Phenotypic Factor Analysis Marleen de Moor Meike Bartels

Phenotypic Factor Analysis Marleen de Moor & Meike Bartels Department of Biological Psychology, VU University Amsterdam mhm. de. moor@psy. vu. nl / m. bartels@psy. vu. nl March 3, 2010 M. de Moor, Twin Workshop Boulder 1

Outline • Introduction to factor analysis – What is factor analysis – Relationship with regression and SEM – Types of factor analysis • Phenotypic factor analysis – 1 factor model – 2 factor model • More advanced models – Factor models for categorical data – Multigroup factor models and measurement invariance • From phenotypic to genetic factor analysis… March 3, 2010 M. de Moor, Twin Workshop Boulder 2

Outline • Introduction to factor analysis – What is factor analysis – Relationship with regression and SEM – Types of factor analysis • Phenotypic factor analysis • More advanced models • From phenotypic to genetic factor analysis… March 3, 2010 M. de Moor, Twin Workshop Boulder 3
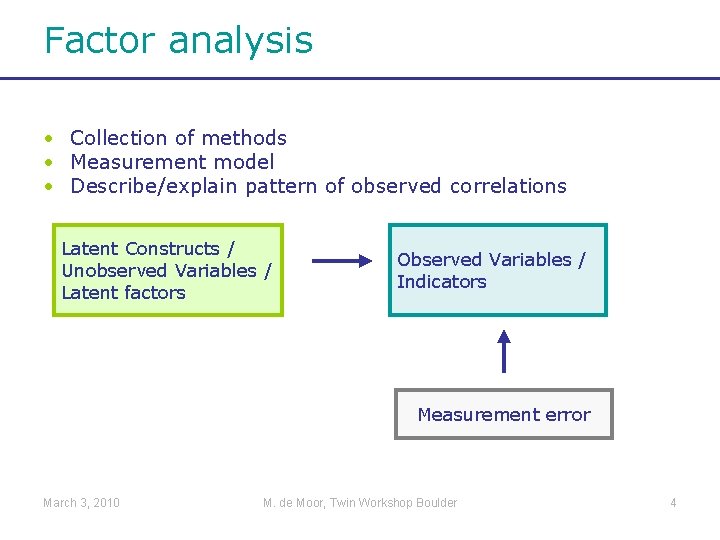
Factor analysis • Collection of methods • Measurement model • Describe/explain pattern of observed correlations Latent Constructs / Unobserved Variables / Latent factors Observed Variables / Indicators Measurement error March 3, 2010 M. de Moor, Twin Workshop Boulder 4
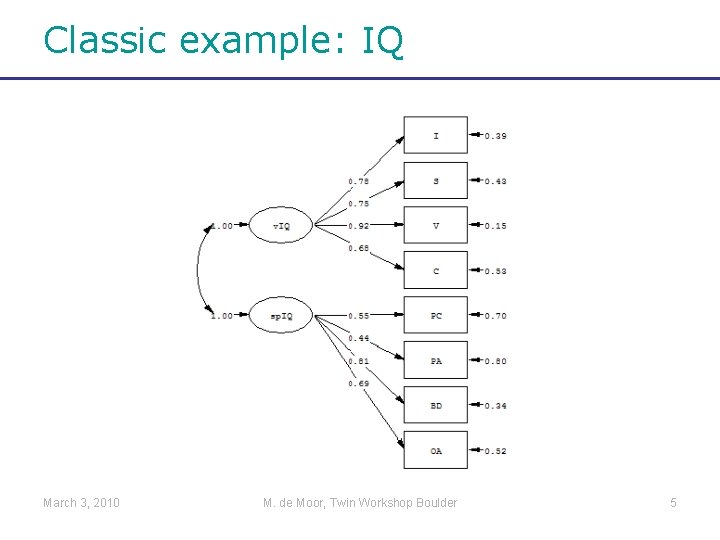
Classic example: IQ March 3, 2010 M. de Moor, Twin Workshop Boulder 5
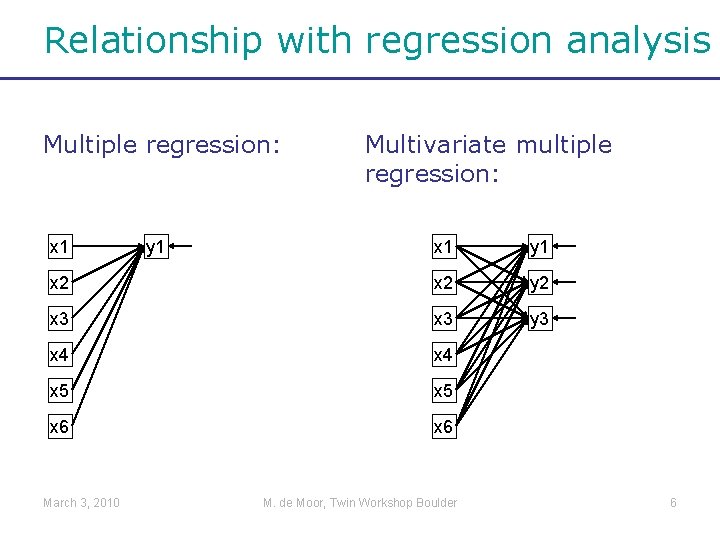
Relationship with regression analysis Multiple regression: x 1 y 1 x 2 y 2 x 3 y 3 x 4 x 5 x 6 March 3, 2010 y 1 Multivariate multiple regression: M. de Moor, Twin Workshop Boulder 6
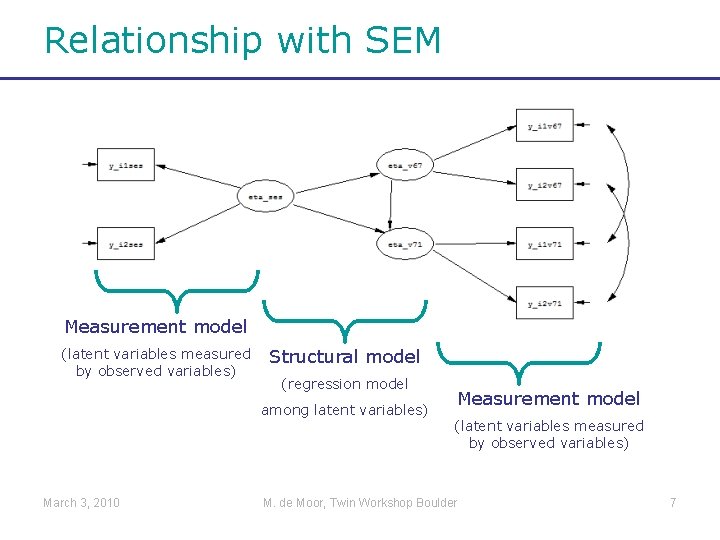
Relationship with SEM Measurement model (latent variables measured by observed variables) Structural model (regression model among latent variables) March 3, 2010 Measurement model (latent variables measured by observed variables) M. de Moor, Twin Workshop Boulder 7

Types of factor analysis • Principal component analysis (PCA) • Exploratory factor analysis (EFA) • Confirmatory factor analysis (CFA) March 3, 2010 M. de Moor, Twin Workshop Boulder 8

