycopersicum GN=TFT 10 PE=2 SV >sp|Q 9 S 9 Z 8|14311_ARATH 14 -3 -3 -like protein GF 14 omicron OS=Arabidopsis thaliana G >sp|Q 9 C 5 W 6|14312_ARATH 14 -3 -3 -like protein GF 14 iota OS=Arabidopsis thaliana GN=G >sp|P 42643|14331_ARATH 14 -3 -3 -like protein GF 14 chi OS=Arabidopsis thaliana GN=GR >sp|P 49106|14331_MAIZE 14 -3 -3 -like protein GF 14 -6 OS=Zea mays GN=GRF 1 PE=1 SV=1 >sp|Q 84 J 55|14331_ORYSJ 14 -3 -3 -like protein GF 14 -A OS=Oryza sativa subsp. japonica >sp|P 85938|14331_PSEMZ 14 -3 -3 -like protein 1 (Fragments) OS=Pseudotsuga menziesii >sp|P 93206|14331_SOLLC 14 -3 -3 protein 1 OS=Solanum lycopersicum GN=TFT 1 PE=3 SV=2 >sp|Q 41418|14331_SOLTU 14 -3 -3 -like protein OS=Solanum tuberosum PE=2 SV=1 >sp|Q 01525|14332_ARATH 14 -3 -3 -like protein GF 14 omega OS=Arabidopsis thaliana GN=

Redirection By placing a “>” with a file name at the end of the command line the output can be redirected to a file.

grep sp uniprot_sprot_plants. fasta > out. txt
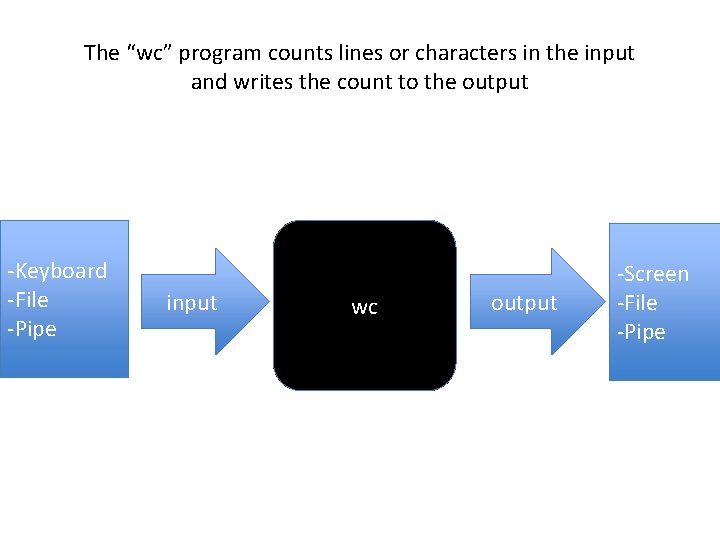
The “wc” program counts lines or characters in the input and writes the count to the output -Keyboard -File -Pipe input wc output -Screen -File -Pipe

wc -l uniprot_sprot_plants. fasta 250177 uniprot_sprot_plants. fasta wc -l out. txt 33851 out. txt

Creating a pipeline With the “|” character the output of one program can be linked to the input of another program
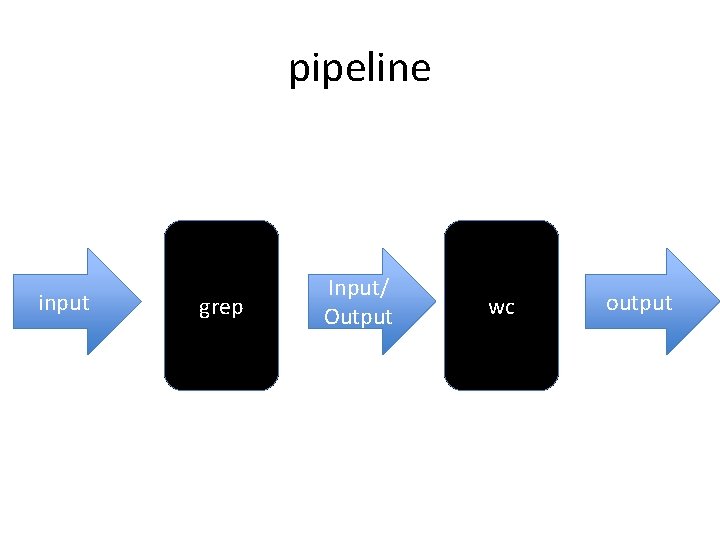
pipeline input grep Input/ Output wc output

