PHANG LAB TALK Tzu L Phang Ph D

PHANG LAB TALK Tzu L Phang Ph. D. Assistant Professor Department of Medicine Division of Pulmonary Sciences & Critical Care Medicine

What I do: • Perform high-throughput data analysis for the scientific community; microarray and Next Generation Sequencing datasets • Provide analysis solution for experts and novice users alike • Develop multi-media approaches to disseminate translational science education • Studying the role of long non-coding RNA; second talk • Establishing the Bioinformatics Consultation and Analysis Core to help researchers and scientists design, analyze and interpret their experiments.

Today’s Talk Layout • The center of my universe: – R and Bioconductor • Collaboration with Biologists • 5 x 5; simple way to teach and contribute • Next Generation Sequencing (NGS)

Today’s Talk Layout • The center of my universe: – R and Bioconductor • Collaboration with Biologists • 5 x 5; simple way to teach and contribute • Next Generation Sequencing (NGS)
R r-project. org
R is hot http: //blog. revolutionanalytics. com/r-is-hot/
R in the media

Bioconductor • www. bioconductor. org • Statistical tools in R for high-throughput data analysis • 6 month update cycle. Release 2. 10 with 554 software package (45 new) • Analysis workflow – – – Oligonucleotide Arrays Sequence Analysis Variants Accessing Annotation Data High-throughput Assays

The Website www. bioconductor. org
Categories
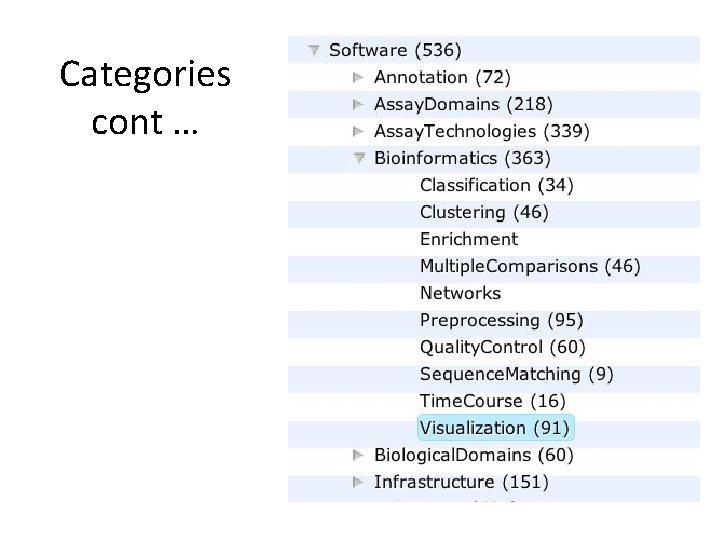
Categories cont …
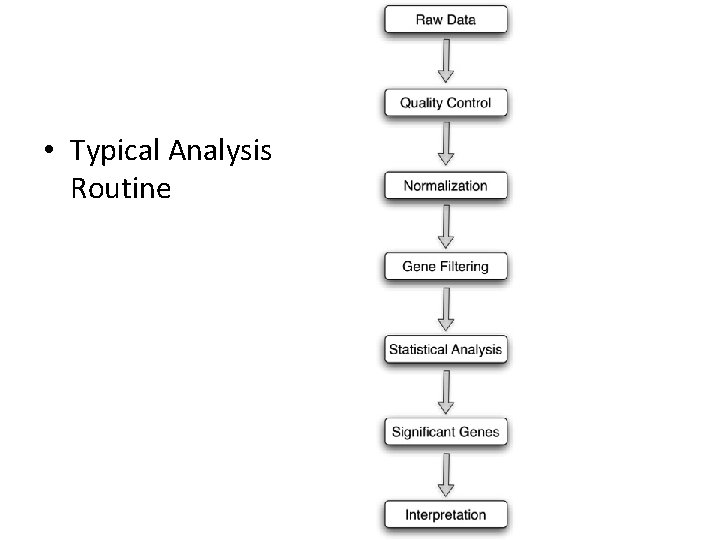
• Typical Analysis Routine

R is easy

Result output

Other Resources http: //www. rseek. org/ http: //crantastic. org/ http: //www. statmethods. net/ http: //stackoverflow. com/

