Phylogenetic Diversity Measures Based on Hill Numbers Anne

Phylogenetic Diversity Measures Based on Hill Numbers Anne Chao National Tsing Hua University Institute of Statistics Hsin-Chu, Taiwan 30043 Eco-Stats Symposium The University of New South Wales Sydney, Australia July 11 -12, 2012
Collaborators of this work: Chun-Huo Chiu National Tsing Hua Univ Taiwan Lou Jost Eco. Minga Foundation Ecuador

Outline: n Traditional Diversity Measures (do not consider species relatedness) Focus on: Doubling property & Hill numbers n Phylogenetic Diversity via Hill Numbers (consider taxonomic or phylogenetic distance between species) Simple Illustrative Examples n Statistical Estimation (brief) n

Bird species diversity

Diversity for the class Crustacea (greatest diversity in the oceans)

Dazzling Orchid Diversity… Some are abundant, some are rare, some still undiscovered

Variance vs. Diversity n Numerical Variables Variance n Categories (Species) Diversity

Biodiversity Definition n n Variety and variability among living organisms and the ecological complexes in which they occur Variation of life at all levels of biological organization

Biodiversity Levels Gene diversity of genes within a species n Species (or taxonomic/phylogenetic) diversity among species in an ecosystem n Ecosystem (or functional) diversity of different ecosystems on Earth. n

Traditional Species Diversity: n n n S species and indexed by 1, 2, . . , S Species absolute abundance/biomass (A 1, A 2, …, AS ) Species relative abundance/biomass (p 1, p 2, …, p. S ) sum=1

Traditional Biodiversity Measures/Indices “Diversity measures” is a diverse issue: Different indices/measures quantify different aspects n Two components - species richness - evenness among abundances n

Species: 4 More diverse Species: 3

Species: 3 Uneven Species: 3 Even More Diverse

4 species Uneven Which one is more diverse? 3 species Even

Gini-Simpson Index (Gini 1912; Simpson 1949) n n (Gini-Simpson Index) Take two individuals, the probability that they belong to different species: (Simpson Index/Concentration, Repeat Rate)

Shannon (1948) entropy Measure of uncertainty in the species identity of a randomly sampled individual

Doubling Property Mac. Arthur (1965), Hill (1973) There are two completely distinct (no overlapped species) communities, each with diversity measure X n Combine these two with equal weight, the diversity should become 2 X An essential minimum requirement for a “diversity” that ecologists expect n “Replication principle” in economics (Dalton 1920): extension to K communities/groups n

What kinds of measures satisfy doubling property? Species richness? Yes!! n Entropy? no!! n Gini-Simpson index? no!! n

n n n Species richness 4 + 4 = 8 Entropy? 1. 39 + 1. 39 > 2. 08 Gini-Simpson index? 0. 75 + 0. 75 > 0. 875
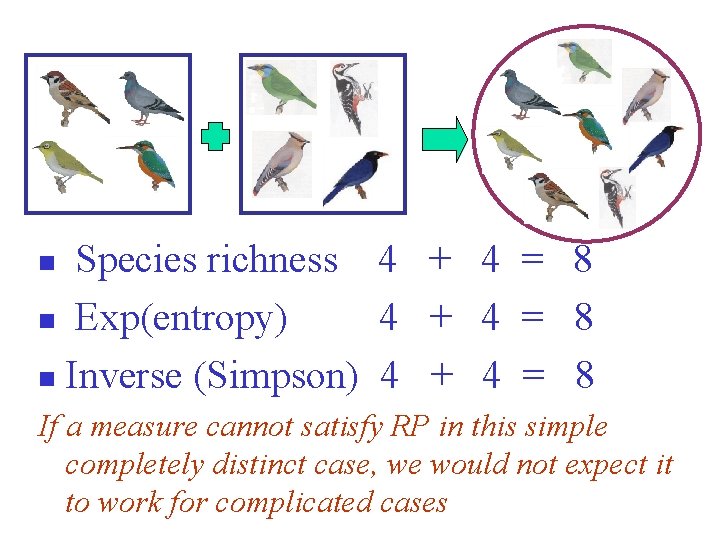
Species richness 4 + 4 = 8 n Exp(entropy) 4 + 4 = 8 n Inverse (Simpson) 4 + 4 = 8 n If a measure cannot satisfy RP in this simple completely distinct case, we would not expect it to work for complicated cases