grep sp uniprot_sprot_plants. fasta| wc –l 33851

grep sp uniprot_sprot_plants. fasta| grep thaliana >sp|P 48347|14310_ARATH 14 -3 -3 -like protein GF 14 epsilon OS=Arabidopsis thaliana G >sp|Q 9 S 9 Z 8|14311_ARATH 14 -3 -3 -like protein GF 14 omicron OS=Arabidopsis thaliana G >sp|Q 9 C 5 W 6|14312_ARATH 14 -3 -3 -like protein GF 14 iota OS=Arabidopsis thaliana GN=G >sp|P 42643|14331_ARATH 14 -3 -3 -like protein GF 14 chi OS=Arabidopsis thaliana GN=GR >sp|Q 01525|14332_ARATH 14 -3 -3 -like protein GF 14 omega OS=Arabidopsis thaliana GN= >sp|P 42644|14333_ARATH 14 -3 -3 -like protein GF 14 psi OS=Arabidopsis thaliana GN=GR >sp|P 46077|14334_ARATH 14 -3 -3 -like protein GF 14 phi OS=Arabidopsis thaliana GN=GR >sp|P 42645|14335_ARATH 14 -3 -3 -like protein GF 14 upsilon OS=Arabidopsis thaliana G >sp|P 48349|14336_ARATH 14 -3 -3 -like protein GF 14 lambda OS=Arabidopsis thaliana GN >sp|Q 96300|14337_ARATH 14 -3 -3 -like protein GF 14 nu OS=Arabidopsis thaliana GN=GRF >sp|P 48348|14338_ARATH 14 -3 -3 -like protein GF 14 kappa OS=Arabidopsis thaliana GN= >sp|Q 96299|14339_ARATH 14 -3 -3 -like protein GF 14 mu OS=Arabidopsis thaliana GN=GRF >sp|Q 9 LQ 10|1 A 110_ARATH Probable aminotransferase ACS 10 OS=Arabidopsis thaliana GN >sp|Q 9 S 9 U 6|1 A 111_ARATH 1 -aminocyclopropane-1 -carboxylate synthase 11 OS=Arabidops >sp|Q 8 GYY 0|1 A 112_ARATH Probable aminotransferase ACS 12 OS=Arabidopsis thaliana GN >sp|Q 06429|1 A 11_ARATH 1 -aminocyclopropane-1 -carboxylate synthase-like protein 1 O >sp|Q 06402|1 A 12_ARATH 1 -aminocyclopropane-1 -carboxylate synthase 2 OS=Arabidopsis >sp|Q 43309|1 A 14_ARATH 1 -aminocyclopropane-1 -carboxylate synthase 4 OS=Arabidopsis >sp|Q 37001|1 A 15_ARATH 1 -aminocyclopropane-1 -carboxylate synthase 5 OS=Arabidopsis >sp|Q 9 SAR 0|1 A 16_ARATH 1 -aminocyclopropane-1 -carboxylate synthase 6 OS=Arabidopsis
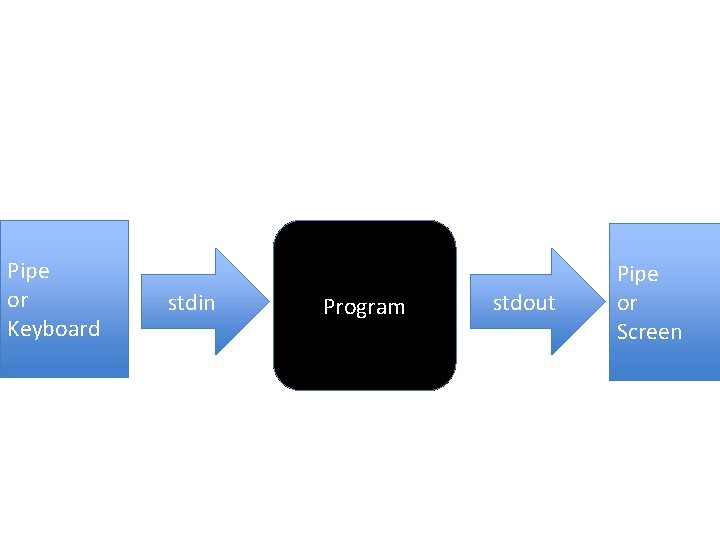
Pipe or Keyboard stdin Program stdout Pipe or Screen
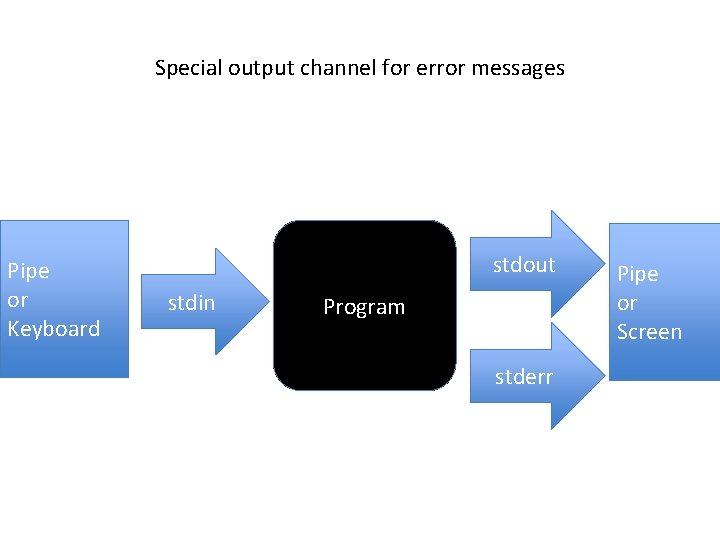
Special output channel for error messages Pipe or Keyboard stdout stdin Program stderr Pipe or Screen