Today’s Talk Layout • The center of my universe: – R and Bioconductor • Collaboration with Biologists • 5 x 5; simple way to teach and contribute • Next Generation Sequencing (NGS

Collaboration • >1000 microarray chips / year • Affymetrix & Illumina platforms • Next Generation Sequencing 25 free Pilot Projects. • Serve the rocky mountain region scientific community

Collaboration - tips • Don’t be a data analyst – be a co-investigator • Suggest analysis approaches that are not obvious • Focus on the result, not method • Always looks for grant writing opportunity • Understand the technical & biological system as thoroughly as possible – you will be surprise what biologists missed informatically

Exmaple 1: Classification of Pituitary Tumors • Pituitary tumors are the most common type of brain tumor in 20% at autopsy and 1/10, 000 persons clinically. Based upon 2010 figures of a veteran population of 22. 7 million, this translates into >225, 000 veterans with pituitary tumors. • Currently no medical therapies exist for these tumors and surgical resection is the treatment of choice. Recurrence rates approach 40%. • Understanding of the pathways to tumorigenesis and markers of aggressiveness and risk of recurrence would alter the intensity and cost of clinical care and may provide novel candidates and pathways to explore for new treatment options for these patients
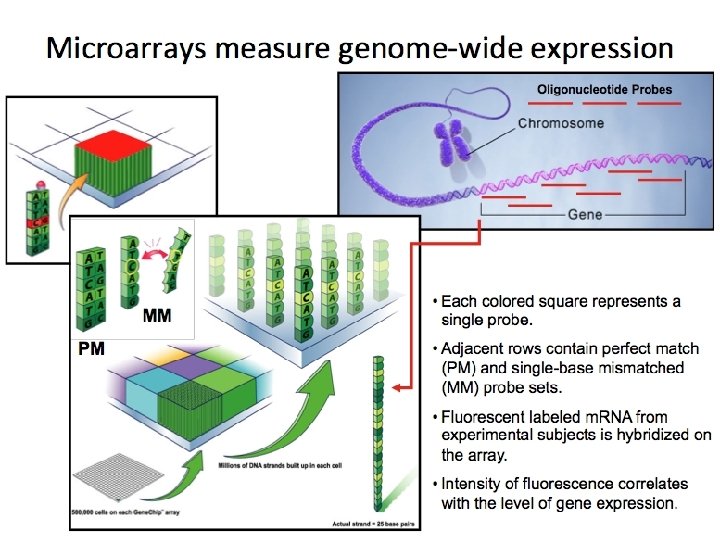
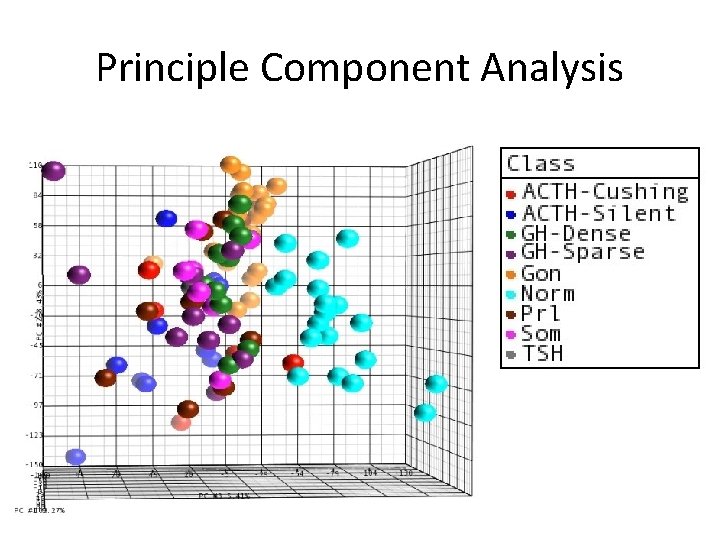
Principle Component Analysis
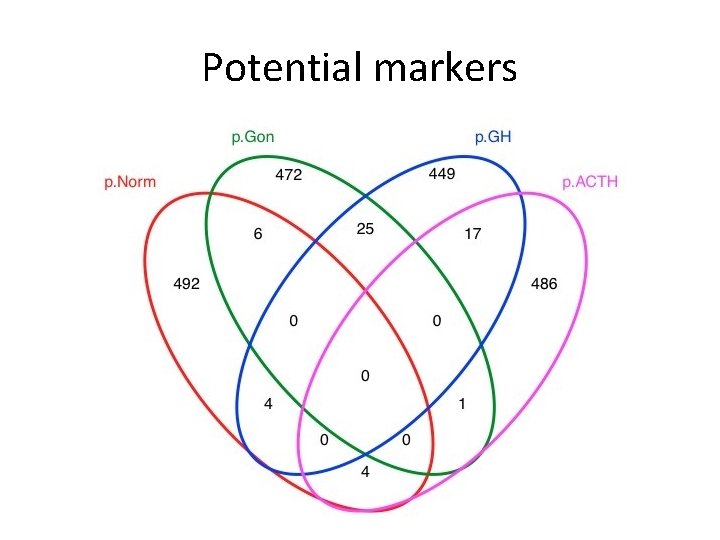
Potential markers