Hill’s (1973) Family of Diversity Indices of order q q = 0, 0 D = species richness n q =1, 1 D = exponential of entropy n n q = 2, 2 D = inverse of Simpson index

“Order” q (Tsallis 2001; Keylock 2005) The order q determines the measure’s sensitivity to species frequencies n q > 1, sensitive to common species n q < 1, sensitive to rare species n q = 1, weighs species by their frequencies, without favoring either common or rare species

Hill numbers: transform to units of “species” n Entropy = 1. 39 is equivalent to exp(1. 39) = 4 “species” n Gini-Simpson index = 0. 75 is equivalent to 1/(1 -0. 75) = 4 “species”

Hill Numbers: “Species Equivalent” n “Effective number of species” The number of equally-common species that would be needed to give the same diversity as the community in study n For equally-common community, Hill numbers are equal to species richness for all orders of q;

“Effective number of species” Community: S species {p 1, p 2, …, p. S} Hill numbers = D for an order q Simple Community: D species with equal relative abundances {1/D, …, 1/D} 25

Hill numbers: An intuitive equivalence Simple community: Complex community: p 1 p 2 … … p. S = Then

Examples: four hypothetical communities There are 100 species, 500 individuals A: equally-common B: slightly uneven C: moderately uneven D: highly uneven
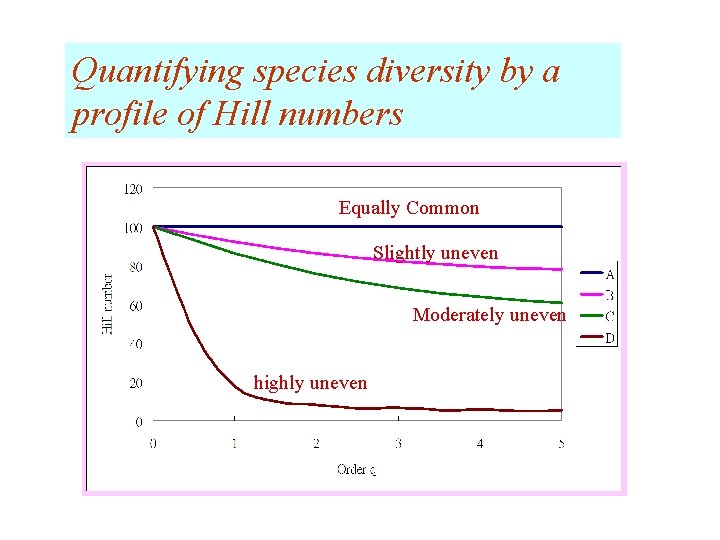
Quantifying species diversity by a profile of Hill numbers Equally Common Slightly uneven Moderately uneven highly uneven

Diversity partitioning via Hill numbers n n n Partitioning gamma (regional) diversity into alpha (within-community) diversity and beta (between-community) diversity Intense debates on additive or multiplicative? Chao et al. (2012) proposed a resolution that both converge to the same classes of similarity measures: Jaccard, Sorenson (q = 0), Horn (q=1) and Morisita-Horn similarity measures (q =2)

Phylogenetic Diversity: All else being equal, which community is more diverse? Community 1 Community 2

Community 1 n n Community 2 Species in community 2 is more phylogenetically diverse than community 1 Pielou (1975, p. 17) was the first to notice the concept of diversity could be broadened to consider taxonomic difference between species.

“I think” Tree of Life The first-known sketch by Charles Darwin of an evolutionary tree describing the relationships among groups of organisms http: //www. amnh. org/exhibitio ns/darwin/idea/treelg. php

Phylogenetic Diversity Measures: We not only consider the relative abundance of species, p 1 p 2 p 3 but also the phylogenetic relationship among species. p 1 p 2 p 3 And, satisfy the essential requirement “replication principle”.