grep sp uniprot_sprot_plants. fas > out. txt grep: uniprot_sprot_plants. fas: No such file or directory

EMBOSS "European Molecular Biology Open Software Suite" http: //emboss. sourceforge. net/ Toolbox with bioinformatics applications
http: //emboss. bioinformatics. nl/

wossname "open reading frame" Finds programs by keywords in their short description SEARCH FOR 'OPEN READING FRAME' getorf Finds and extracts open reading frames (ORFs) plotorf Plot potential open reading frames in a nucleotide sequence

wossname documentation Finds programs by keywords in their short description SEARCH FOR 'DOCUMENTATION' tfm Displays full documentation for an application

tfm getorf Function Finds and extracts open reading frames (ORFs) Description This program finds and outputs the sequences of open reading frames (ORFs) in one or more nucleotide sequences. An ORF may be defined as a region of a specified minimum size between two STOP codons, or between a START and a STOP codon. The ORFs can be output as the nucleotide sequence or as the protein translation. Optionally, the program will output the region around the START codon, the first STOP codon, or the final STOP codon of an ORF. The START and STOP codons are defined in a Genetic Code table; a suitable can be selected for the organism you are investigating. The output is a sequence file containing predicted open reading frames longer than the minimum size, which defaults to 30 bases (i. e. 10 amino acids).