PCA • Data reduction technique • Linear transformation of the data • Summarize the observed pattern of correlations among variables with a smaller number of principal components March 3, 2010 M. de Moor, Twin Workshop Boulder 9
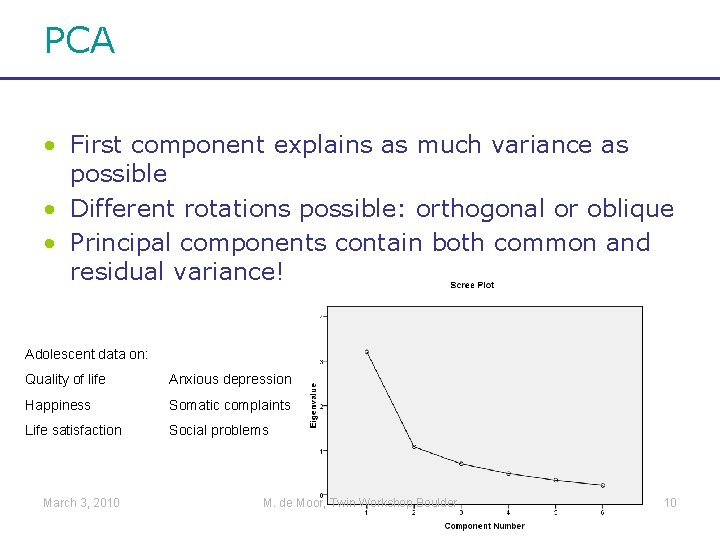
PCA • First component explains as much variance as possible • Different rotations possible: orthogonal or oblique • Principal components contain both common and residual variance! Adolescent data on: Quality of life Anxious depression Happiness Somatic complaints Life satisfaction Social problems March 3, 2010 M. de Moor, Twin Workshop Boulder 10
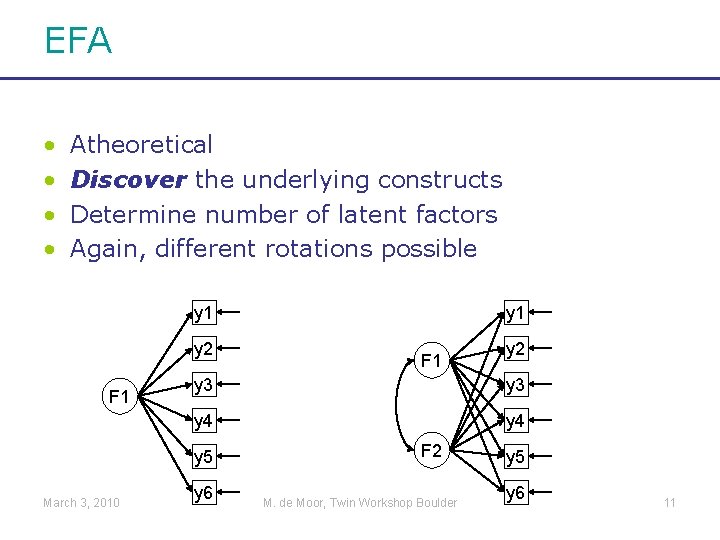
EFA • • Atheoretical Discover the underlying constructs Determine number of latent factors Again, different rotations possible y 1 y 2 F 1 y 2 y 3 y 4 y 5 March 3, 2010 y 1 y 6 F 2 M. de Moor, Twin Workshop Boulder y 5 y 6 11
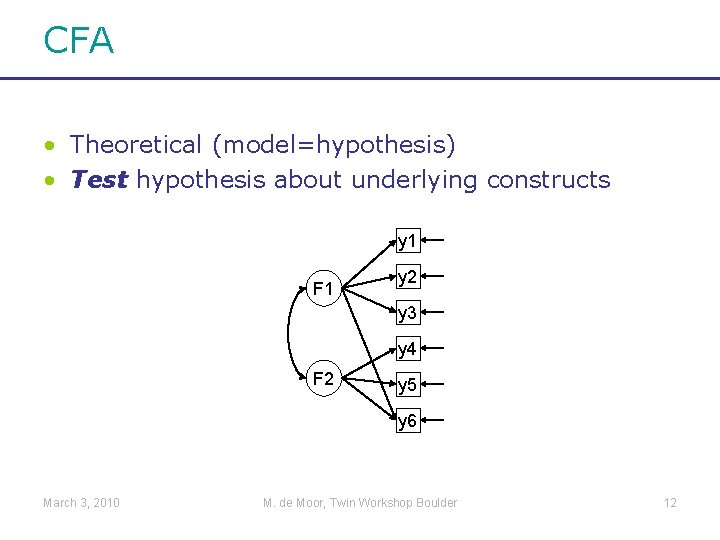
CFA • Theoretical (model=hypothesis) • Test hypothesis about underlying constructs y 1 F 1 y 2 y 3 y 4 F 2 y 5 y 6 March 3, 2010 M. de Moor, Twin Workshop Boulder 12

Outline • Introduction to factor analysis • Phenotypic factor analysis – 1 factor model – 2 factor model • More advanced models • From phenotypic to genetic factor analysis… March 3, 2010 M. de Moor, Twin Workshop Boulder 13
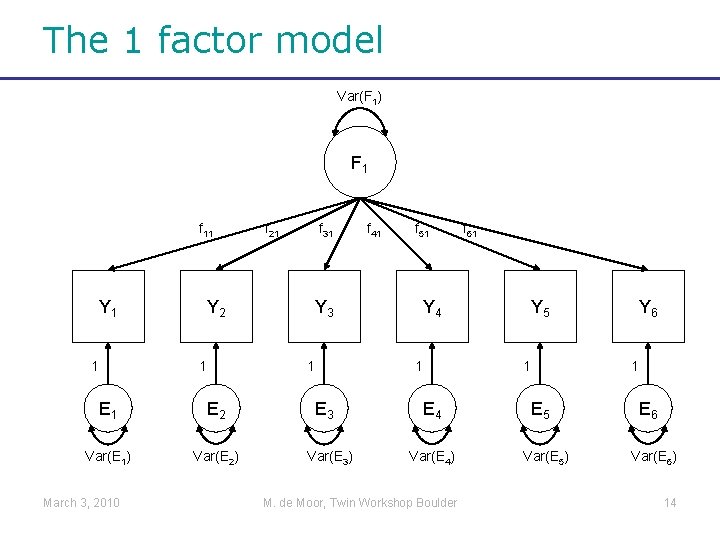
The 1 factor model Var(F 1) F 1 f 11 Y 1 1 Y 2 1 E 2 Var(E 1) Var(E 2) March 3, 2010 f 21 f 31 Y 3 1 E 3 Var(E 3) f 41 f 51 Y 4 1 E 4 Var(E 4) M. de Moor, Twin Workshop Boulder f 61 Y 5 1 E 5 Var(E 5) Y 6 1 E 6 Var(E 6) 14
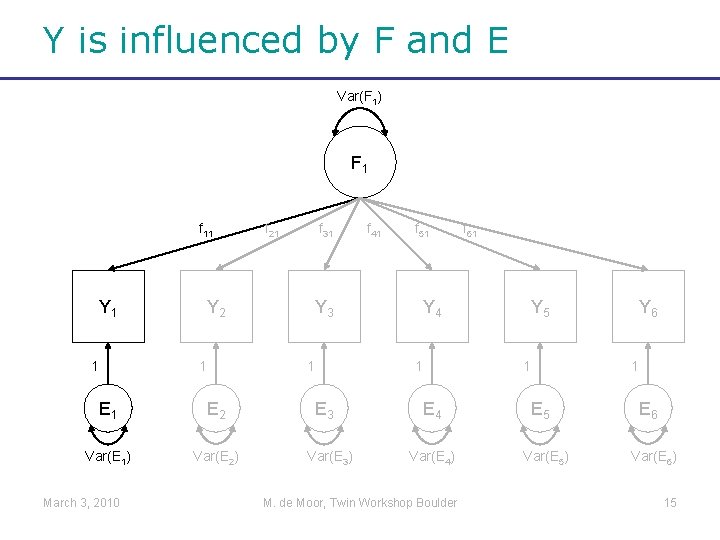
Y is influenced by F and E Var(F 1) F 1 f 11 Y 1 1 Y 2 1 E 2 Var(E 1) Var(E 2) March 3, 2010 f 21 f 31 Y 3 1 E 3 Var(E 3) f 41 f 51 Y 4 1 E 4 Var(E 4) M. de Moor, Twin Workshop Boulder f 61 Y 5 1 E 5 Var(E 5) Y 6 1 E 6 Var(E 6) 15