Outputs

Example 2: Explore the artistic side!
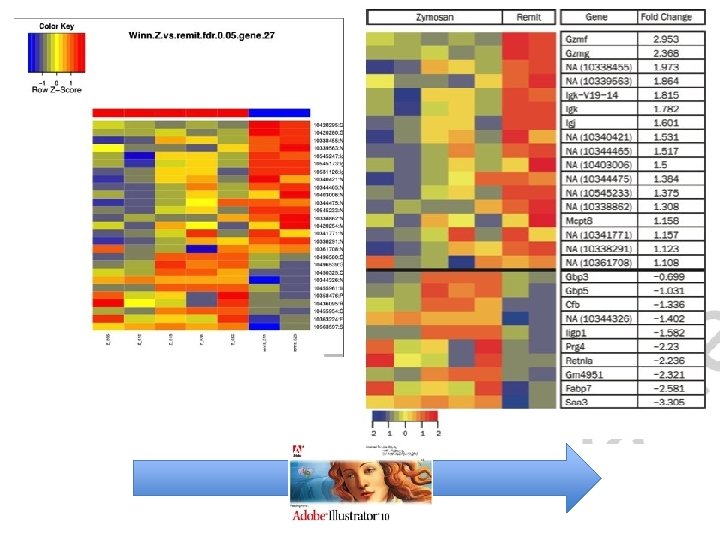

Example 3: Unconventional Usage
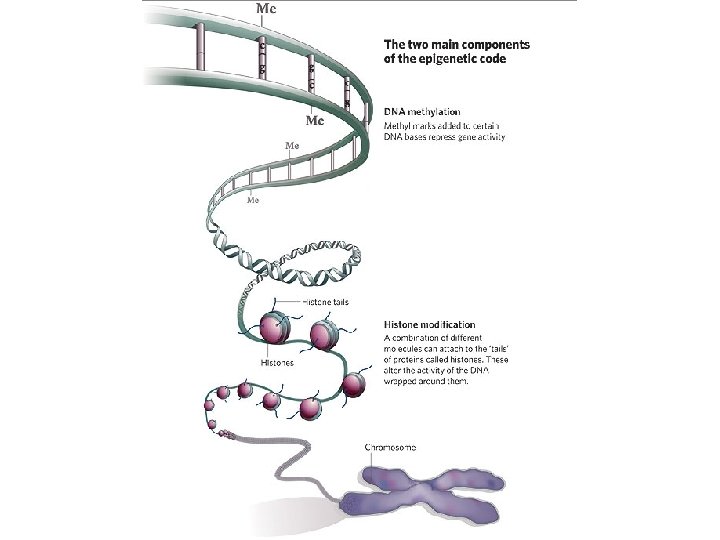

Introduction • Crohn’s Disease (CD) is an Inflammatory Bowel Disease (IBD) that affecting up to one million Americans (15 to 30 ages). • Discordance between monozygotic twins affected by CD provide evidence for epigenetic role in etiology of disease. • We combined 2 microarray technologies to study these roles – CHARM array (Comprehensive High-throughout Array for Relative Methylation) – Gene Expression (Affymetix Gene 1. 0 ST)
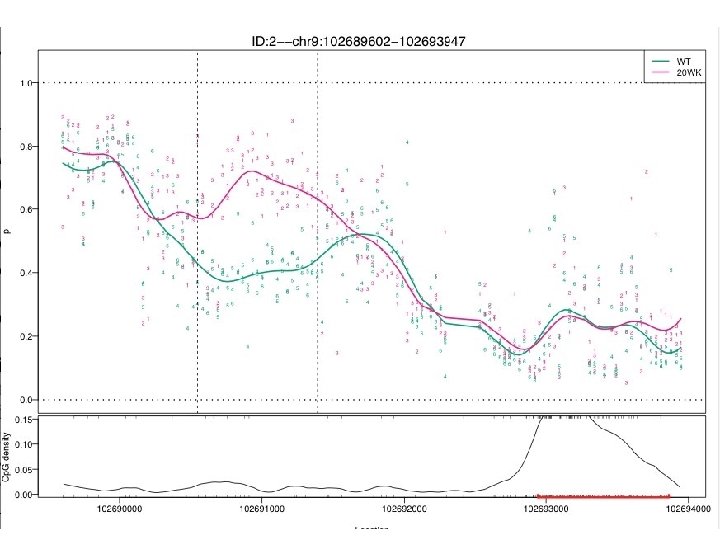

Research Informatics Integrated Core (RIIC) Michael G. Kahn MD, Ph. D CCTSI Co-Director & RIIC Core Director Michael. Kahn@ucdenver. edu
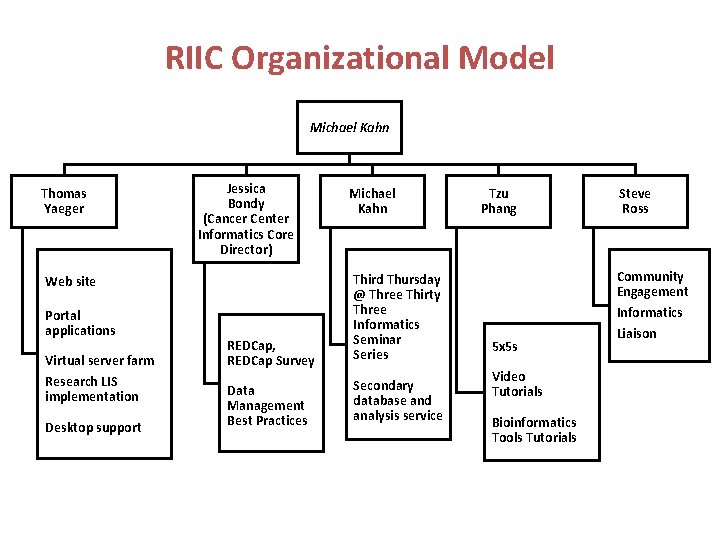
RIIC Organizational Model Michael Kahn Thomas Yaeger Jessica Bondy (Cancer Center Informatics Core Director) REDCap, REDCap Survey Third Thursday @ Three Thirty Three Informatics Seminar Series Data Management Best Practices Secondary database and analysis service Web site Portal applications Virtual server farm Research LIS implementation Desktop support Michael Kahn Tzu Phang 5 x 5 s Video Tutorials Bioinformatics Tools Tutorials Steve Ross Community Engagement Informatics Liaison

http: //cctsi. ucdenver. edu/RIIC/Pages/Consultation. Data. Analysis. aspx


5 X 5 http: //cctsi. ucdenver. edu/5 x 5

Demonstration http: //gcrc. ucdenver. edu/Videos/Informatics/5 x 5/Social. Networking 5 x 5. wmv

Tools
Podcast

TIES – Translational Informatics Education Support (TIES) • Bridging the gap in translational research through education • Training biologist informatics • Enhance collaboration through education and knowledge exchange • Bring awareness in latest technical advances • Disseminate knowledge through innovation

Next Generation Sequencing The future is here ….

High Throughput Parallel Sequencing • http: //www. youtube. com/watch? v=77 r 5 p 8 IBw. Jk

Paradigm Shift • Standard “Sanger” sequencing – 96 sample/day – Read length ~650 bp – Total = 450, 000 bases of sequence data • 454 – the game changer! – ~400, 000 different templates (reads)/day – Read length ~ 250 (at that time) – Total = 100, 000 bases of sequence data