Doubling Property for Phylogenetic Diversity n n n Two completely phylogenetically distinct assemblages (no shared lineages), with the same phylogenetic diversity =X. Assemblages are pooled in equal proportions, then the pooled assemblage has phylogenetic diversity 2 X. Similar extension to N assemblages

Doubling Property in phylogenetic version n n Two completely phylogenetically distinct (no overlapped tree branch) across assemblages, each with diversity measure X Combine these two, the diversity becomes 2 X 35

Pioneering Work in phylogenetic diversity (1) Branch-length-based measure: n Phylogenetic Diversity PD (Faith 1992) sum of the branch lengths of the phylogeny connecting all species from tips to root Satisfy “replication principle”.
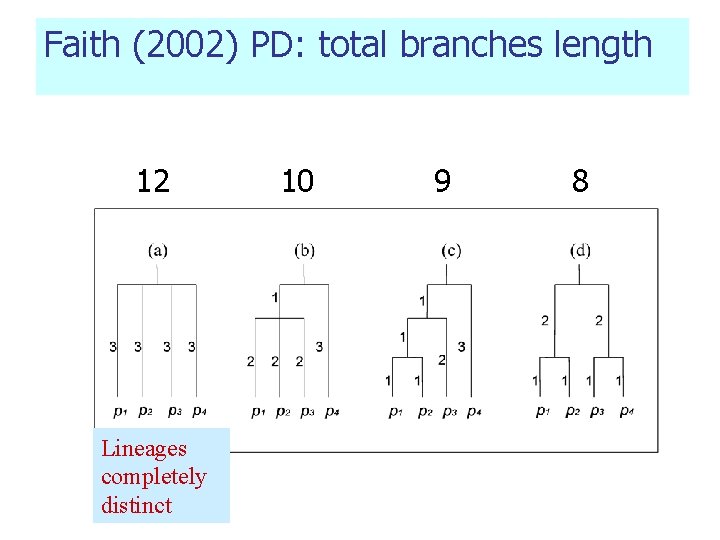
Faith (2002) PD: total branches length 12 Lineages completely distinct 10 9 8

Pioneering work (2) n n n Weitzman (1992, 1993, 1998) from a perspective of economic theory of biodiversity preservation “Unfortunately, Noah’s Ark has a limited capacity…. and a (limited) budget available for biodiversity preservation…” What to preserve?

The Noah’s Ark: the agony of choice The woodpecker might have to go! Courtesy of Ramon Teja, http: //www. livepencil. com/

Traditional n Species richness n Entropy n Gini-Simpson n Hill Numbers Phylogenetic Faith PD (Faith 1992) Phylogenetic entropy (Allen et al. 2009) Quadratic entropy (Rao 1982) Chao, Chiu and Jost (2010)

Pioneering Work (3) n Quadratic entropy (Rao 1982) dij : phylogenetic distance between species i and j, pi and pj denote species relative abundance of species i and j. Q: mean phylogenetic distance between any two randomly chosen individuals in a community n Phylogenetic entropy (Allen et. al. 2009) Li : length of branch i, ai : the abundance descending from branch i. n A parametric class based on Tsallis entropy (Pavoine et. al. 2009) I 0 = Faith’s PD minus the tree height I 1 = phylogenetic entropy Hp I 2 = Rao’s Q measure

Phylogenetic diversity measures Except for Faith’s PD, all indices mentioned above do NOT satisfy the “replication principle”. (Need transformations!) n n. Chao et al. (2010) were motivated to develop a unified class of phylogenetic diversity measures based on Hill numbers Satisfy “replication principle”
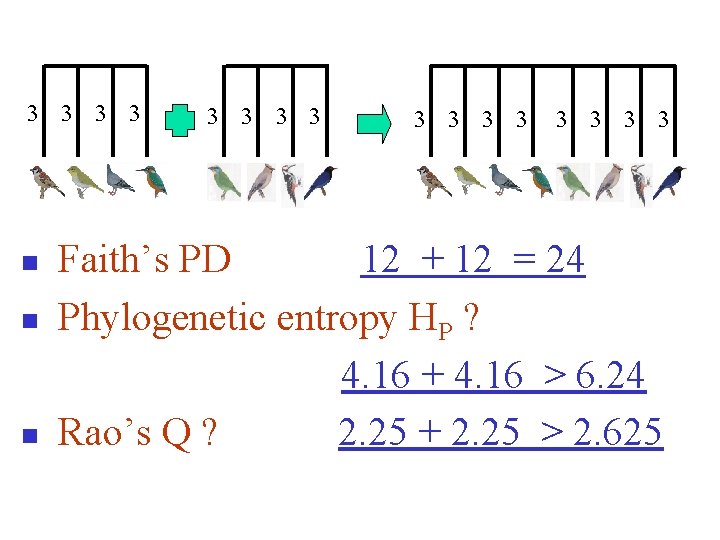
3 3 n n n 3 3 3 Faith’s PD 12 + 12 = 24 Phylogenetic entropy HP ? 4. 16 + 4. 16 > 6. 24 Rao’s Q ? 2. 25 + 2. 25 > 2. 625