More formally… • Yi 1=f 11*Fi 1+Ei 1 Random variables (varies across individuals i=1…N) March 3, 2010 M. de Moor, Twin Workshop Boulder 16

More formally… • Yi 1=f 11*Fi 1+Ei 1 Random variables (varies across individuals i=1…N) Fixed parameter (constant across individuals) March 3, 2010 M. de Moor, Twin Workshop Boulder 17

More formally… • Yi 1=f 11*Fi 1+Ei 1 • Yi 2=f 21*Fi 1+Ei 2 • Yi 3=f 31*Fi 1+Ei 3 … • Yi 6=f 61*Fi 1+Ei 6 March 3, 2010 M. de Moor, Twin Workshop Boulder 18
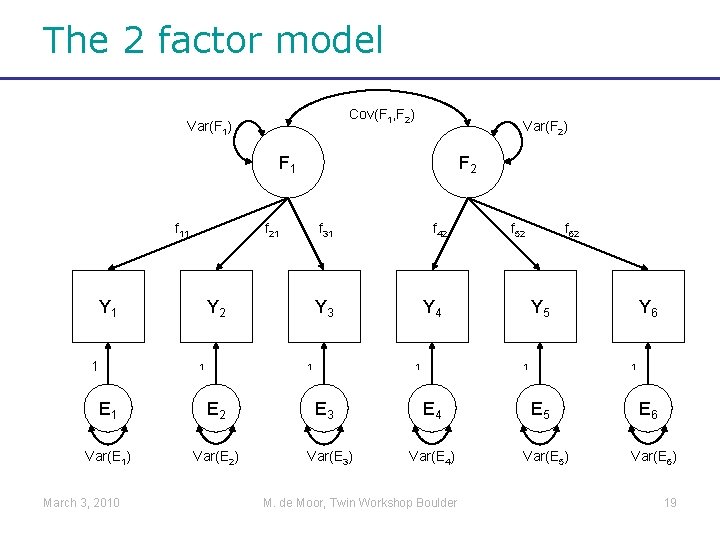
The 2 factor model Cov(F 1, F 2) Var(F 1) Var(F 2) F 1 f 11 f 21 Y 1 1 f 31 Y 2 1 E 2 Var(E 1) Var(E 2) f 42 Y 3 1 E 1 March 3, 2010 F 2 Y 4 1 E 3 Var(E 3) f 52 f 62 Y 5 1 E 4 Var(E 4) M. de Moor, Twin Workshop Boulder Y 6 1 E 5 Var(E 5) E 6 Var(E 6) 19
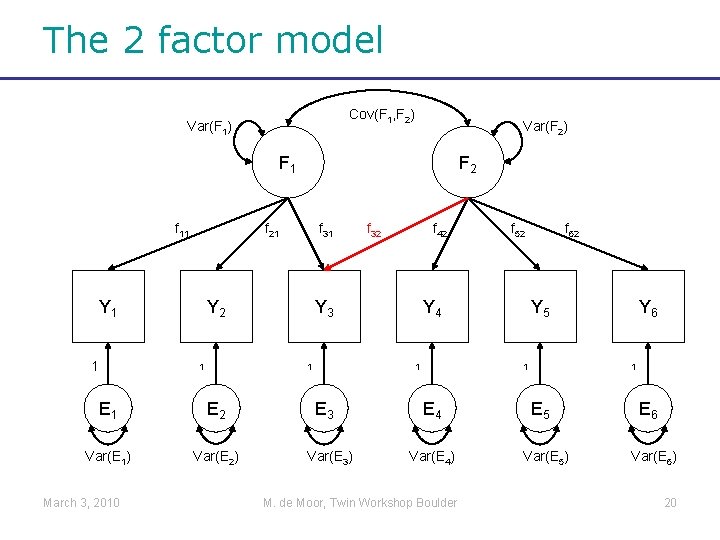
The 2 factor model Cov(F 1, F 2) Var(F 1) Var(F 2) F 1 f 11 f 21 Y 1 1 f 31 Y 2 1 E 2 Var(E 1) Var(E 2) f 32 f 42 Y 3 1 E 1 March 3, 2010 F 2 Y 4 1 E 3 Var(E 3) f 52 f 62 Y 5 1 E 4 Var(E 4) M. de Moor, Twin Workshop Boulder Y 6 1 E 5 Var(E 5) E 6 Var(E 6) 20
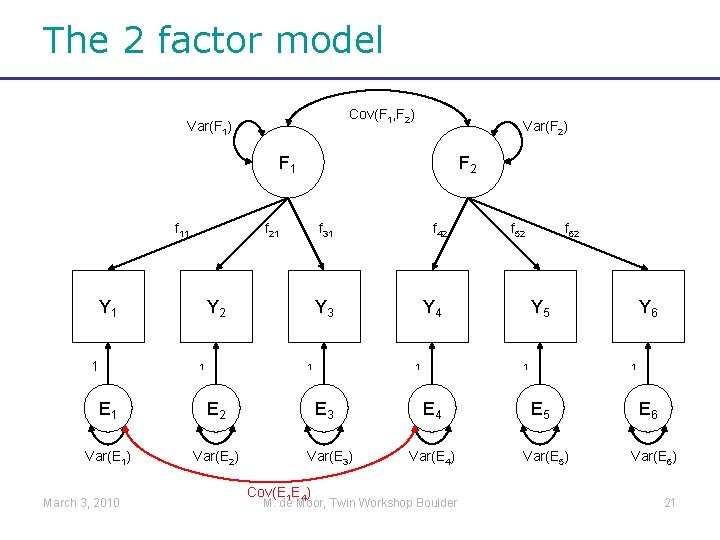
The 2 factor model Cov(F 1, F 2) Var(F 1) Var(F 2) F 1 f 11 f 21 Y 1 1 f 31 Y 2 1 E 2 Var(E 1) Var(E 2) f 42 Y 3 1 E 1 March 3, 2010 F 2 Var(E 3) Cov(E 1 E 4) Y 4 1 E 3 f 52 f 62 Y 5 1 E 4 Var(E 4) M. de Moor, Twin Workshop Boulder Y 6 1 E 5 Var(E 5) E 6 Var(E 6) 21
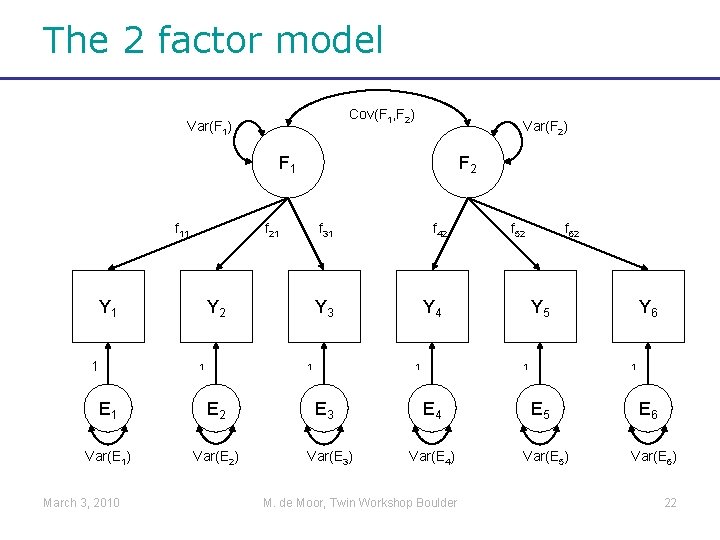
The 2 factor model Cov(F 1, F 2) Var(F 1) Var(F 2) F 1 f 11 f 21 Y 1 1 f 31 Y 2 1 E 2 Var(E 1) Var(E 2) f 42 Y 3 1 E 1 March 3, 2010 F 2 Y 4 1 E 3 Var(E 3) f 52 f 62 Y 5 1 E 4 Var(E 4) M. de Moor, Twin Workshop Boulder Y 6 1 E 5 Var(E 5) E 6 Var(E 6) 22

