Pfam DAS and the future Rob Finn DAS

Pfam, DAS and the future Rob Finn DAS Workshop 2009

What is Pfam? – Protein families/domain database • Complete and accurate classification of protein space • Each family represented by alignments and profile HMMs – Two Distinct Parts • Pfam-A - high quality, curated, annotation • Pfam-B - low quality, automated, unannotated – Additional Features • Active site, coiled-coils, low complexity, transmembrane regions
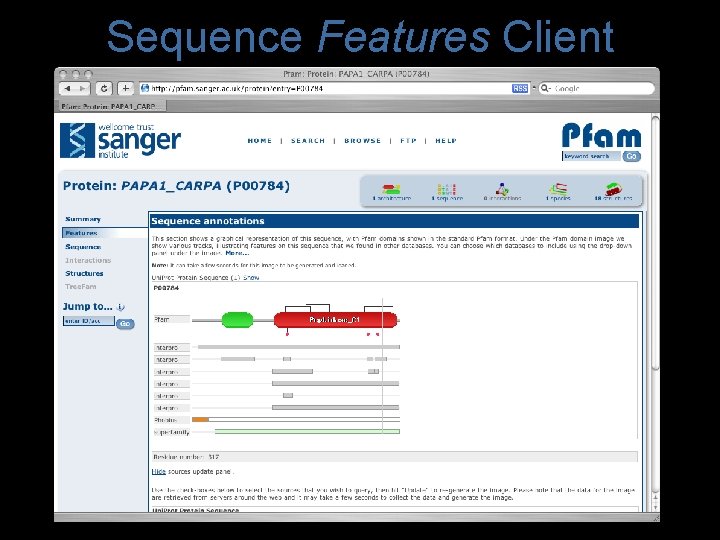
Sequence Features Client

Sequence Features Client • Motivation – Include Other annotations • Identify where we are missing domains – Reduce data duplication – Enrich single protein data in Pfam – Allow tailored views

Tailored Features Views • Updates from DAS registry
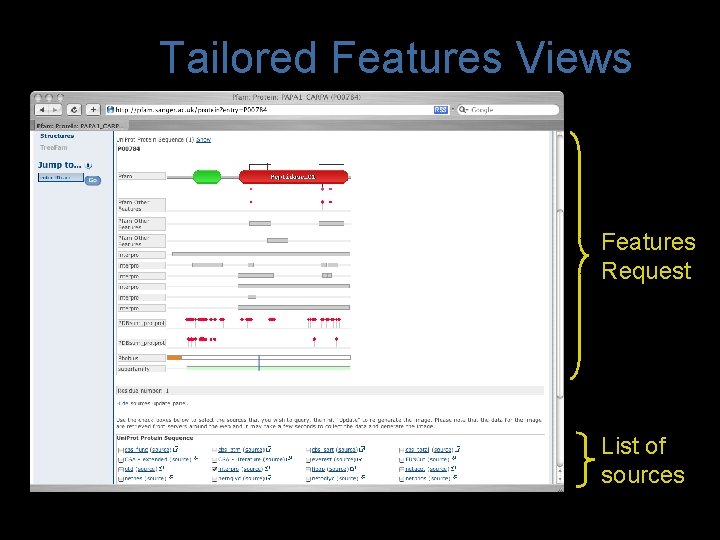
Tailored Features Views Features Request List of sources
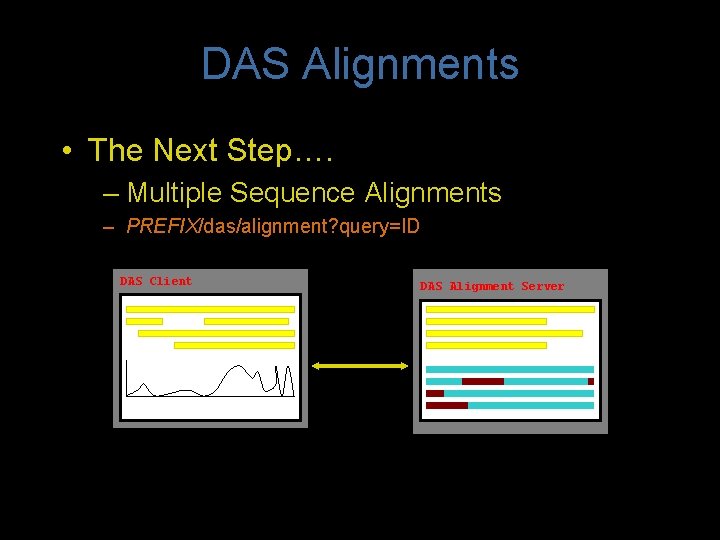
DAS Alignments • The Next Step…. – Multiple Sequence Alignments – PREFIX/das/alignment? query=ID DAS Client DAS Alignment Server
![DAS Alignments • Dealing with large alignments – PREFIX/das/alignment? query=ID[&subject=ID[RANGE]] or/and [&rows=START-END} DAS Alignment DAS Alignments • Dealing with large alignments – PREFIX/das/alignment? query=ID[&subject=ID[RANGE]] or/and [&rows=START-END} DAS Alignment](http://slidetodoc.com/presentation_image/49299287cfa8bed7660b3a256a52ac27/image-8.jpg)
DAS Alignments • Dealing with large alignments – PREFIX/das/alignment? query=ID[&subject=ID[RANGE]] or/and [&rows=START-END} DAS Alignment Server DAS Client X
![DAS Alignments • Dealing with large alignments – PREFIX/das/alignment? query=ID[&rows=START-END] DAS Alignment Server DAS DAS Alignments • Dealing with large alignments – PREFIX/das/alignment? query=ID[&rows=START-END] DAS Alignment Server DAS](http://slidetodoc.com/presentation_image/49299287cfa8bed7660b3a256a52ac27/image-9.jpg)
DAS Alignments • Dealing with large alignments – PREFIX/das/alignment? query=ID[&rows=START-END] DAS Alignment Server DAS Client DAS Align Feature Server