Phylogenetic Diversity Measures: Two parameters: n Order q in Hill number slice 3 L 7 p 1+p 2+p 3+p 4 slice 2 L 6 n Time parameter T: Consider the phylogenetic diversity through T years ago L 4 p 1+p 2+p 3 L 5 L 1 p 2+p 3 L 2 slice 1 L 3 t=0 (Present time) p 1 p 2 p 3 p 4

Basic approach based on Hill Numbers for shared lineages n At any given moment t , slice the tree, we can find the lineage (branch cuts, “species”) and its relative abundance (measure of their importance in the present-day community) n Obtain Hill number q. D(t) at moment t. n Average over from the present time to T years ago n Call this average diversity as “Mean Diversity of order q over T years”, it is in units of “lineage” (or “species”).

Conceptual framework for q = 0 Connect Faith’s PD to mean species richness n For a fixed T, the nodes divide the phylogenetic tree into Segment 1, 2 and 3 with duration (length) T 1, T 2, and T 3 n In any moment of Segment 1, there are 4 lineages (i. e. , 4 branches cut) n Segment 2, there are 3 lineages n Segment 3, there are 2 lineages The mean lineage (species) richness over the time interval [−T, 0] is (T 1/T) × 4 + (T 2/T) × 3 + (T 3/T) × 2 = total branch length in [-T, 0] / T (Mean Phylogenetic Diversity of order 0 over T years) If T = height of tree, then

Conceptual framework for q > 0 To incorporating abundance, use lineage abundance: sum of the relative abundances descended from the branch n There are T 1 assemblages with abundance vector{p 1, p 2, p 3, p 4 }, n T 2 assemblages with abundance vector {p 1, p 2+p 3, p 4 } and n T 3 assemblages with abundance vector {p 1+p 2+p 3, p 4 }. There a total of T 1+T 2+T 3 = T assemblages and each is given the same weight 1/T. The “Mean diversity of order q over T years” is the following average

Mean Phylogenetic Diversity of order q over T years General Formula BT : all branches in the time interval [-T, 0] Li : the length (duration) of Branch i in the set BT ai : the total relative abundance descended from Branch i

Interpretation of mean diversity n n Mean effective number of completely distinct lineages (species) over T years Link to traditional diversity: When all species are completely equally distinct with branch lengths T (including T = 0, ignoring phylogeny)
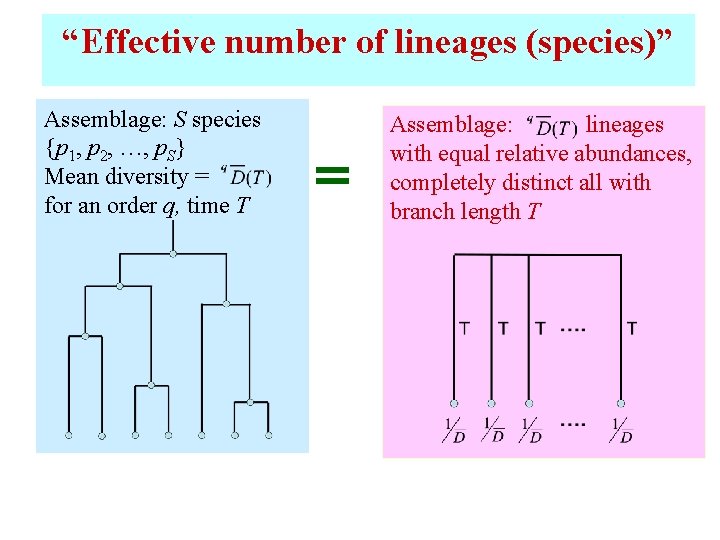
“Effective number of lineages (species)” Assemblage: S species {p 1, p 2, …, p. S} Mean diversity = for an order q, time T Assemblage: lineages with equal relative abundances, completely distinct all with branch length T

