Pfam DAS Rob Finn 26 th Feb 2007

Pfam & DAS Rob Finn 26 th Feb 2007

Pfam Acknowledgements • • • John Tate Roger Pettett Andreas Prlic Andy Jenkinson But takes data from community…. . !

Pfam New Pfam Website!

Pfam DA New Pfam Website! S En ab l ed

Pfam What do we provide? • • • Pfam Domains Active Sites Other sequence Features Alignment features

Pfam Alignment Client • Motivation – Simple alignment client • Not trying to reinvent Jalview – Markup alignment – Pfam alignments vary in size • Paging

Pfam Alignment Client • Example - combines alignment and features requests

Pfam Features Client • Motivation – Include Other annotations • Identify where we are missing domains – Reduce data duplication – Allow tailored views – Enrich single protein data in Pfam
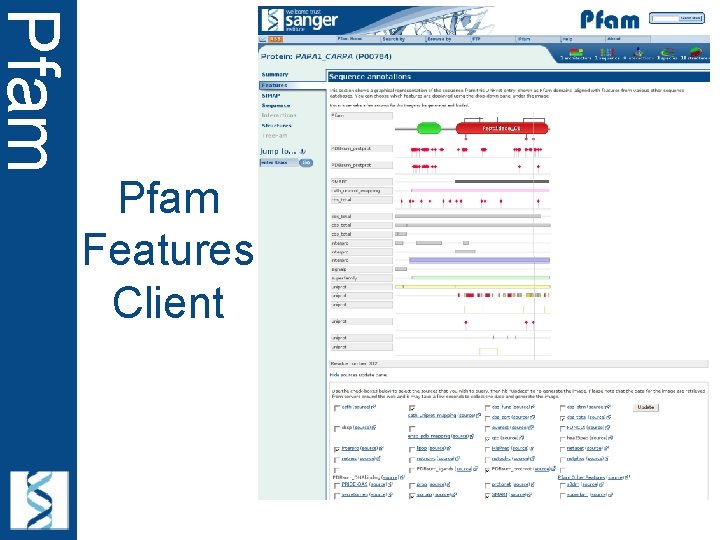
Pfam Features Client
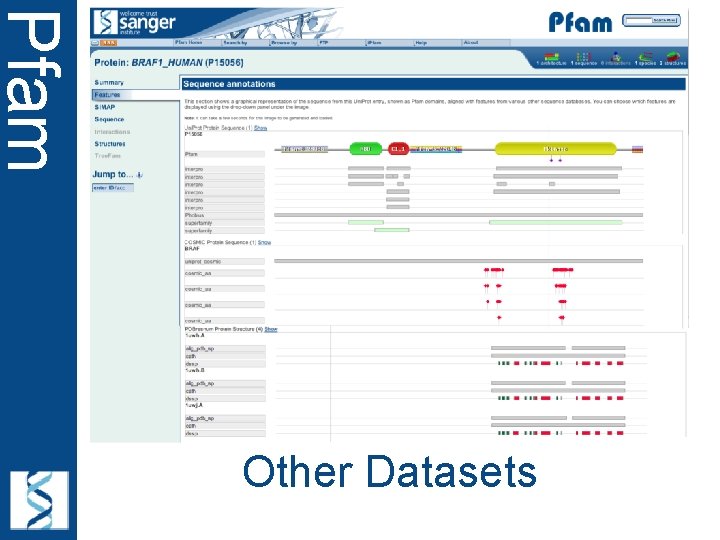
Pfam Other Datasets

Pfam What Code Base? • Pfam is Perl orientated – Server • Pro. Server (CPAN) – Client • Bio: : Das: : Lite (CPAN) • Bio: : Pfam: : Drawing – Break out?

Pfam What Can’t I Do (with Das)? • Asynchronous requests – Return pfam_scan results via DAS • Mapping experimental datasets to public datasets • Serve Pfam Textual Annotation – SOAP implementation

Pfam What is Hard to Do? • Large scale analyses • Filter out rubbish! – But what is one man’s rubbish…. • Requirement for stylesheet? • Standard Graphics library?

Pfam What Worries Me? • Data deluge – Make Views…. ? • Other people serve Pfam data – Confusion…. . ? – Redundancy • People always want more from clients

Pfam Thank you!
- Slides: 15