Phusion 2 and The Genome Assembly of Tasmanian

Phusion 2 and The Genome Assembly of Tasmanian Devil Zemin Ning The Wellcome Trust Sanger Institute

Outline of the Talk: q q q Challenges in genome assemblies from pure Illumina reads The Phusion 2 pipeline The Tasmanian devil genome project The Devil genome assembly Other assemblies: human cancer, zebrafish, rice, etc

Challenges in Whole Genome Assembly using Pure Illumina Reads Ø Large genome and huge datasets For human: 100 Gb at 30 x Ø Repetitive/Duplication structures, Alus, LINES, SVAs 30 -40% such as human, mouse; 50 -60% such as rice and other plant genomes. Ø Tandem repeats: how many copies they have? TATATATATATATA GCGCGCGCGCGCGCGCG GTGTGTGTGTGTGTGTGTG AGTAGTAGTAGTAGTAGTAGT
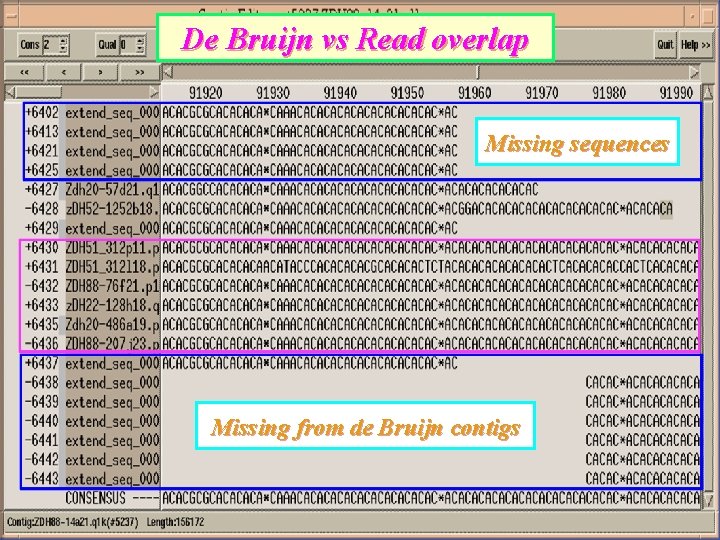
De Bruijn vs Read overlap Missing sequences Missing from de Bruijn contigs
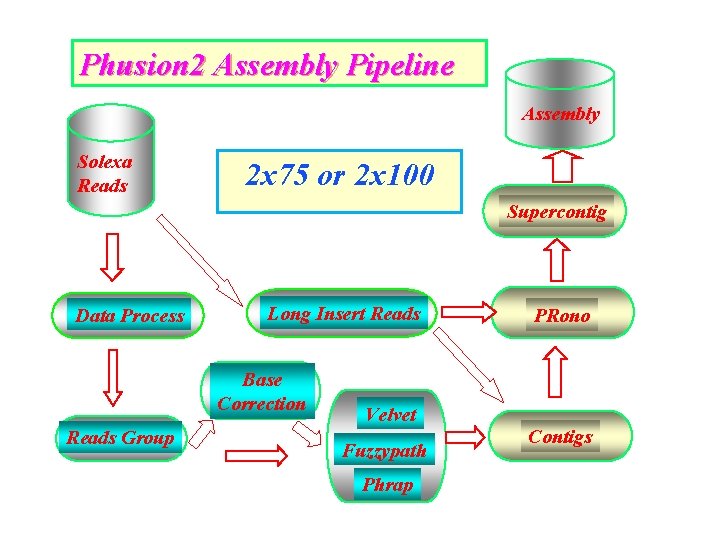