Related Measure: Branch Diversity n n q = 0, branch diversity reduces to Faith’s PD Branch diversity: the amount of evolutionary “work” done on the assemblage or the effective lineageyears or lineage-length (or other units) contained in the tree in the time period [−T, 0]

Generalize and unify existing measures: n Order q = 0 = Total branch lengths in [-T, 0] / T n Order q =1 n Order q = 2
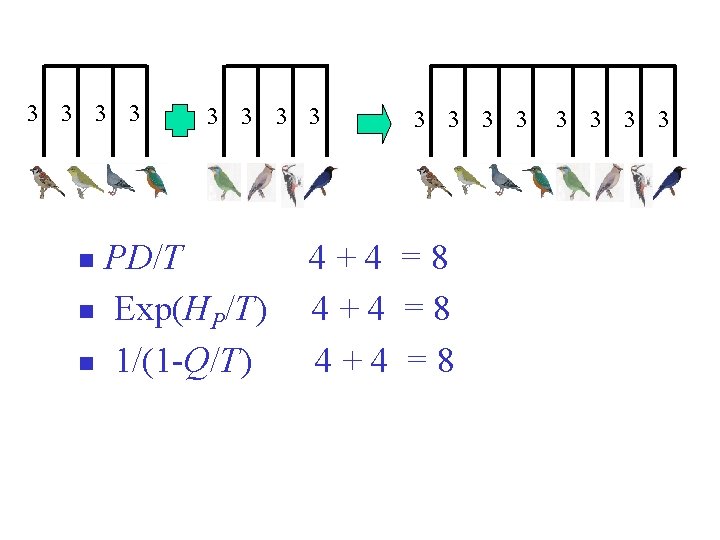
3 3 3 3 PD/T n Exp(HP/T) n 1/(1 -Q/T) n 3 3 4+4 =8 3 3
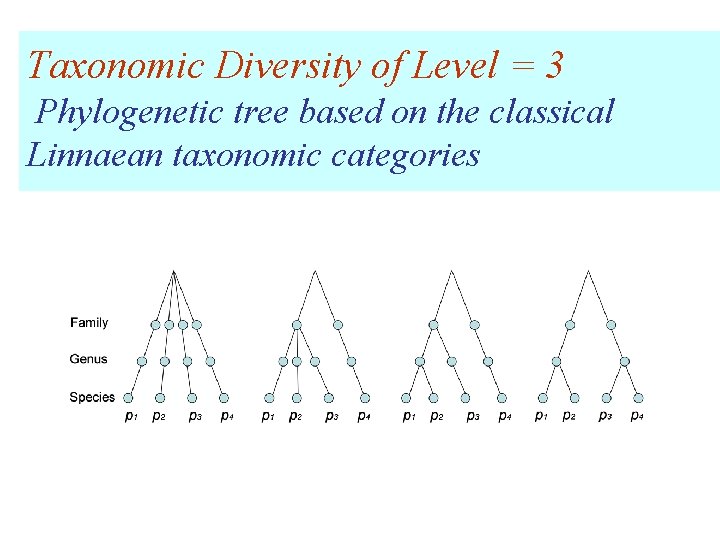
Taxonomic Diversity of Level = 3 Phylogenetic tree based on the classical Linnaean taxonomic categories
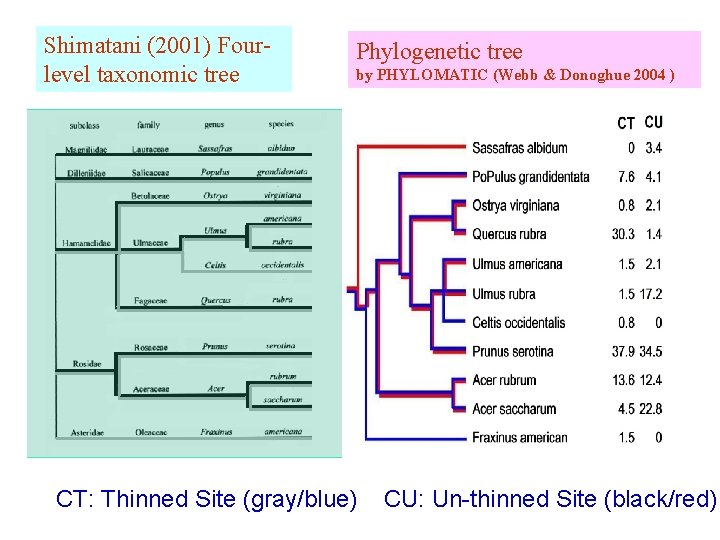