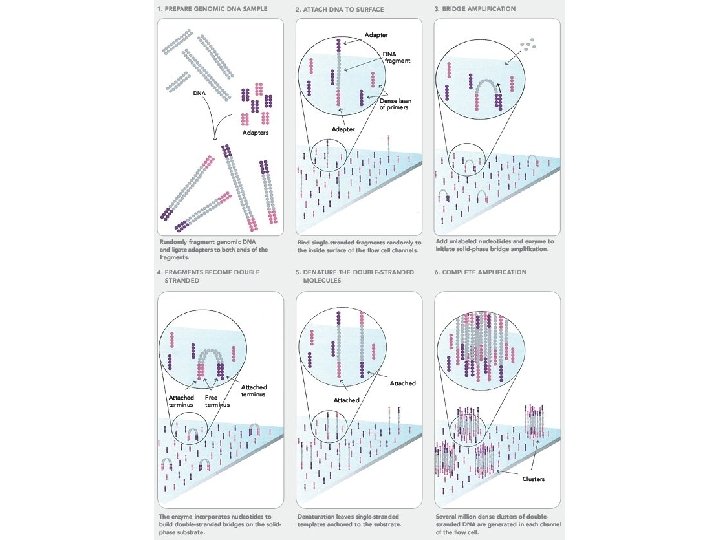
The second generation Roche (454) http: //454. com/ – First on the market – Emulsion PCR and pyrosequencing Illumina (Solexa) http: //www. illumina. com/ – Second on the market – Bridge PCR and polymerase based SBS Abi (Solid) http: //solid. appliedbiosystems. com/ – Third on the market – Emulsion PCR and ligase based sequencing

Single molecule sequencing Helicos Biosciences http: //helicosbio. com true Single Molecule Sequencing technology Pacific Biosciences http: //www. pacificbiosciences. com Single Molecule Real Time sequencing

Portable Sequencer • Ion Torrent

Others Polonator http: //www. polonator. org Emulsion PCR and ligase based sequencing Used in the Personal Genome Project Open platform, open source Cheap/affordable Complete Genomics http: //www. completegenomics. com Specializing in human genome sequencing
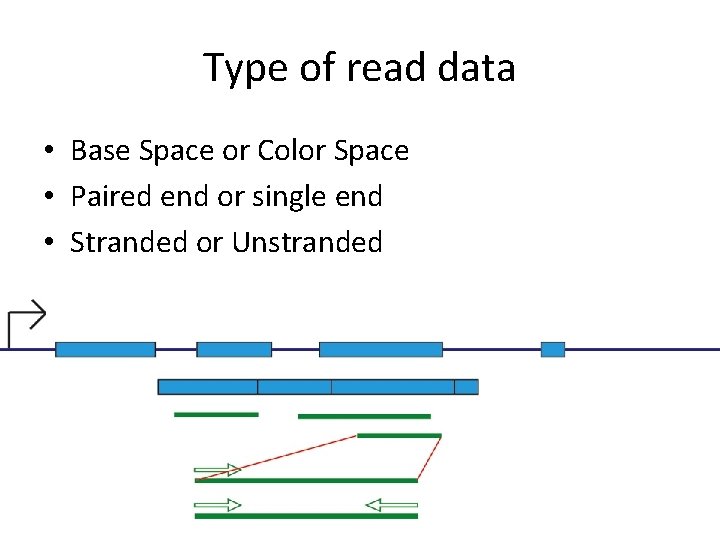
Type of read data • Base Space or Color Space • Paired end or single end • Stranded or Unstranded

Short Reads • Short reads from NGS are challenging (Solexa ~36 bp, now Hi. Seq 100 bp single pass) – Very hard to assemble whole genome – Especially on repeat regions • Requires many fold coverage • New and faster algorithm for many traditional bioinformatics operations • Reads are getting longer – another moving target. (2 x 250)

Applications • An explosion of scientific innovation!! • New usages not directly foreseen by the original developers of the technology • Some envision the beginning of next revolution – such as PCR – NGS machine in every lab!! • Cheap high-volume sequencing – revisiting data collection and management system

RNA Sequencing • “Digital Gene Expression” or “RNA-Seq” • Truly accurate gene expression measurements – Can replace gene expression microarrays • 25% more sensitive • Does not rely on hybridization (no %GC bias, no crosshybridization between related genes) • Discover novel genes (and other kinds of RNA molecules) – one experiment found that 34% of human transcripts were not from known genes • Sultan et al, Science. 2008 Aug 15; 321(5891): 956 -60.