Command line options All EMBOSS programs have a number of command line options. To get started: –help –stdout –filter Get help Write to standard output Read stdin, write stdout
![getorf -help Standard (Mandatory) qualifiers: [-sequence] seqall Nucleotide sequence(s) filename and optional format, or getorf -help Standard (Mandatory) qualifiers: [-sequence] seqall Nucleotide sequence(s) filename and optional format, or](http://slidetodoc.com/presentation_image_h/db4a6ff4ebd55cb1c0215503b238c34f/image-27.jpg)
getorf -help Standard (Mandatory) qualifiers: [-sequence] seqall Nucleotide sequence(s) filename and optional format, or reference (input USA) [-outseq] seqoutall [<sequence>. <format>] Protein sequence set(s) filename and optional format (output USA) Additional (Optional) qualifiers: -table menu [0] Code to use (Values: 0 (Standard); 1 (Standard (with alternative initiation codons)); 2 (Vertebrate Mitochondrial); 3 (Yeast Mitochondrial); 4 (Mold, Protozoan, Coelenterate Mitochondrial and Mycoplasma/Spiroplasma); 5 (Invertebrate Mitochondrial); 6 (Ciliate Macronuclear and Dasycladacean); 9 (Echinoderm Mitochondrial); 10 (Euplotid Nuclear); 11 (Bacterial); 12 (Alternative Yeast Nuclear); 13 (Ascidian Mitochondrial); 14 (Flatworm Mitochondrial); 15 (Blepharisma Macronuclear); 16 (Chlorophycean Mitochondrial); 21 (Trematode Mitochondrial); 22 (Scenedesmus obliquus); 23 (Thraustochytrium Mitochondrial))
![cat example 1. fasta | getorf -filter -find 1 >BTBSCRYR_1 [72 - 110] Bovine cat example 1. fasta | getorf -filter -find 1 >BTBSCRYR_1 [72 - 110] Bovine](http://slidetodoc.com/presentation_image_h/db4a6ff4ebd55cb1c0215503b238c34f/image-28.jpg)
cat example 1. fasta | getorf -filter -find 1 >BTBSCRYR_1 [72 - 110] Bovine m. RNA for lens beta-s-crystallin. . . MTAIATVQISTCT >BTBSCRYR_2 [11 - 544] Bovine m. RNA for lens beta-s-crystallin. . . MSKAGTKITFFEDKNFQGRHYDSDCDCADFHMYLSRCNSIRVEGGTWAVYERPNFAGYMY ILPRGEYPEYQHWMGLNDRLSSCRAVHLSSGGQYKLQIFEKGDFNGQMHETTEDCPSIME QFHMREVHSCKVLEGAWIFYELPNYRGRQYLLDKKEYRKPVDWGAASPAVQSFRRIVE >BTBSCRYR_3 [159 - 590] Bovine m. RNA for lens beta-s-crystallin. . . MKGPILLGTCTSYPGASILSTSTGWASTTASAPAGLFTCLVEASISFRSLRKGILMVRCM RPRKTALPSWSSSTCGRSTPVRCWRAPGSSMSCPTTEAGSTCWTRRSTGSPSTGVQLPQL SSLSAALWSDDTDAAKRWLALSSK >BTBSCRYR_4 [547 - 603] Bovine m. RNA for lens beta-s-crystallin. . . MIQMRPNAGWPCHPNKHYK >BTBSCRYR_5 [618 - 445] (REVERSE SENSE) Bovine m. RNA for lens beta-s-crystallin. . . MPIVLFIMLIWMTRPASVWPHLYHHSTMRRKDWTAGEAAPQSTGFRYSFLSSRYCLPR >BTBSCRYR_6 [381 - 331] (REVERSE SENSE) Bovine m. RNA for lens beta-s-crystallin. . . MWNCSMMEGQSSVVSCI >BTBSCRYR_7 [337 - 197] (REVERSE SENSE) Bovine m. RNA for lens beta-s-crystallin. . . MHLTIKIPFLKDLKLILASTRQVNSPAGAEAVVEAHPVLVLRILAPG >BTBSCRYR_8 [192 - 73] (REVERSE SENSE) Bovine m. RNA for lens beta-s-crystallin. . . MYMYPAKLGLSYTAQVPPSTLMELQRLRYMWKSAQSQSLS

Exercise Make a pipeline that reports (only) the size in residues of the longest protein in this file: uniprot_sprot_plants. fasta It can be done using these applications as building blocks: sizeseq nthseq pepstats grep cut
http: //main. g 2. bx. psu. edu/
- Slides: 30