Phusion Assembler and Its Application in Schistosome Genomes

Phusion Assembler and Its Application in Schistosome Genomes By Zemin Ning Informatics Division The Wellcome Trust Sanger Institute
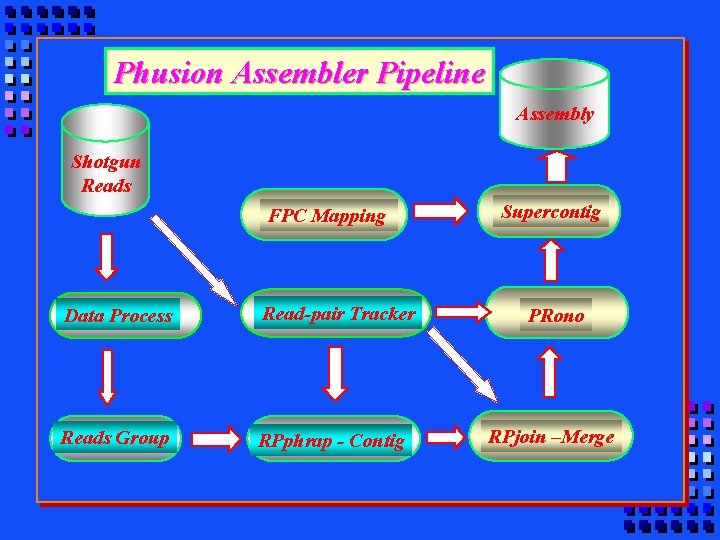
Phusion Assembler Pipeline Assembly Shotgun Reads FPC Mapping Data Process Read-pair Tracker Reads Group RPphrap - Contig Supercontig PRono RPjoin –Merge
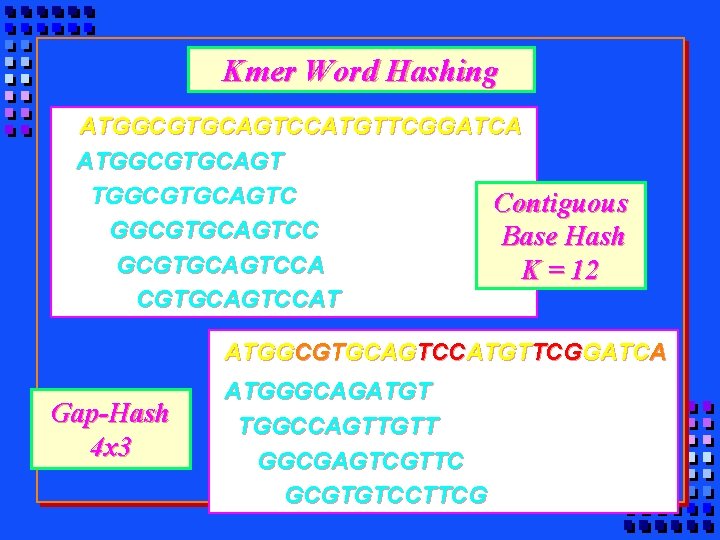
Kmer Word Hashing ATGGCGTGCAGTCCATGTTCGGATCA ATGGCGTGCAGTC Contiguous GGCGTGCAGTCC Base Hash GCGTGCAGTCCA K = 12 CGTGCAGTCCAT ATGGCGTGCAGTCCATGTTCGGATCA Gap-Hash 4 x 3 ATGGGCAGATGT TGGCCAGTTGTT GGCGAGTCGTTC GCGTGTCCTTCG
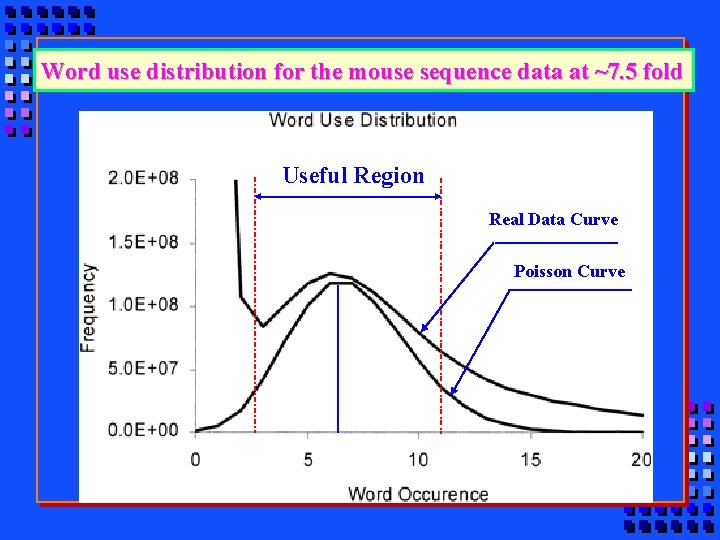
Word use distribution for the mouse sequence data at ~7. 5 fold Useful Region Real Data Curve Poisson Curve
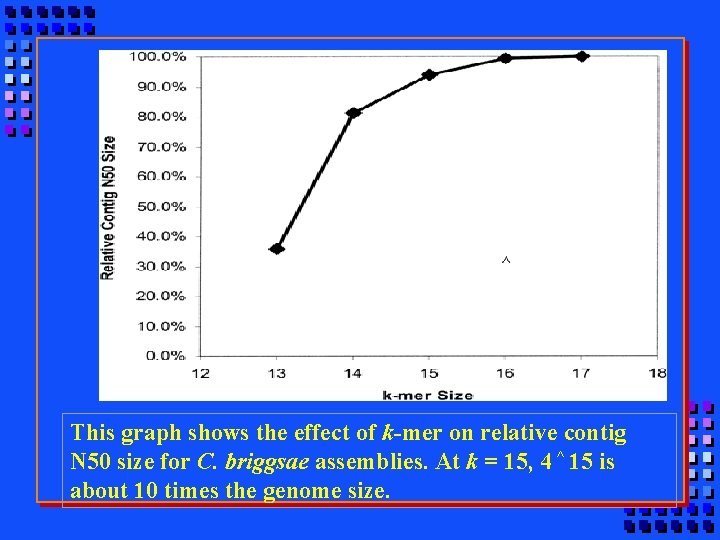
This graph shows the effect of k-mer on relative contig N 50 size for C. briggsae assemblies. At k = 15, 4 ^ 15 is about 10 times the genome size.