Why RNAseq better then microarray? • Not predefine gene annotation — make discovering novel transcripts possible • Low, if any, background • Large dynamic range of expression levels, no upper limit for quantification • Reveal sequence variation, such as SNP, in the transcript region • In Helico — single molecule sequencing — no PCR step, remove amplification bias

More information from RNA • Can capture true alternative splicing information – Sequence of splice-junctions • One study found 4, 096 previously unknown splice junctions in 3, 106 human genes – Different transcription start and end points for RNA molecules • Allelic variation (SNPs) • Small RNAs


Bottleneck: Data Analysis

Informatics is the Bottleneck • Scientists are currently able to generate sequence data much faster/more easily than they are able to analyze it • Customized analysis / Bioinformatics consulting is needed for every project

Bioinformatics Challenges • Need for large amount of CPU power – Informatics groups must manage compute clusters – Challenges in parallelizing existing software or redesign of algorithms to work in a parallel environment – Another level of software complexity and challenges to interoperability • VERY large text files (million lines long) – Can’t do ‘business as usual’ with familiar tools such as Microsoft Excel. – Impossible memory usage and execution time • Sequence Quality filtering
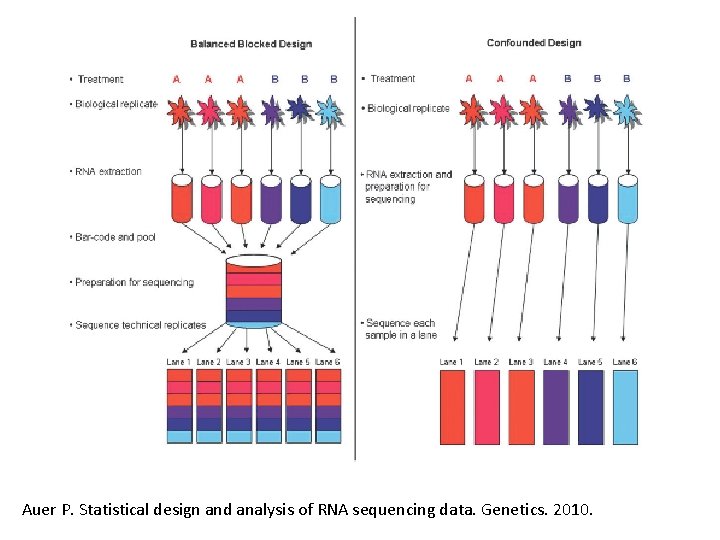
Auer P. Statistical design and analysis of RNA sequencing data. Genetics. 2010.

Data formats • • • Images “raw” basecalls with quality scores Sequence reads aligned to reference genomes Assemblies (contigs) Variants (SNPs, indels, copy number variants)