Phusion 2 Assembly Pipeline Assembly Solexa Reads 2 x 75 or 2 x 100 Supercontig Data Process Long Insert Reads Base Correction Reads Group PRono Velvet Fuzzypath Phrap Contigs
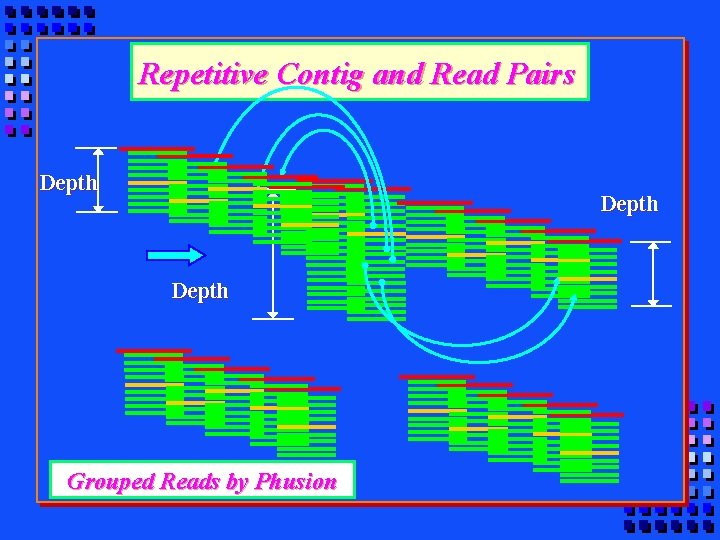
Repetitive Contig and Read Pairs Depth Grouped Reads by Phusion
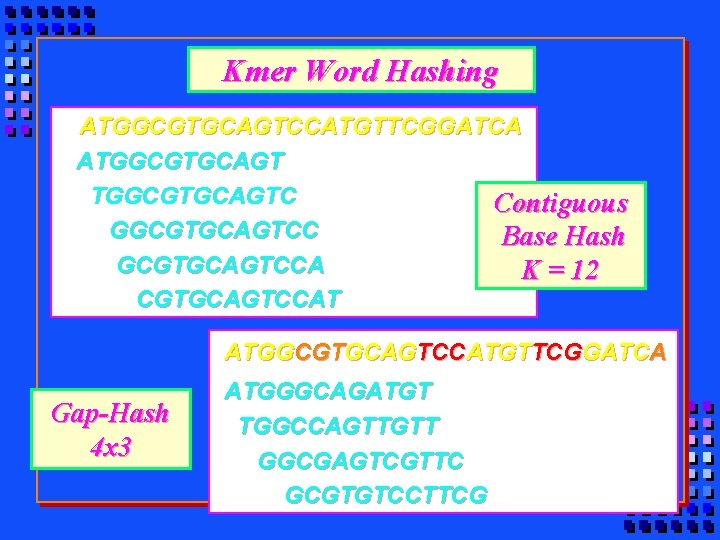
Kmer Word Hashing ATGGCGTGCAGTCCATGTTCGGATCA ATGGCGTGCAGTC Contiguous GGCGTGCAGTCC Base Hash GCGTGCAGTCCA K = 12 CGTGCAGTCCAT ATGGCGTGCAGTCCATGTTCGGATCA Gap-Hash 4 x 3 ATGGGCAGATGT TGGCCAGTTGTT GGCGAGTCGTTC GCGTGTCCTTCG
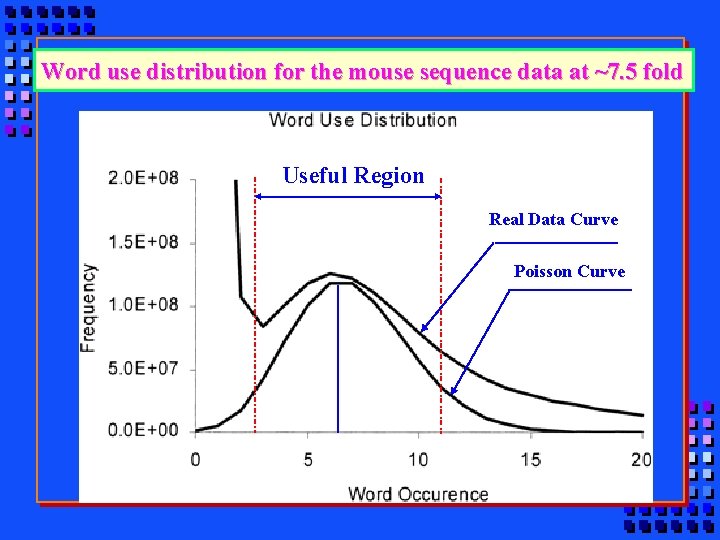
Word use distribution for the mouse sequence data at ~7. 5 fold Useful Region Real Data Curve Poisson Curve
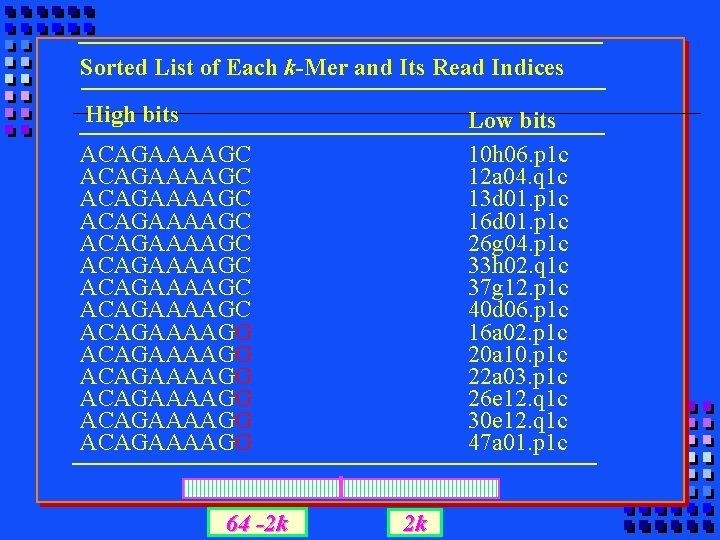
Sorted List of Each k-Mer and Its Read Indices High bits Low bits 10 h 06. p 1 c 12 a 04. q 1 c 13 d 01. p 1 c 16 d 01. p 1 c 26 g 04. p 1 c 33 h 02. q 1 c 37 g 12. p 1 c 40 d 06. p 1 c 16 a 02. p 1 c 20 a 10. p 1 c 22 a 03. p 1 c 26 e 12. q 1 c 30 e 12. q 1 c 47 a 01. p 1 c ACAGAAAAGC ACAGAAAAGC ACAGAAAAGG ACAGAAAAGG 64 -2 k 2 k