From equations to matrices • Yi 1=f 11*Fi 1+Ei 1 • Yi 2=f 21*Fi 1+Ei 2 • Yi 3=f 31*Fi 1+Ei 3 • Yi 4=f 42*Fi 2+Ei 4 • Yi 5=f 52*Fi 2+Ei 5 • Yi 6=f 62*Fi 2+Ei 6 i=1…N number of individuals j=1…J number of observed variables k=1…K number of factors Assumption: Data follow a multivariate normal distribution March 3, 2010 M. de Moor, Twin Workshop Boulder 23

Expected (co)variances Can be obtained in 2 ways: • Using path diagram (Wright’s rules) • Using equations (algebraic derivation) March 3, 2010 M. de Moor, Twin Workshop Boulder 24
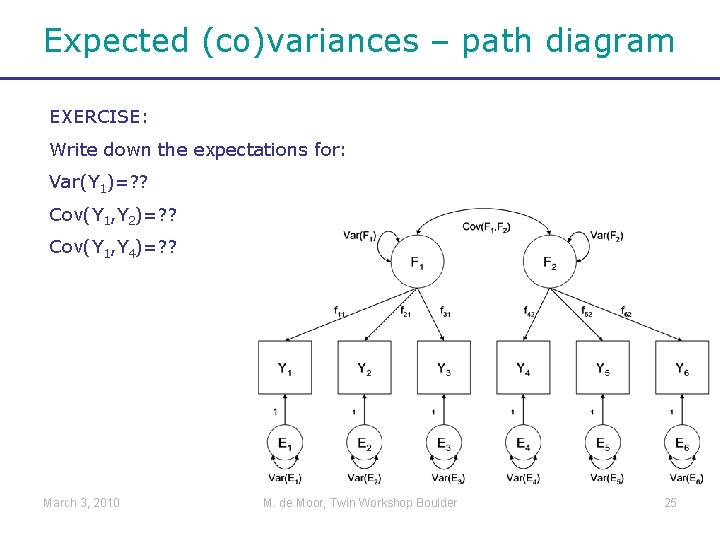
Expected (co)variances – path diagram EXERCISE: Write down the expectations for: Var(Y 1)=? ? Cov(Y 1, Y 2)=? ? Cov(Y 1, Y 4)=? ? March 3, 2010 M. de Moor, Twin Workshop Boulder 25
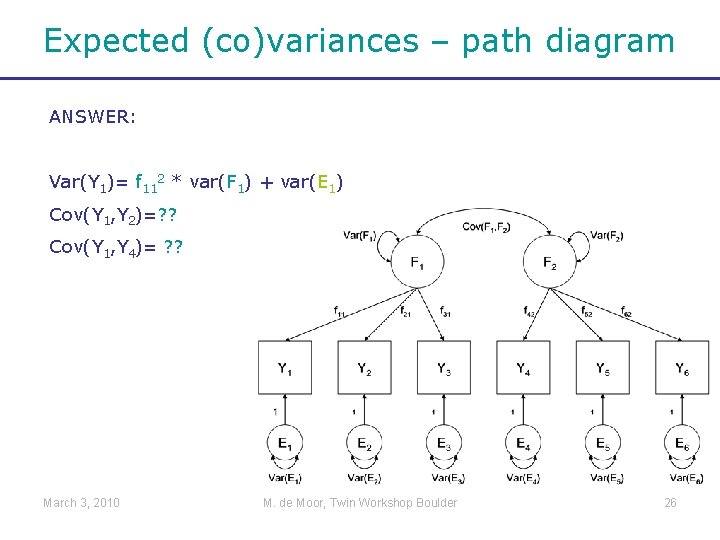
Expected (co)variances – path diagram ANSWER: Var(Y 1)= f 112 * var(F 1) + var(E 1) Cov(Y 1, Y 2)=? ? Cov(Y 1, Y 4)= ? ? March 3, 2010 M. de Moor, Twin Workshop Boulder 26
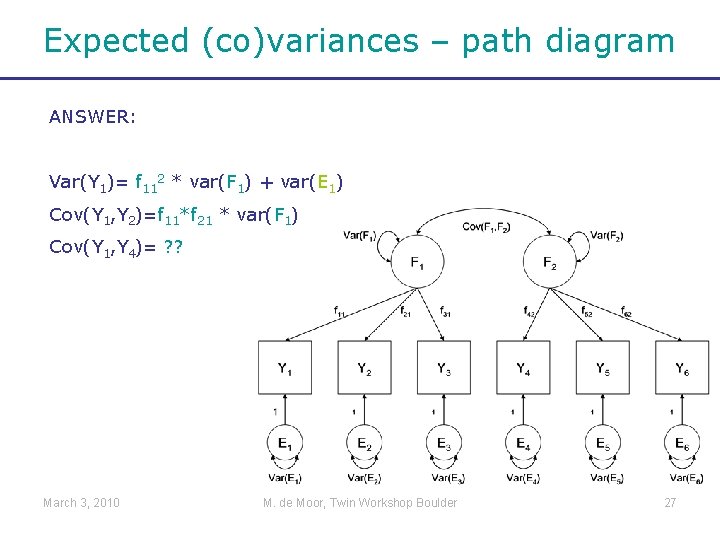
Expected (co)variances – path diagram ANSWER: Var(Y 1)= f 112 * var(F 1) + var(E 1) Cov(Y 1, Y 2)=f 11*f 21 * var(F 1) Cov(Y 1, Y 4)= ? ? March 3, 2010 M. de Moor, Twin Workshop Boulder 27
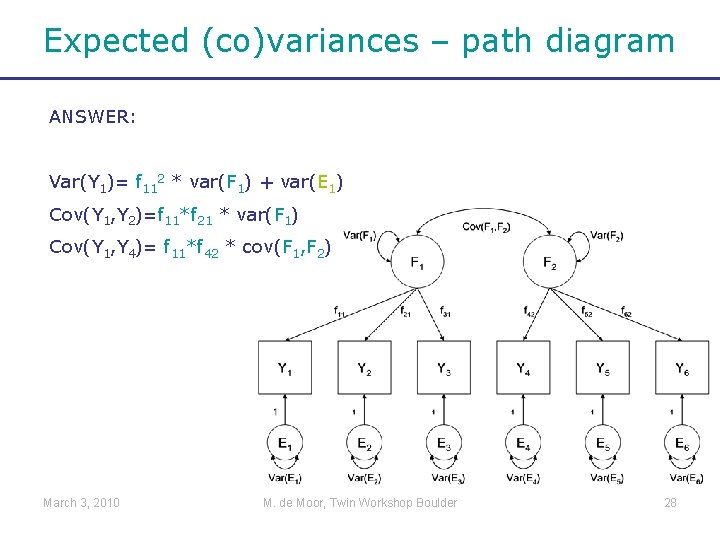
Expected (co)variances – path diagram ANSWER: Var(Y 1)= f 112 * var(F 1) + var(E 1) Cov(Y 1, Y 2)=f 11*f 21 * var(F 1) Cov(Y 1, Y 4)= f 11*f 42 * cov(F 1, F 2) March 3, 2010 M. de Moor, Twin Workshop Boulder 28