Shimatani (2001) Fourlevel taxonomic tree Phylogenetic tree by PHYLOMATIC (Webb & Donoghue 2004 ) CT: Thinned Site (gray/blue) CU: Un-thinned Site (black/red)
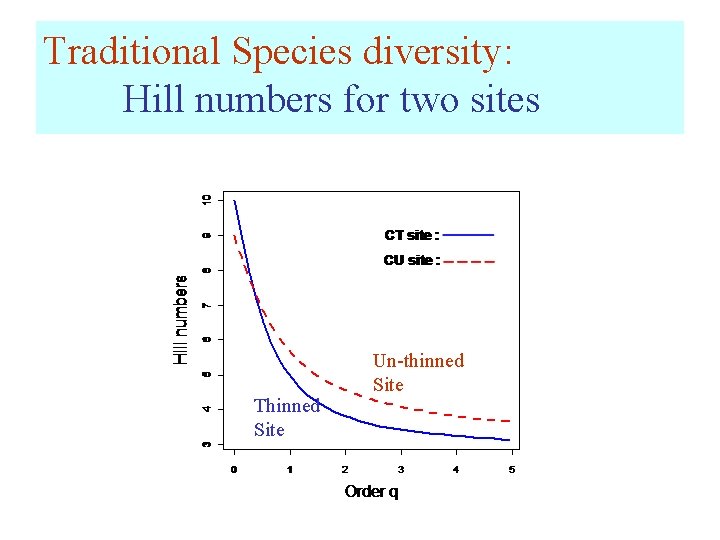
Traditional Species diversity: Hill numbers for two sites Thinned Site Un-thinned Site
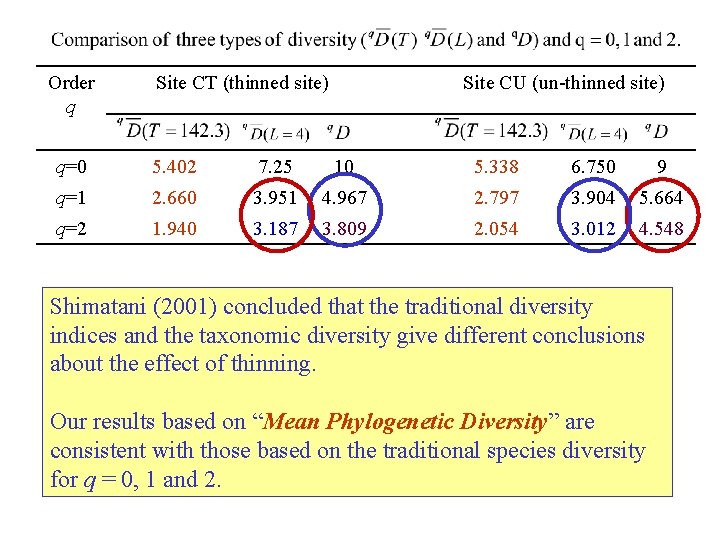
Order q Site CT (thinned site) Site CU (un-thinned site) q=0 5. 402 7. 25 10 5. 338 6. 750 9 q=1 2. 660 3. 951 4. 967 2. 797 3. 904 5. 664 q=2 1. 940 3. 187 3. 809 2. 054 3. 012 4. 548 Shimatani (2001) concluded that the traditional diversity indices and the taxonomic diversity give different conclusions about the effect of thinning. Our results based on “Mean Phylogenetic Diversity” are consistent with those based on the traditional species diversity for q = 0, 1 and 2.