Relation Matrix: R(i, j) – number of kmer words shared between read i and read j 1 2 3 4 5 6 … j … N 1 2 3 4 5 6 41 0 0 41 37 0 0 37 0 22 0 0 0 27 0 0 22 0 0 27 0 Group 2: (4, 6) i N Group 1: (1, 2, 3, 5) R(i, j)
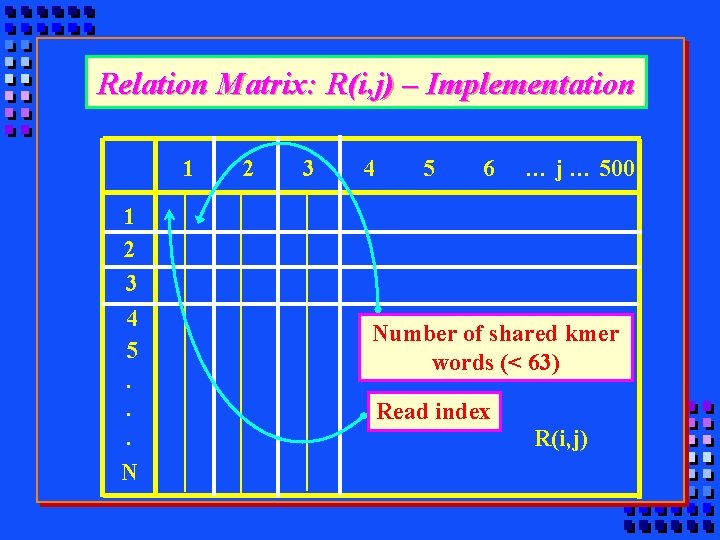
Relation Matrix: R(i, j) – Implementation 1 2 3 4 5 6 … j … 500 1 2 3 4 5. . . N Number of shared kmer words (< 63) Read index R(i, j)
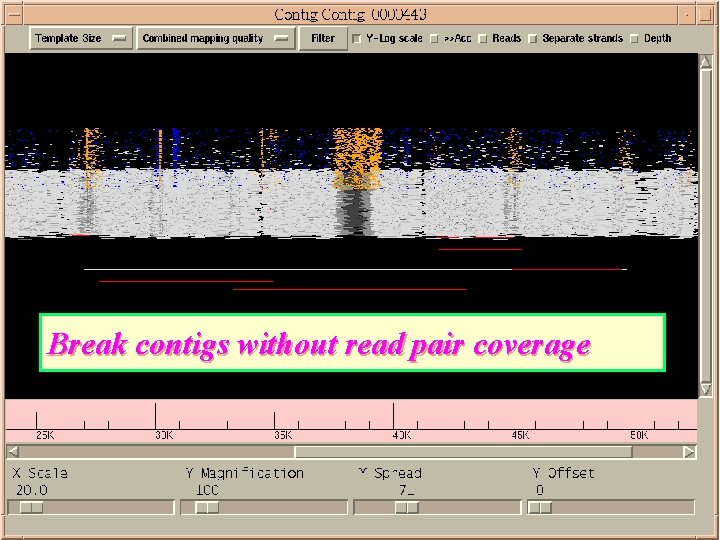
Break contigs without read pair coverage
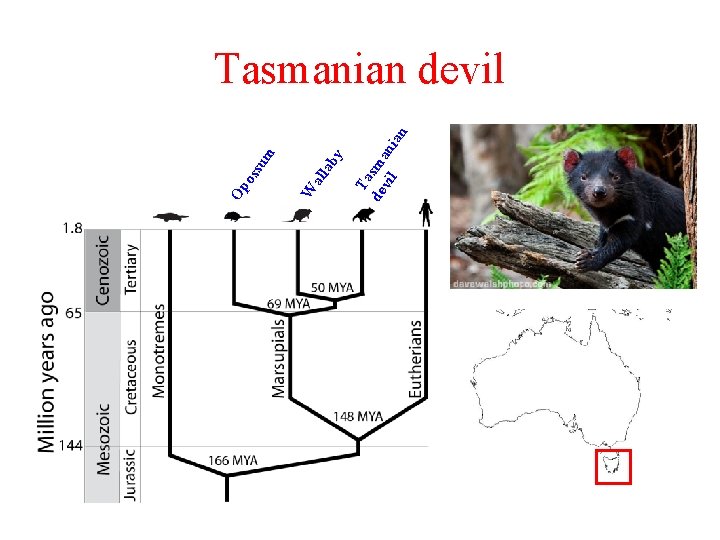
Ta de sma vil n ian ab y all W su m Op os Tasmanian devil
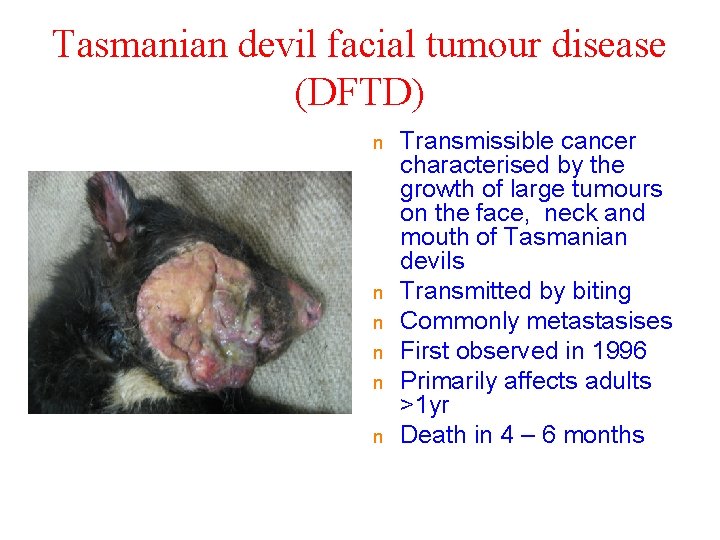
Tasmanian devil facial tumour disease (DFTD) n n n Transmissible cancer characterised by the growth of large tumours on the face, neck and mouth of Tasmanian devils Transmitted by biting Commonly metastasises First observed in 1996 Primarily affects adults >1 yr Death in 4 – 6 months
