Phenotypic Characterization of lrb Mutants in Arabidopsis thaliana
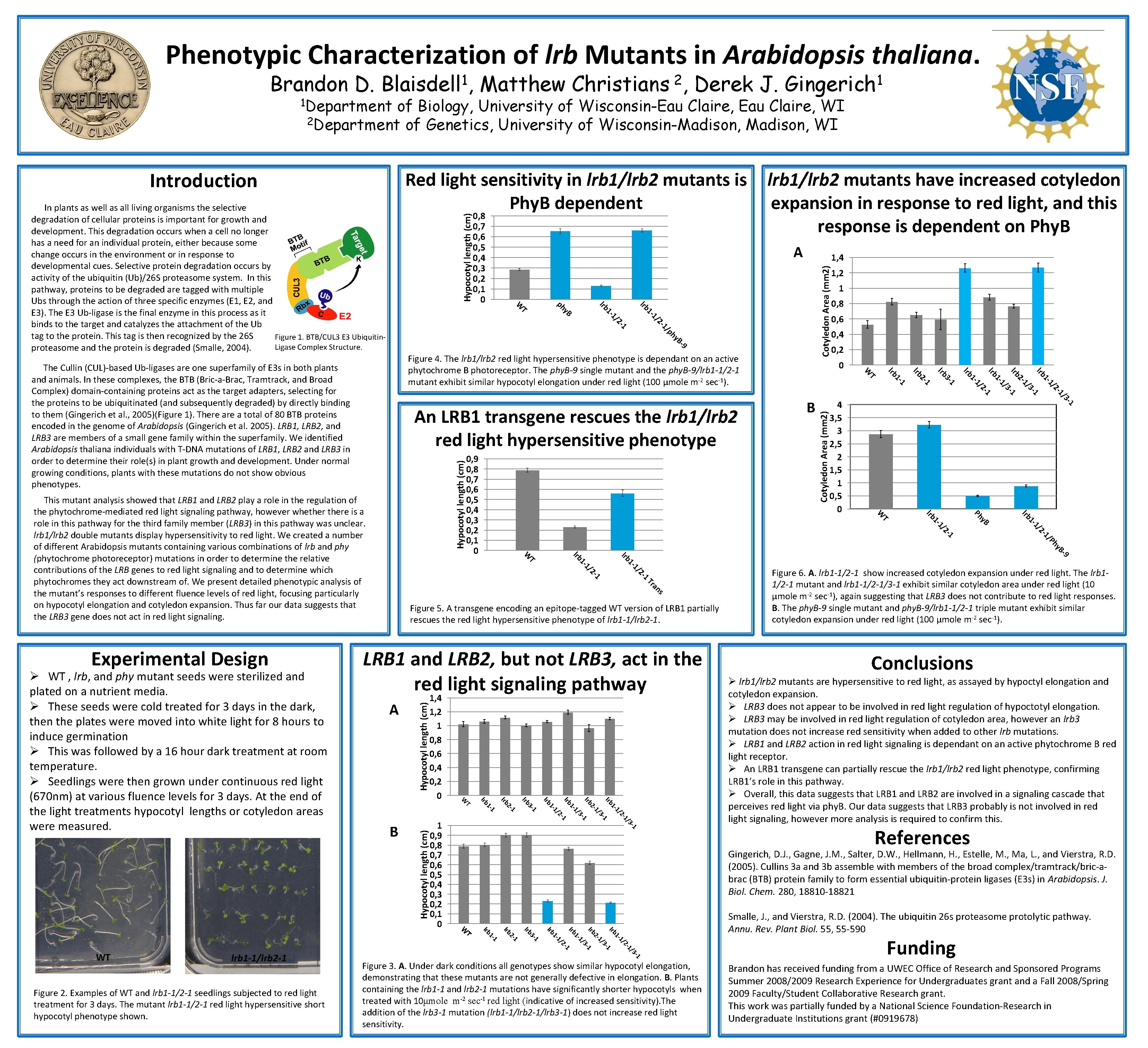
Phenotypic Characterization of lrb Mutants in Arabidopsis thaliana. Brandon D. 1 Blaisdell , Derek J. 1 Gingerich of Biology, University of Wisconsin-Eau Claire, WI 2 Department of Genetics, University of Wisconsin-Madison, WI Introduction Red light sensitivity in lrb 1/lrb 2 mutants is Phy. B dependent Hypocotyl length (cm) 1, 4 9 B- hy /p 1 2 - 1/ 1/ 1 - 1 - lrb 1, 2 1 0, 8 0, 6 0, 4 0, 2 0 1/ 1 1 3 - Cotyledon Area (mm 2) 2 - 3 - -9 y. B Ph 1/ 2 - 1/ 1 - lrb Hypocotyl length (cm) 1/ 1/ Hypocotyl length (cm) 1 - 21 y. B Hypocotyl length (cm) lrb 3 - 1 2 Ph 1 2 - 1/ 1 - lrb T W ns a Tr -1 /3 -1 /2 1 1 - lrb -1 /3 1 2 - lrb -1 /3 1 1 - lrb -1 /2 1 1 - 1 0, 9 0, 8 0, 7 0, 6 0, 5 0, 4 0, 3 0, 2 0, 1 0 lrb 1 3 - lrb 1 2 - lrb 1 1 - lrb B 4 3, 5 3 2, 5 2 1, 5 1 0, 5 0 1/ 1 - 1 1/ 1 - lrb 3 - 1 2 - 1 1 - lrb lrb T W 1 1 2 - 2 - 1/ 1/ 1 - 1 - A 1, 4 1, 2 1 0, 8 0, 6 0, 4 0, 2 0 B lrb T LRB 1 and LRB 2, but not LRB 3, act in the red light signaling pathway Figure 6. A. lrb 1 -1/2 -1 show increased cotyledon expansion under red light. The lrb 11/2 -1 mutant and lrb 1 -1/2 -1/3 -1 exhibit similar cotyledon area under red light (10 μmole m-2 sec-1), again suggesting that LRB 3 does not contribute to red light responses. B. The phy. B-9 single mutant and phy. B-9/lrb 1 -1/2 -1 triple mutant exhibit similar cotyledon expansion under red light (100 μmole m-2 sec-1). Conclusions Ø lrb 1/lrb 2 mutants are hypersensitive to red light, as assayed by hypoctyl elongation and cotyledon expansion. Ø LRB 3 does not appear to be involved in red light regulation of hypoctotyl elongation. Ø LRB 3 may be involved in red light regulation of cotyledon area, however an lrb 3 mutation does not increase red sensitivity when added to other lrb mutations. Ø LRB 1 and LRB 2 action in red light signaling is dependant on an active phytochrome B red light receptor. Ø An LRB 1 transgene can partially rescue the lrb 1/lrb 2 red light phenotype, confirming LRB 1’s role in this pathway. Ø Overall, this data suggests that LRB 1 and LRB 2 are involved in a signaling cascade that perceives red light via phy. B. Our data suggests that LRB 3 probably is not involved in red light signaling, however more analysis is required to confirm this. References Gingerich, D. J. , Gagne, J. M. , Salter, D. W. , Hellmann, H. , Estelle, M. , Ma, L. , and Vierstra, R. D. (2005). Cullins 3 a and 3 b assemble with members of the broad complex/tramtrack/bric-abrac (BTB) protein family to form essential ubiquitin-protein ligases (E 3 s) in Arabidopsis. J. Biol. Chem. 280, 18810 -18821 Smalle, J. , and Vierstra, R. D. (2004). The ubiquitin 26 s proteasome protolytic pathway. Annu. Rev. Plant Biol. 55, 55 -590 -1 /3 -1 /2 1 1 - lrb -1 /3 1 2 - lrb -1 /3 1 1 - lrb -1 /2 1 1 - lrb 1 3 - lrb 1 2 - lrb 1 1 - lrb T Figure 2. Examples of WT and lrb 1 -1/2 -1 seedlings subjected to red light treatment for 3 days. The mutant lrb 1 -1/2 -1 red light hypersensitive short hypocotyl phenotype shown. W Figure 5. A transgene encoding an epitope-tagged WT version of LRB 1 partially rescues the red light hypersensitive phenotype of lrb 1 -1/lrb 2 -1. W lrb 1 -1/lrb 2 -1 y. B 0, 9 0, 8 0, 7 0, 6 0, 5 0, 4 0, 3 0, 2 0, 1 0 T W WT ph T An LRB 1 transgene rescues the lrb 1/lrb 2 red light hypersensitive phenotype This mutant analysis showed that LRB 1 and LRB 2 play a role in the regulation of the phytochrome-mediated red light signaling pathway, however whethere is a role in this pathway for the third family member (LRB 3) in this pathway was unclear. lrb 1/lrb 2 double mutants display hypersensitivity to red light. We created a number of different Arabidopsis mutants containing various combinations of lrb and phy (phytochrome photoreceptor) mutations in order to determine the relative contributions of the LRB genes to red light signaling and to determine which phytochromes they act downstream of. We present detailed phenotypic analysis of the mutant’s responses to different fluence levels of red light, focusing particularly on hypocotyl elongation and cotyledon expansion. Thus far our data suggests that the LRB 3 gene does not act in red light signaling. Ø WT , lrb, and phy mutant seeds were sterilized and plated on a nutrient media. Ø These seeds were cold treated for 3 days in the dark, then the plates were moved into white light for 8 hours to induce germination Ø This was followed by a 16 hour dark treatment at room temperature. Ø Seedlings were then grown under continuous red light (670 nm) at various fluence levels for 3 days. At the end of the light treatments hypocotyl lengths or cotyledon areas were measured. A Figure 4. The lrb 1/lrb 2 red light hypersensitive phenotype is dependant on an active phytochrome B photoreceptor. The phy. B-9 single mutant and the phy. B-9/lrb 1 -1/2 -1 mutant exhibit similar hypocotyl elongation under red light (100 μmole m-2 sec-1). The Cullin (CUL)-based Ub-ligases are one superfamily of E 3 s in both plants and animals. In these complexes, the BTB (Bric-a-Brac, Tramtrack, and Broad Complex) domain-containing proteins act as the target adapters, selecting for the proteins to be ubiquitinated (and subsequently degraded) by directly binding to them (Gingerich et al. , 2005)(Figure 1). There a total of 80 BTB proteins encoded in the genome of Arabidopsis (Gingerich et al. 2005). LRB 1, LRB 2, and LRB 3 are members of a small gene family within the superfamily. We identified Arabidopsis thaliana individuals with T-DNA mutations of LRB 1, LRB 2 and LRB 3 in order to determine their role(s) in plant growth and development. Under normal growing conditions, plants with these mutations do not show obvious phenotypes. Experimental Design 0, 8 0, 7 0, 6 0, 5 0, 4 0, 3 0, 2 0, 1 0 W In plants as well as all living organisms the selective degradation of cellular proteins is important for growth and development. This degradation occurs when a cell no longer has a need for an individual protein, either because some change occurs in the environment or in response to developmental cues. Selective protein degradation occurs by activity of the ubiquitin (Ub)/26 S proteasome system. In this pathway, proteins to be degraded are tagged with multiple Ubs through the action of three specific enzymes (E 1, E 2, and E 3). The E 3 Ub-ligase is the final enzyme in this process as it binds to the target and catalyzes the attachment of the Ub tag to the protein. This tag is then recognized by the 26 S Figure 1. BTB/CUL 3 E 3 Ubiquitin. Ligase Complex Structure. proteasome and the protein is degraded (Smalle, 2004). lrb 1/lrb 2 mutants have increased cotyledon expansion in response to red light, and this response is dependent on Phy. B Cotyledon Area (mm 2) 1 Department Matthew Christians 2, Figure 3. A. Under dark conditions all genotypes show similar hypocotyl elongation, demonstrating that these mutants are not generally defective in elongation. B. Plants containing the lrb 1 -1 and lrb 2 -1 mutations have significantly shorter hypocotyls when treated with 10μmole m-2 sec-1 red light (indicative of increased sensitivity). The addition of the lrb 3 -1 mutation (lrb 1 -1/lrb 2 -1/lrb 3 -1) does not increase red light sensitivity. Funding Brandon has received funding from a UWEC Office of Research and Sponsored Programs Summer 2008/2009 Research Experience for Undergraduates grant and a Fall 2008/Spring 2009 Faculty/Student Collaborative Research grant. This work was partially funded by a National Science Foundation-Research in Undergraduate Institutions grant (#0919678)
- Slides: 1