Expected (co)variances - equations Var (Y 1) = E [ (f 11*Fi 1+Ei 1) * (f 11*Fi 1+Ei 1) ] = E [ (f 11*Fi 1)2 + 2*f 11*Fi 1*Ei 1 + (Ei 1)2 ] = f 112 * var(F 1) + var(E 1) Cov (Y 1, Y 2) = E [ (f 11*Fi 1+Ei 1) * (f 21*Fi 1+Ei 2) ] = E [ f 11*Fi 1*f 21*Fi 1 + f 11*Fi 1 * Ei 2 + Ei 1*f 21*Fi 1 + Ei 1*Ei 2 ] = f 11*f 21 * var(F 1) Cov (Y 1, Y 4) = f 11*f 42 * cov(F 1, F 2) March 3, 2010 M. de Moor, Twin Workshop Boulder 29

Expected (co)variances - equations j=1…J number of observed variables k=1…K number of factors Jx. J symmetric Jx. K full Kx. K symm (LISREL notation) March 3, 2010 Kx. J full Jx. J diag (Open. Mx notation) M. de Moor, Twin Workshop Boulder 30

Identification • Latent factors have no scale: means, variances? For each latent factor: • Mean: fix to zero • Variance: two most commonly used options – Fix to one, estimate all factor loadings – Estimate variance, fix first factor loading to one March 3, 2010 M. de Moor, Twin Workshop Boulder 31

Identification of (co)variances March 3, 2010 M. de Moor, Twin Workshop Boulder 32

Identification of (co)variances March 3, 2010 M. de Moor, Twin Workshop Boulder 33

Identification • Count number of observed statistics • Count number of free parameters • If #obs. stat. < #free par. Model unidentified • If #obs. stat. = #free par. Model justidentified • If #obs. stat. > #free par. Model identified March 3, 2010 M. de Moor, Twin Workshop Boulder 34

Identification of 1 factor model Observed statistics: #obs. var. = J = 6 #obs. cov. = J(J-1)/2 = 6*5/2 = 15 #obs. var/cov. = J(J+1)/2 = 6*7/2 = 21 Free parameters: # residual variances = 6 # factor loadings = 6 Degrees of freedom: df = 21 -12 = 9 March 3, 2010 M. de Moor, Twin Workshop Boulder 35

Identification of 2 factor model Observed statistics: #obs. var. = J = 6 #obs. cov. = J(J-1)/2 = 6*5/2 = 15 #obs. var/cov. = J(J+1)/2 = 6*7/2 = 21 Free parameters: # residual variances = 6 # factor loadings = 6 # covariances among factors = 1 Degrees of freedom: df = 21 -13 = 8 March 3, 2010 M. de Moor, Twin Workshop Boulder 36

Practical – Description of data March 3, 2010 M. de Moor, Twin Workshop Boulder 37

DATASET - Netherlands Twin Register (www. tweelingenregister. org) - Dutch Health and Behavior Questionnaire (DHBQ) - Adolescent Twins and non-twin Siblings - Aged 14 and 16 (siblings between 12 and 25) - Online & Paper and Pencil March 3, 2010 M. de Moor, Twin Workshop Boulder 38

CONTENT DHBQ - Psychopathology - Leisure time activities - Exercise - Family Size (no of sibs) - Self –esteem - Family situation (divorce) - Optimism - Zygosity - Life Events - Height, Weight - Loneliness - Eating Disorders - Number of peers and peer relation - General Health and Illnesses (astma, migraine, etc) - Pubertal Development - Hours sleep - Personality (age 16) - Family Functioning (Family Functioning, Family Conflict) - Life style (smoking, alcohol use, marihuna use) - Educational Achievement (incl truancy) - Wellbeing (Happiness, Satisfaction with Life, Quality of Life) March 3, 2010 M. de Moor, Twin Workshop Boulder 39
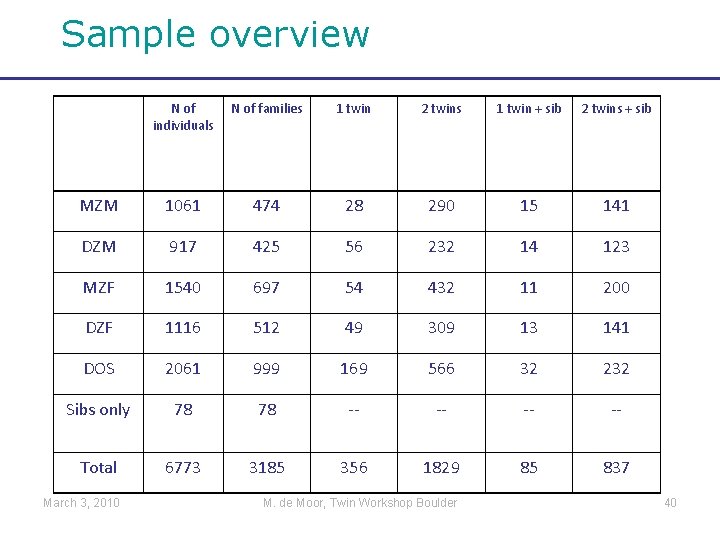
Sample overview N of individuals N of families 1 twin 2 twins 1 twin + sib 2 twins + sib MZM 1061 474 28 290 15 141 DZM 917 425 56 232 14 123 MZF 1540 697 54 432 11 200 DZF 1116 512 49 309 13 141 DOS 2061 999 169 566 32 232 Sibs only 78 78 -- -- Total 6773 3185 356 1829 85 837 March 3, 2010 M. de Moor, Twin Workshop Boulder 40