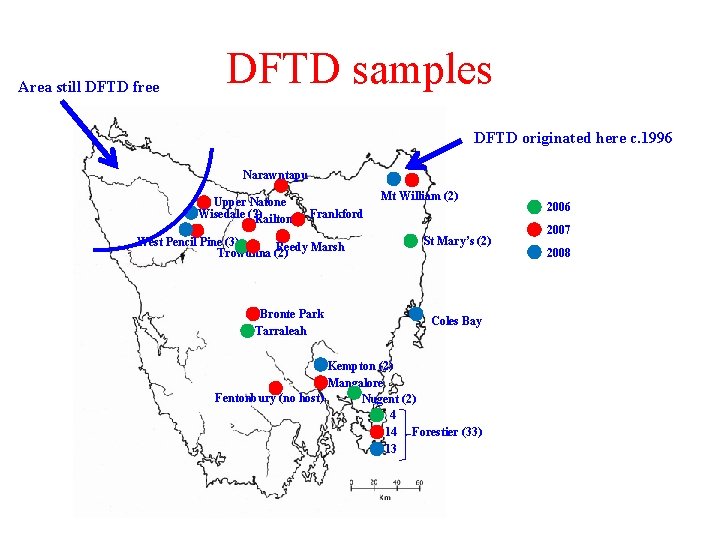
Area still DFTD free DFTD samples DFTD originated here c. 1996 Narawntapu Upper Natone Wisedale (? ) Railton Mt William (2) Frankford West Pencil Pine (3) Reedy Marsh Trowunna (2) Bronte Park Tarraleah St Mary’s (2) Coles Bay Kempton (2) Mangalore Fentonbury (no host) Nugent (2) 4 14 Forestier (33) 13 2006 2007 2008
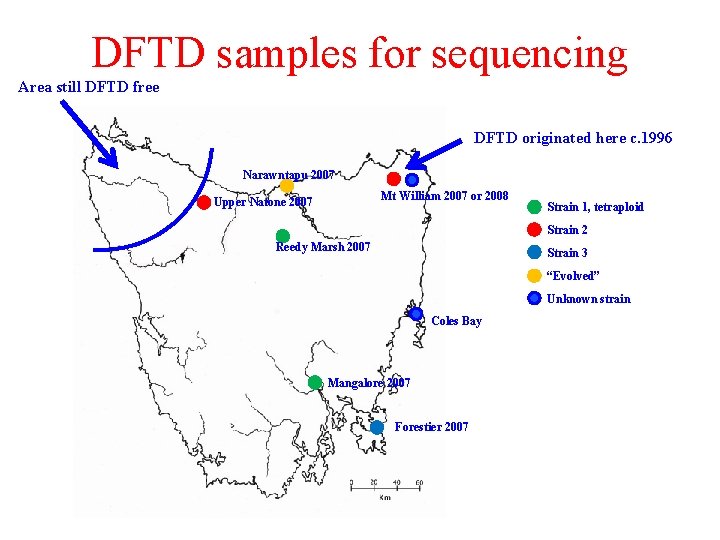
DFTD samples for sequencing Area still DFTD free DFTD originated here c. 1996 Narawntapu 2007 Mt William 2007 or 2008 Upper Natone 2007 Strain 1, tetraploid Strain 2 Reedy Marsh 2007 Strain 3 “Evolved” Unknown strain Coles Bay Mangalore 2007 Forestier 2007
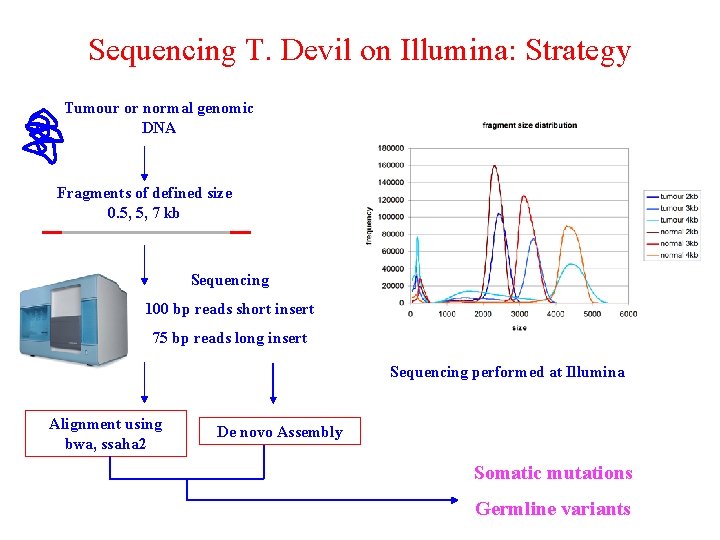