Hexadecimal mode Decimal mode
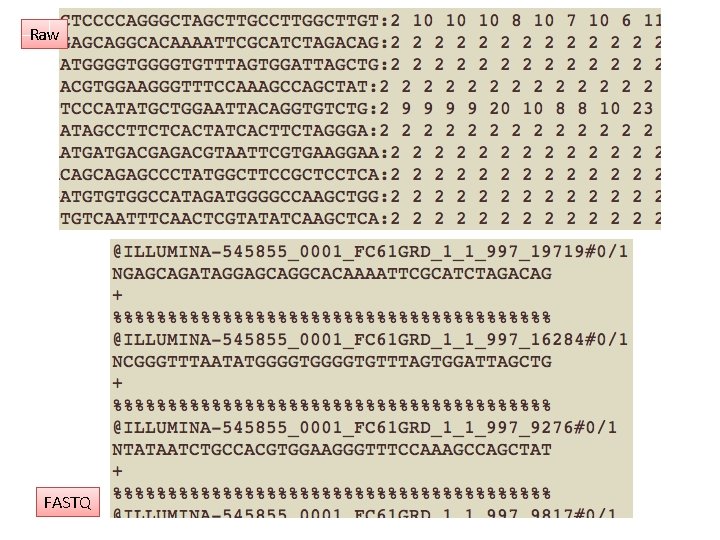
Raw FASTQ

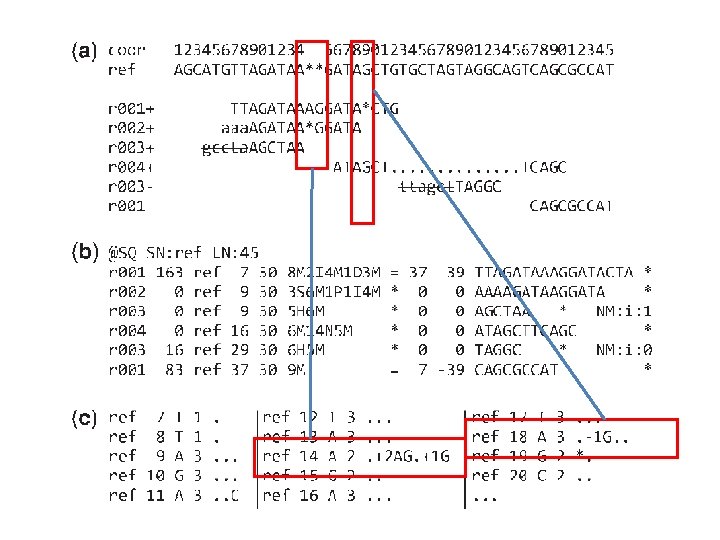
Example SAM format Pileup format
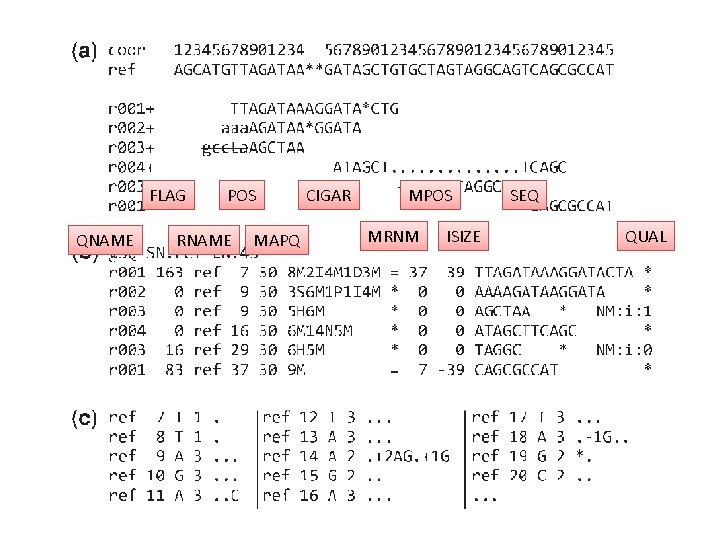
FLAG QNAME POS RNAME MAPQ CIGAR MPOS MRNM ISIZE SEQ QUAL

CIGAR • • M : match/mismatch I : Insertion compared with reference D : Deletion compared with reference N : Skipped bases on reference S : soft clipping (unaligned) H : hard clipping P : padding





![File Size • • • s_1_ILS 4_sequence. txt [5. 2 GB] s_1_ILS 4_sequence. fastq File Size • • • s_1_ILS 4_sequence. txt [5. 2 GB] s_1_ILS 4_sequence. fastq](http://slidetodoc.com/presentation_image_h/8c9a10839e2f2faefa5c997121a9cb95/image-73.jpg)
File Size • • • s_1_ILS 4_sequence. txt [5. 2 GB] s_1_ILS 4_sequence. fastq [3. 3 GB] s_1_ILS 4_sequence. sam [4. 5 GB] s_1_ILS 4_sequence. bam [995 MB] s_1_ILS 4_sequence. sorted. bam [696 MB]
The Bible