Today’s Focus - Youth Self Report (YSR) - Subjective Wellbeing * subjective happiness * satisfaction with life * quality of life - General Family Functioning March 3, 2010 M. de Moor, Twin Workshop Boulder 41

Practical 1: Single group factor models Data: CFA_family_wellbeing. dat • 1000 adolescent twins (one twin per family) • Observed variables: – – – Quality of life Happiness Satisfaction with life Anxious depression scale (YSR) Somatic complaints scale (YSR) Social problems scale (YSR) Files are on F: marleenBoulder 2010CFA March 3, 2010 M. de Moor, Twin Workshop Boulder 42
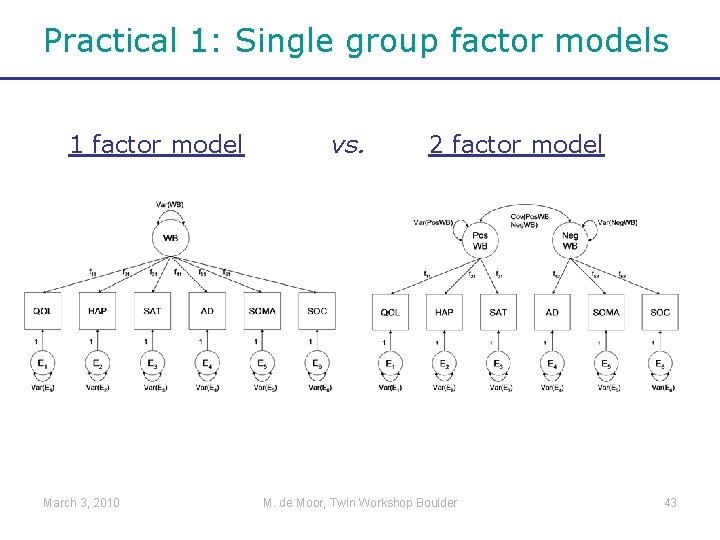
Practical 1: Single group factor models 1 factor model March 3, 2010 vs. 2 factor model M. de Moor, Twin Workshop Boulder 43

Practical 1 a: Single group 1 factor model Open. Mx script One. Factor. Model. Matrix_WELLBEING. R require(Open. Mx) # Prepare Data # -----------------------------------all. Data<-read. table("CFA_family_wellbeing. dat", header=TRUE, na. strings=-999) Read in data, -999 are treated as missing NA cfa. Data<-all. Data[, c('qol', 'hap', 'sat', 'ad', 'soma', 'soc')] Select variables to use in CFA col. Means(cfa. Data, na. rm=TRUE) cov(cfa. Data[, c('qol', 'hap', 'sat', 'ad', 'soma', 'soc')], use="pairwise. complete. obs") cor(cfa. Data[, c('qol', 'hap', 'sat', 'ad', 'soma', 'soc')], use="pairwise. complete. obs") Compute descriptives of variables nvar<-6 nfac<-1 Specify number of variables and factors March 3, 2010 M. de Moor, Twin Workshop Boulder 44
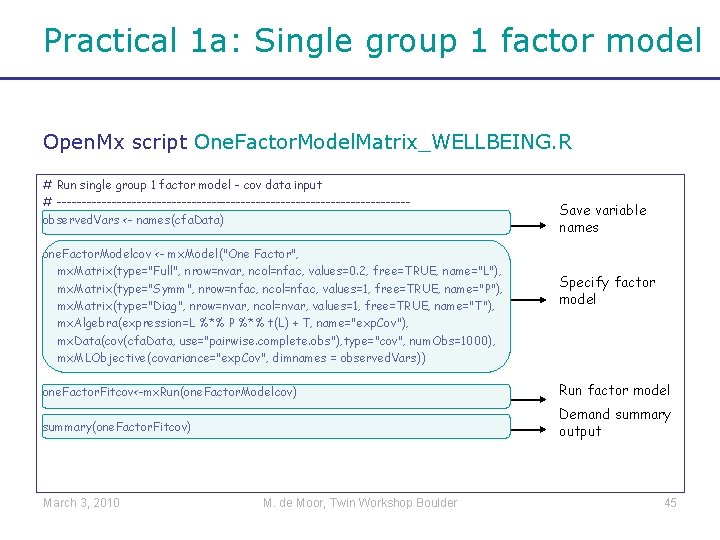
Practical 1 a: Single group 1 factor model Open. Mx script One. Factor. Model. Matrix_WELLBEING. R # Run single group 1 factor model - cov data input # -----------------------------------observed. Vars <- names(cfa. Data) one. Factor. Modelcov <- mx. Model("One Factor", mx. Matrix(type="Full", nrow=nvar, ncol=nfac, values=0. 2, free=TRUE, name="L"), mx. Matrix(type="Symm", nrow=nfac, ncol=nfac, values=1, free=TRUE, name="P"), mx. Matrix(type="Diag", nrow=nvar, ncol=nvar, values=1, free=TRUE, name="T"), mx. Algebra(expression=L %*% P %*% t(L) + T, name="exp. Cov"), mx. Data(cov(cfa. Data, use="pairwise. complete. obs"), type="cov", num. Obs=1000), mx. MLObjective(covariance="exp. Cov", dimnames = observed. Vars)) Save variable names Specify factor model one. Factor. Fitcov<-mx. Run(one. Factor. Modelcov) Run factor model summary(one. Factor. Fitcov) Demand summary output March 3, 2010 M. de Moor, Twin Workshop Boulder 45

Practical 1 a: Single group 1 factor model • Copy files from F: marleenBoulder 2010CFA to own directory • Check whether your own directory is your working directory! 1. 2. 3. 4. Open script One. Factor. Model. Matrix_WELLBEING. R Identify the factor model by constraining Var(WB)=1 Run the 1 factor model Write down the following information: Model: #obs. stat. #free par. Chi 2 df AIC BIC RMSEA 1 factor March 3, 2010 M. de Moor, Twin Workshop Boulder 46

Practical 1 a: Single group 1 factor model one. Factor. Modelcov <- mx. Model("One Factor", mx. Matrix(type="Full", nrow=nvar, ncol=nfac, values=0. 2, free=TRUE, name="L"), mx. Matrix(type="Symm", nrow=nfac, ncol=nfac, values=1, free=FALSE, name="P"), mx. Matrix(type="Diag", nrow=nvar, ncol=nvar, values=1, free=TRUE, name="T"), mx. Algebra(expression=L %*% P %*% t(L) + T, name="exp. Cov"), mx. Data(cov(cfa. Data, use="pairwise. complete. obs"), type="cov", num. Obs=1000), mx. MLObjective(covariance="exp. Cov", dimnames = observed. Vars)) Model: #obs. stat. #free par. Chi 2 df AIC BIC RMSEA 1 factor 21 12 508. 6 9 490. 6 223. 2 0. 24 March 3, 2010 M. de Moor, Twin Workshop Boulder 47