Sequencing T. Devil on Illumina: Strategy Tumour or normal genomic DNA Fragments of defined size 0. 5, 5, 7 kb Sequencing 100 bp reads short insert 75 bp reads long insert Sequencing performed at Illumina Alignment using bwa, ssaha 2 De novo Assembly Somatic mutations Germline variants

Paired Reads Separated by “NN”
Error Bases Correction

Genome Assembly – T. Devil Solexa reads: Number of read pairs: Finished genome size: Read length: Estimated read coverage: Insert size: Number of reads clustered: 528 Million; 3. 5 GB; 2 x 100 bp; ~30 X; 410/50 -600 bp; 458 Million Assembly features: - contig stats Phusion 2 ABy. SS Total number of contigs: Total bases of contigs: N 50 contig size: Largest contig: Averaged contig size: Contig coverage on genome: Mis-assembly errors: 7, 796, 722 2. 28 Gb 2, 013 31045 292 65% ? 1, 420, 262 3. 29 Gb 7, 618 76, 418 2, 314 ~94 % ?
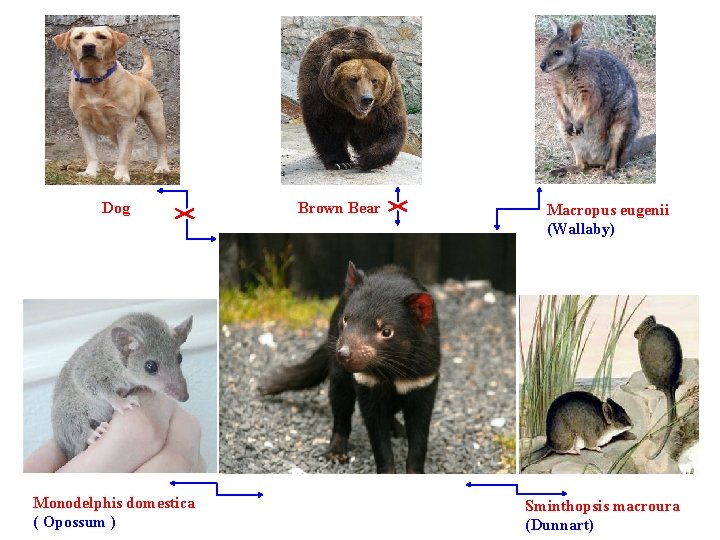
Dog Monodelphis domestica ( Opossum ) Brown Bear Macropus eugenii (Wallaby) Sminthopsis macroura (Dunnart)
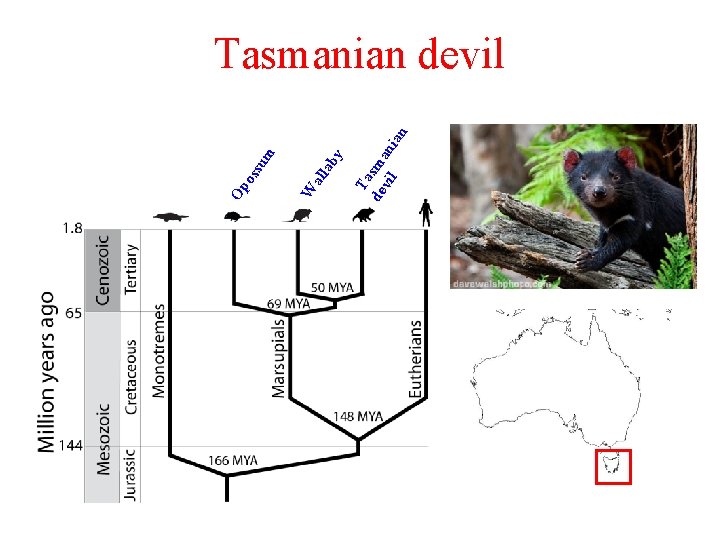