Utility Tools • • Sam. Tools Picard Useq Etc …

Bioconductor Solution
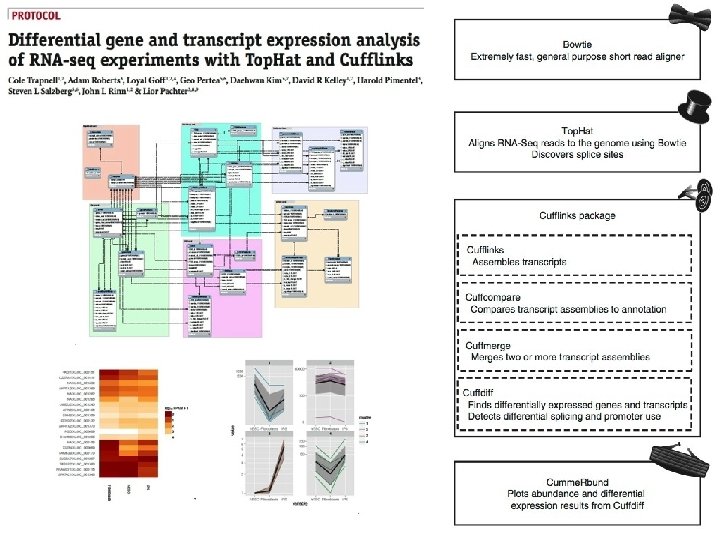

A demonstration
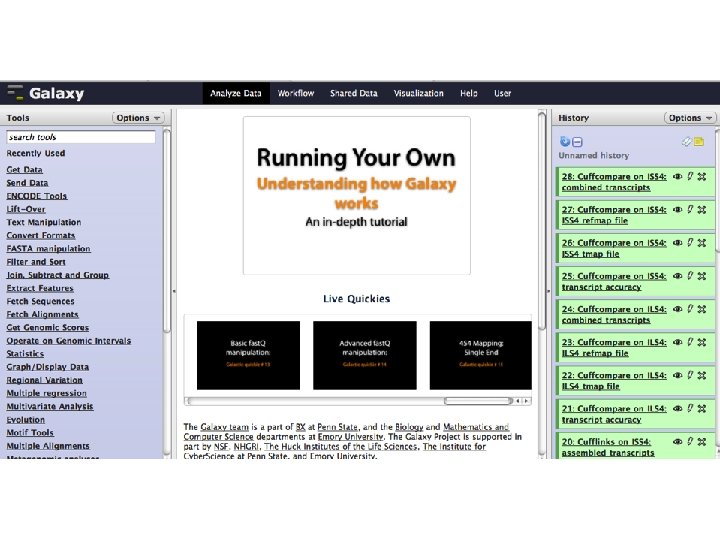

Secondary Tools Laboratory Management Data mining and visualization Project management for genome assembly Pathway mapping (functional analysis of groups of genes) • Motif finding (for Chip-Seq) • •

Integration • Integrate information from different technologies on a single genome map – Genetic variation – Gene expression (m. RNA levels) – Alternative splicing – Transcription factor binding – Methylation/histone status – Small RNA levels (gene regulatory molecules) – Non-coding RNA levels!

Speed/Efficiency • New emphasis on efficient data structures and algorithms • Use of “old style” tools such as grep/sed/awk • Machine language programming • Currently a huge burst of programming creativity in an “anything goes” environment • A desperate scramble for tools that work • Huge duplication of effort in programming, but also in evaluating new software

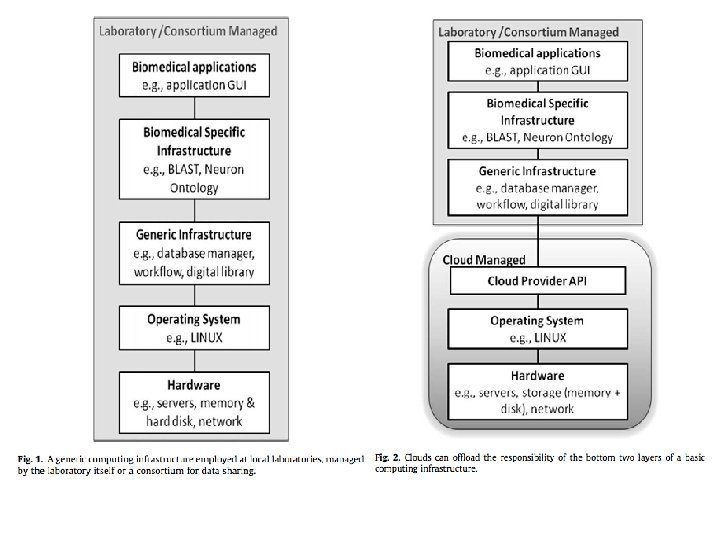
Amazon Web Services http: //aws. amazon. com/education/

Future Directions • Sequencing will continue to get much faster and cheaper, by 4 -10 x per year for several more years. • Affordable complete human genome sequencing will be available as a clinical diagnostic tool within 2 -3 years. • Data storage and analysis bottleneck • Data security/privacy issues
Move to 1: 52

Field Trip
- Slides: 91