Practical 1 b: Single group 2 factor model Open. Mx script Two. Factor. Model. Matrix_WELLBEING. R require(Open. Mx) # Prepare Data # -----------------------------------all. Data<-read. table("CFA_family_wellbeing. dat", header=TRUE, na. strings=-999) Read in data, -999 are treated as missing NA cfa. Data<-all. Data[, c('qol', 'hap', 'sat', 'ad', 'soma', 'soc')] Select variables to use in CFA col. Means(cfa. Data, na. rm=TRUE) cov(cfa. Data[, c('qol', 'hap', 'sat', 'ad', 'soma', 'soc')], use="pairwise. complete. obs") cor(cfa. Data[, c('qol', 'hap', 'sat', 'ad', 'soma', 'soc')], use="pairwise. complete. obs") Compute descriptives of variables nvar<-6 nfac<-2 Specify number of variables and factors March 3, 2010 M. de Moor, Twin Workshop Boulder 48
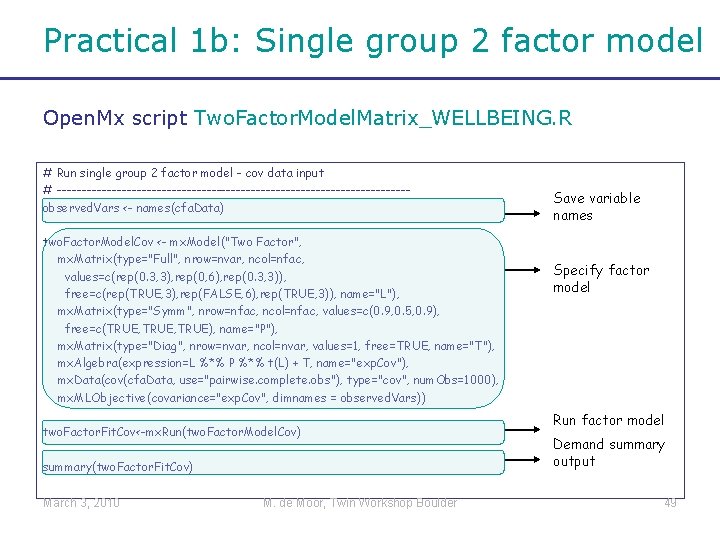
Practical 1 b: Single group 2 factor model Open. Mx script Two. Factor. Model. Matrix_WELLBEING. R # Run single group 2 factor model - cov data input # -----------------------------------observed. Vars <- names(cfa. Data) two. Factor. Model. Cov <- mx. Model("Two Factor", mx. Matrix(type="Full", nrow=nvar, ncol=nfac, values=c(rep(0. 3, 3), rep(0, 6), rep(0. 3, 3)), free=c(rep(TRUE, 3), rep(FALSE, 6), rep(TRUE, 3)), name="L"), mx. Matrix(type="Symm", nrow=nfac, ncol=nfac, values=c(0. 9, 0. 5, 0. 9), free=c(TRUE, TRUE), name="P"), mx. Matrix(type="Diag", nrow=nvar, ncol=nvar, values=1, free=TRUE, name="T"), mx. Algebra(expression=L %*% P %*% t(L) + T, name="exp. Cov"), mx. Data(cov(cfa. Data, use="pairwise. complete. obs"), type="cov", num. Obs=1000), mx. MLObjective(covariance="exp. Cov", dimnames = observed. Vars)) two. Factor. Fit. Cov<-mx. Run(two. Factor. Model. Cov) summary(two. Factor. Fit. Cov) March 3, 2010 M. de Moor, Twin Workshop Boulder Save variable names Specify factor model Run factor model Demand summary output 49

Factor loading matrix “L” mx. Matrix(type="Full", nrow=nvar, ncol=nfac, values=c(rep(0. 3, 3), rep(0, 6), rep(0. 3, 3)), free=c(rep(TRUE, 3), rep(FALSE, 6), rep(TRUE, 3)), name="L"), Factor loading matrix L: March 3, 2010 Starting values for elements in this matrix: M. de Moor, Twin Workshop Boulder Free parameters in this matrix: 50

Covariance matrix latent factors “P” mx. Matrix("Symm", nfac, values=c(0. 9, 0. 5, 0. 9), free=c(TRUE, TRUE), name="P"), Covariance matrix P: Starting values for elements in this matrix: Free parameters in this matrix: March 3, 2010 M. de Moor, Twin Workshop Boulder 51

Practical 1 b: Single group 2 factor model 1. Open script Two. Factor. Model. Matrix_WELLBEING. R 2. Identify the factor model by constraining Var(Pos. WB)=1 and Var(Neg. WB)=1 3. Run the 2 factor model 4. Write down the following information: Model: #obs. stat. #free par. Chi 2 df AIC BIC RMSEA 1 factor 21 12 508. 6 9 490. 6 223. 2 0. 24 2 factor 5. How do the models fit? Which model fits best? M. de Moor, Twin Workshop Boulder Files are on F: marleenBoulder 2010CFA March 3, 2010 52

Practical 1 b: Single group 2 factor model two. Factor. Model. Cov <- mx. Model("Two Factor", mx. Matrix(type="Full", nrow=nvar, ncol=nfac, values=c(rep(0. 3, 3), rep(0, 6), rep(0. 3, 3)), free=c(rep(TRUE, 3), rep(FALSE, 6), rep(TRUE, 3)), name="L"), mx. Matrix(type="Symm", nrow=nfac, ncol=nfac, values=c(1, 0. 5, 1), free=c(FALSE, TRUE, FALSE), name="P"), mx. Matrix(type="Diag", nrow=nvar, ncol=nvar, values=1, free=TRUE, name="T"), mx. Algebra(expression=L %*% P %*% t(L) + T, name="exp. Cov"), mx. Data(cov(cfa. Data, use="pairwise. complete. obs"), type="cov", num. Obs=1000), mx. MLObjective(covariance="exp. Cov", dimnames = observed. Vars)) Model: #obs. stat. #free par. Chi 2 df AIC BIC RMSEA 1 factor 21 12 508. 6 9 490. 6 223. 2 0. 24 2 factor 21 13 69. 9 8 53. 9 7. 3 0. 09 March 3, 2010 M. de Moor, Twin Workshop Boulder 53

Outline • Introduction to factor analysis • Phenotypic factor analysis • More advanced models – Factor models for categorical data – Multigroup factor models and measurement invariance • From phenotypic to genetic factor analysis… March 3, 2010 M. de Moor, Twin Workshop Boulder 54