Ta de sma vil n ian ab y all W su m Op os Tasmanian devil
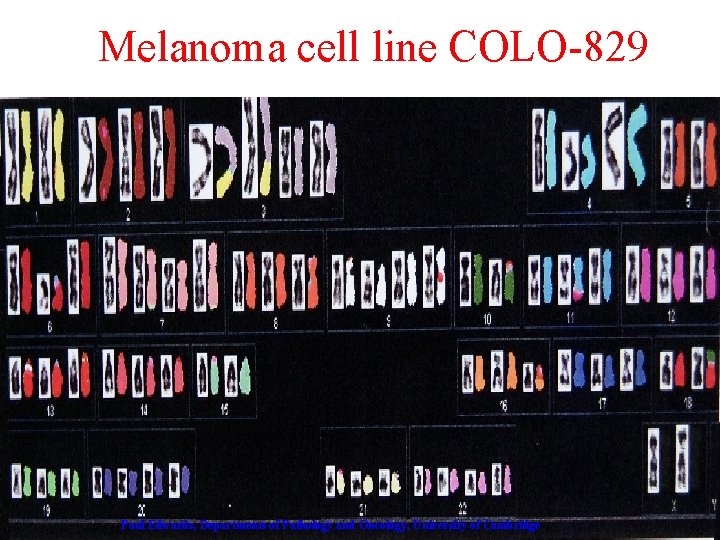
Melanoma cell line COLO-829 Paul Edwards, Departments of Pathology and Oncology, University of Cambridge

Human Cancer Genome Assembly – Normal Cell Solexa reads: Number of read pairs: Finished genome size: Read length: Estimated read coverage: Insert size: Number of reads clustered: 557 Million; 3. 0 GB; 2 x 75 bp; ~25 X; 190/50 -300 bp; 458 Million Assembly features: - contig stats Total number of contigs: Total bases of contigs: N 50 contig size: Largest contig: Averaged contig size: Contig coverage over the genome: Mis-assembly errors: 1, 020, 346; 2. 713 Gb 8, 344; 107, 613 2, 659; ~90 %; ?

Genome Assembly – Tumour Cell Solexa reads: Number of read pairs: Finished genome size: Read length: Estimated read coverage: Insert size: Number of reads clustered: 562 Million; 3. 0 GB; 2 x 75 bp; ~25 X; 190/50 -300 bp; 449 Million Assembly features: - contig stats Total number of contigs: Total bases of contigs: N 50 contig size: Largest contig: Averaged contig size: Contig coverage over the genome: Mis-assembly errors: 1, 249, 719; 2. 690 Gb 6, 073; 72, 123 2, 152; ~90 %; ?

Rice Genome Assembly One Of the most difficult Genomes on earth? Solexa reads: Number of read pairs: Finished genome size: Read length: Estimated read coverage: Insert size: Number of reads clustered: 97. 9 Million; 440 MB; 2 x 76 bp; ~33 X; 500/50 -600 bp; 81. 2 Million Assembly features: - contig stats Total number of contigs: Total bases of contigs: N 50 contig size: Largest contig: Averaged contig size: Contig coverage over the genome: Mis-assembly errors: 374, 713; 365 Mb 7, 639; 72, 321 973; ~83 %; ?

Acknowledgements: q q q q Elizabeth Murchuson Erin Preasance Mike Stratton Dirk Evers Ole Schulz-Trieglaff Qi Feng Bin Han
- Slides: 28