Unique and Repetitive DNA Sections A Depth X’ Unique Section B X’’ C Repetitive Section Depth
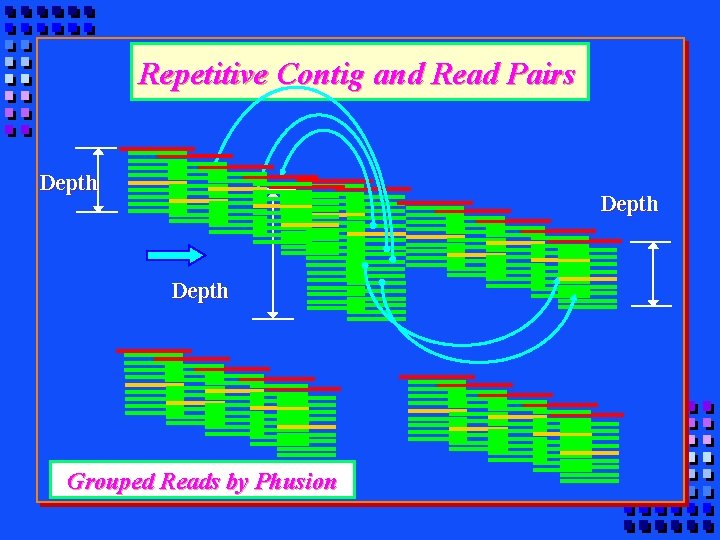
Repetitive Contig and Read Pairs Depth Grouped Reads by Phusion
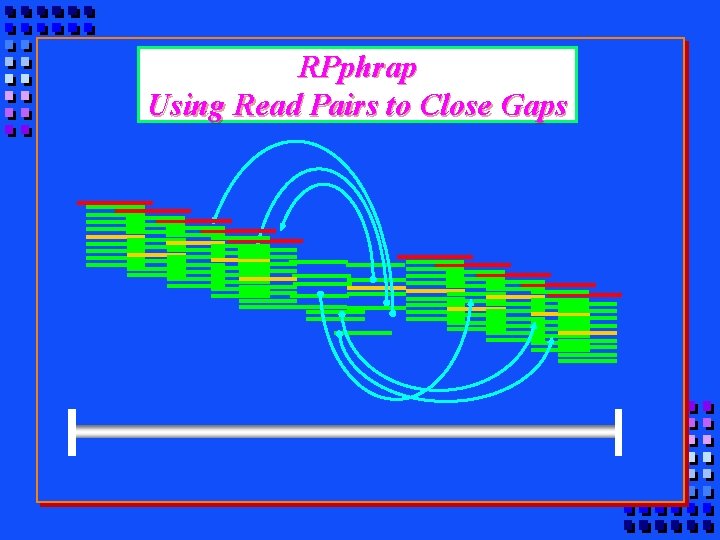
RPphrap Using Read Pairs to Close Gaps
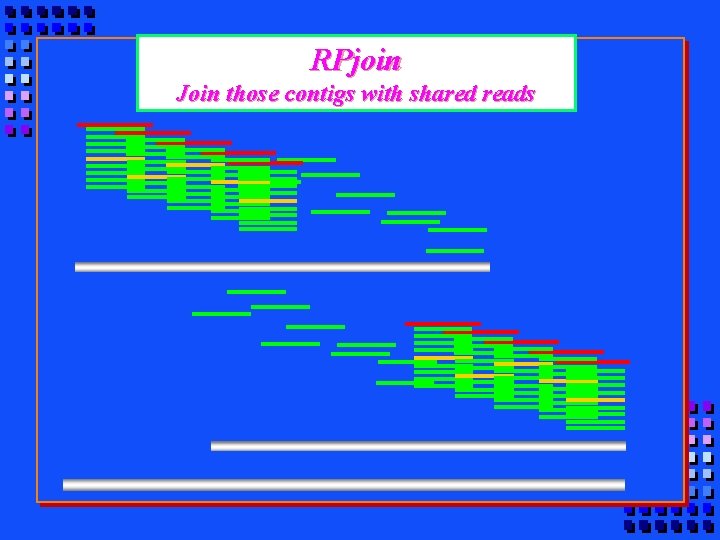
RPjoin Join those contigs with shared reads

S. Mansoni WGS Assembly WGS reads: Number of shotgun reads: Estimated genome size: Estimated read coverage: Number of reads placed: Ratio of placed reads: 3, 362, 074; 380 Mbp; ~7. 5 X; 2, 725, 204; 81. 1 %; Assembly features: - contig stats Total number of contigs: Total bases of contigs: N 50 contig size: Averaged contig size: Contig coverage over the genome: 50, 376; 375. 7 MB 16, 315; 7, 457; >95 % ? ; 8 Assembly: - supercontig stats Total number of supercontigs: Total bases of supercontigs: N 50 supercontig size: Averaged supercontig size: Supercontig coverage over the genome: 19, 031; 381. 8 MB; 824, 505; 20, 006; >95 % ?

Acknowledgements: q q q Yong Gu Jim Mullkin Matthew Berriman Jane Rogers Sanger Systems Support Sanger Sequencing Facilities
- Slides: 11