Factor models for categorical data • What if my observed data are categorical? • For example, multiple items of one scale Threshold models = Latent response variable models March 3, 2010 M. de Moor, Twin Workshop Boulder 55
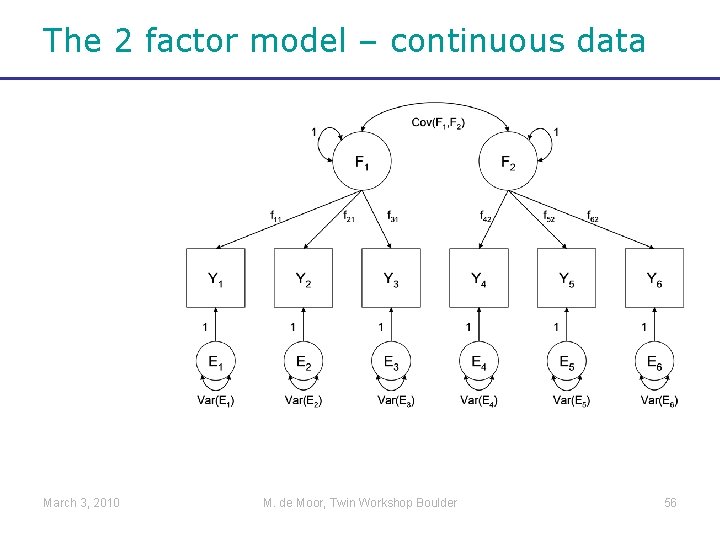
The 2 factor model – continuous data March 3, 2010 M. de Moor, Twin Workshop Boulder 56
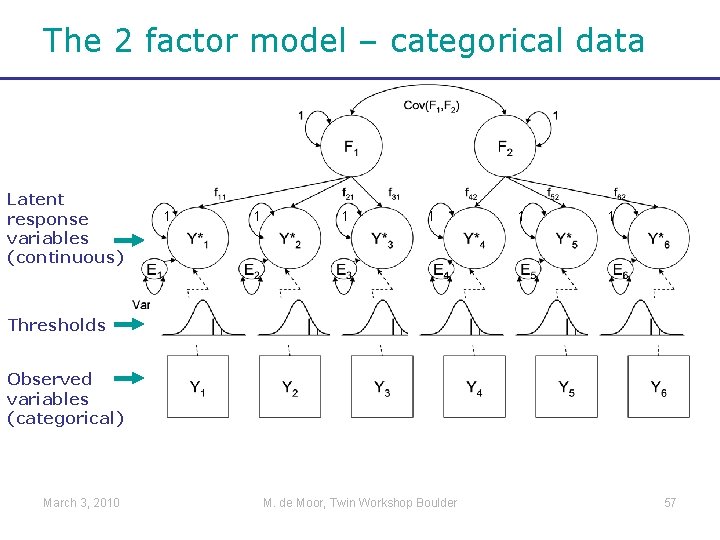
The 2 factor model – categorical data Latent response variables (continuous) 1 1 1 Thresholds Observed variables (categorical) March 3, 2010 M. de Moor, Twin Workshop Boulder 57

Multigroup factor model • Fit factor model in multiple groups Group 1: Boys Group 2: Girls March 3, 2010 M. de Moor, Twin Workshop Boulder 58

Group comparisons • Group comparisons of latent constructs: – Means For example: v IQ differences across ethnic groups – Covariance structure For example: v Covariance differences in negative and positive wellbeing in adolescent boys and girls • Only meaningful if shown that same constructs are measured in all groups! March 3, 2010 M. de Moor, Twin Workshop Boulder 59

Modeling means and covariances Means model: March 3, 2010 Covariance model: M. de Moor, Twin Workshop Boulder 60

Measurement invariance (MI) = Absence of measurement bias = Same measurement model holds in each group Group differences in observed variables are only caused by group differences in latent factors, and not by other differences in the model, such as differences in factor loadings March 3, 2010 M. de Moor, Twin Workshop Boulder 61

Types of MI models Four models (models 2 -4 are nested under 1) Most complex 1. Configural Invariance model 2. Metric Invariance model 3. Strict Invariance model Most parsimonious 4. Strong Invariance model March 3, 2010 M. de Moor, Twin Workshop Boulder 62

Types of MI models Four models (models 2 -4 are nested under 1) 1. Configural Invariance model • Fit same factor model in each group 2. Metric Invariance model • Constrain factor loadings equal across groups 3. Strict Invariance model • Constrain factor loadings and intercepts equal across groups 4. Strong Invariance model • Constrain factor loadings, intercepts and residual variances equal across groups March 3, 2010 M. de Moor, Twin Workshop Boulder 63

Outline • • Introduction to factor analysis Phenotypic factor analysis More advanced models From phenotypic to genetic factor analysis… March 3, 2010 M. de Moor, Twin Workshop Boulder 64
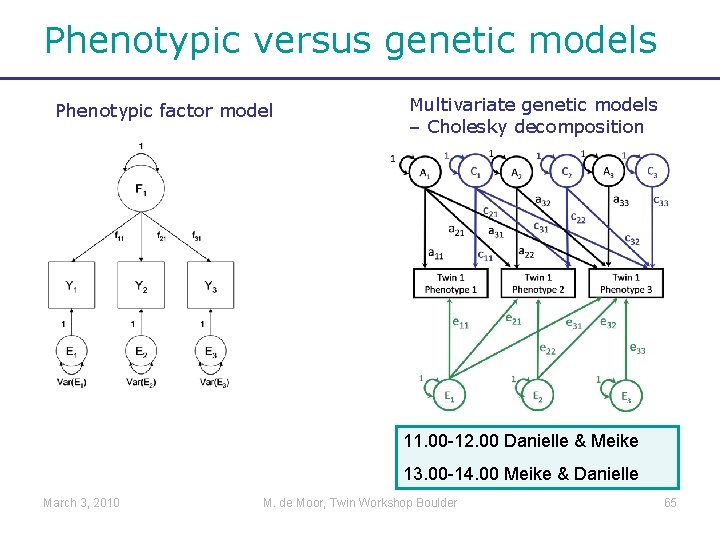
Phenotypic versus genetic models Phenotypic factor model Multivariate genetic models – Cholesky decomposition 11. 00 -12. 00 Danielle & Meike 13. 00 -14. 00 Meike & Danielle March 3, 2010 M. de Moor, Twin Workshop Boulder 65
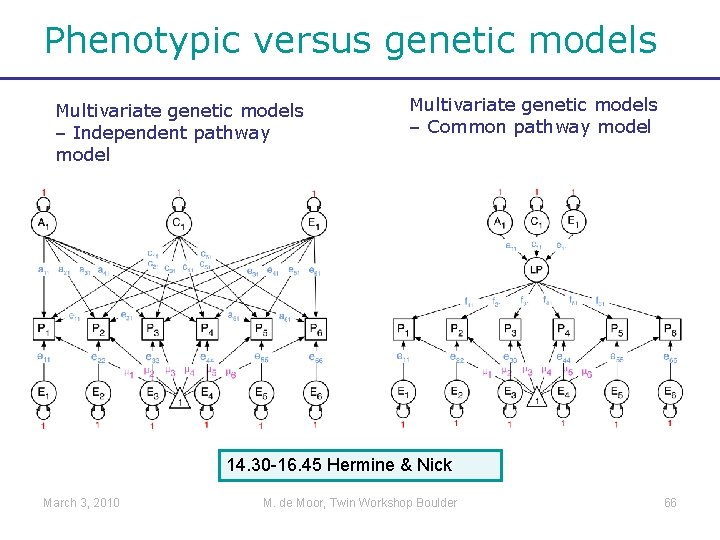
Phenotypic versus genetic models Multivariate genetic models – Independent pathway model Multivariate genetic models – Common pathway model 14. 30 -16. 45 Hermine & Nick March 3, 2010 M. de Moor, Twin Workshop Boulder 66

March 3, 2010 M. de Moor, Twin Workshop Boulder 67
- Slides: 67