Diversity profile n n n Non-phylogenetic: Use a profile of Hill numbers (as a function of order q) to quantify diversity of a community Phylogenetic: Use three profiles (q = 0, 1, 2); each is a function of time T to quantify phylogenetic diversity All these measures satisfy “doubling property”
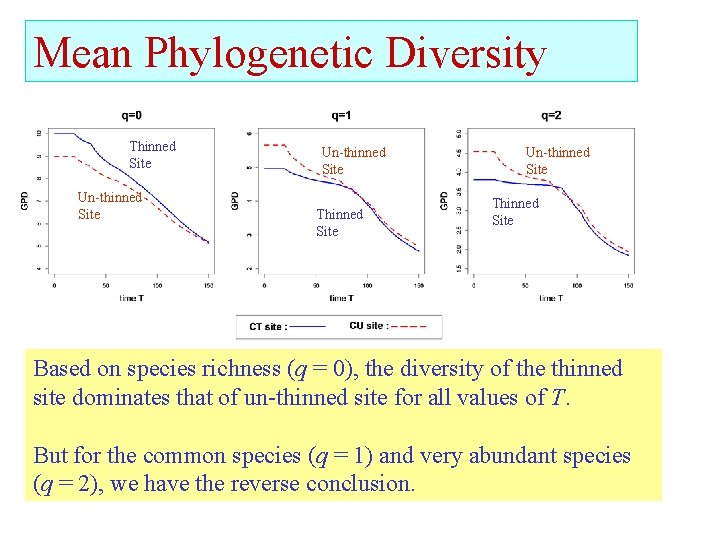
Mean Phylogenetic Diversity Thinned Site Un-thinned Site Thinned Site Based on species richness (q = 0), the diversity of the thinned site dominates that of un-thinned site for all values of T. But for the common species (q = 1) and very abundant species (q = 2), we have the reverse conclusion.

Extensions n n The general cases of non-ultrametric trees Partitioning phylogenetic Hill numers: phylogenetic alpha, beta, gamma diversity measures and related similarity measures (Chiu, Jost & Chao 2013) Extension to dendrogram-based functional diversity (Petchey and Gaston, 2002) Extension to distance-based functional diversity

Statistical Estimation for traditional diversity measures Depends on the order q Non-surprisingly Non-trivial n q = 0 “species richness estimation” n q = 1 “Shannon entropy estimation” and its exponential Surprisingly Non-trivial n q = 2 widely used in genetics (gene identity, or heterozygosity) Non-surprisingly Nearly unbiased estimator exists trivial

q = 0 “species richness estimation” Since Fisher, Corbert and Williams (1943) n Curve fitting (fitting a parametric curve to SAC) n Parametric models for species abundances n Non-parametric approach n Rarefaction/extrapolation of species accumulation curve (by estimating expected species richness for a finite size sample or sample completeness

q = 1 “Entropy estimation” Since Shannon (1948) n Traditional bias-reduction n Jackknife for bias-reduction n Bayesian approaches n Coverage-adjusted estimator n Estimation via Renyi’s entropies n Polynomial representation

Other Related Estimation Issues n n Hill numbers: Estimation of gamma, alpha and beta diversity and related similarity/differentiation measures Their phylogenetic generalization

Main References: n n n Chao, A. , Chiu C. -H. and Jost, L. (2010). Phylogenetic diversity measures based on Hill numbers. Philosophical Transactions of the Royal Society B. , 365, 3599 -3609. Chiu, C. -H. , Jost, L. and Chao, A. (2013). Phylogenetic beta diversity, similarity, and differentiation measures based on Hill numbers. To appear in Ecological Monographs. Chao, A. , Gotelli, N. G. , Hsieh, T. C. , Sander, E. L. , Ma, K. H. , Colwell, R. K. and Ellison, A. M. (2013). Rarefaction and extrapolation with Hill numbers: a framework for sampling and estimation in species biodiversity studies. To appear in Ecological Monographs.

Nanney (2004) “We are all blind men (and women) trying to describe a monstrous elephant of ecological and evolutionary diversity. . . ”

Heaven is under our feet as well as over our heads Henry David Thoreau , Writer and Naturalist (1817 -1862) THANK YOU VERY MUCH!!
- Slides: 67