In Practice – Pfam alignments vary in size • 2 - 80, 000+ sequences • Paging Essential – Simple DAS alignment client • HTML, AJAX Pfam Alignments

Future Directions • More alignment sources are on their way! – Develop standalone, generic application – Paging replaced for ‘Live Grid’ • Issues – Genomics alignments! – Layering on features

HMMER 3 • Faster and more sensitive version of underlying software – Make use of new features? Query Length Pfam (140 X 11000) 20 0. 02 400 0. 41 35000 35. 93 Real time DAS searches!

Hot Alignments Can we scale efficiently?

Bringing in other datasets • Pfam – NCBI NR (gen. Pept) – Metagenomics • COSMIC - Catalogue Of Somatic Mutations In Cancer
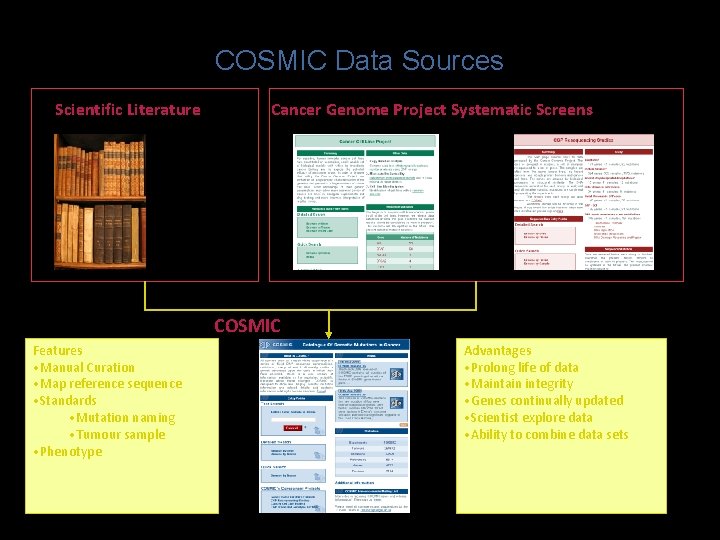
COSMIC Data Sources Scientific Literature Cancer Genome Project Systematic Screens COSMIC Features • Manual Curation • Map reference sequence • Standards • Mutation naming • Tumour sample • Phenotype Advantages • Prolong life of data • Maintain integrity • Genes continually updated • Scientist explore data • Ability to combine data sets
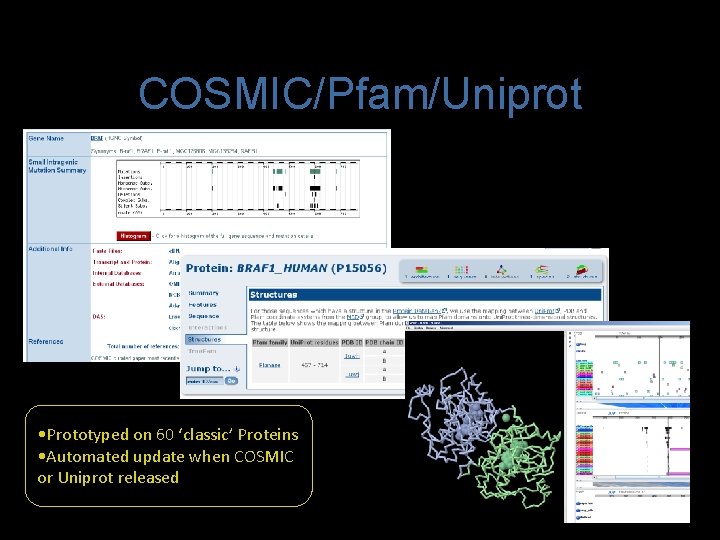
COSMIC/Pfam/Uniprot • Prototyped on 60 ‘classic’ Proteins • Automated update when COSMIC or Uniprot released
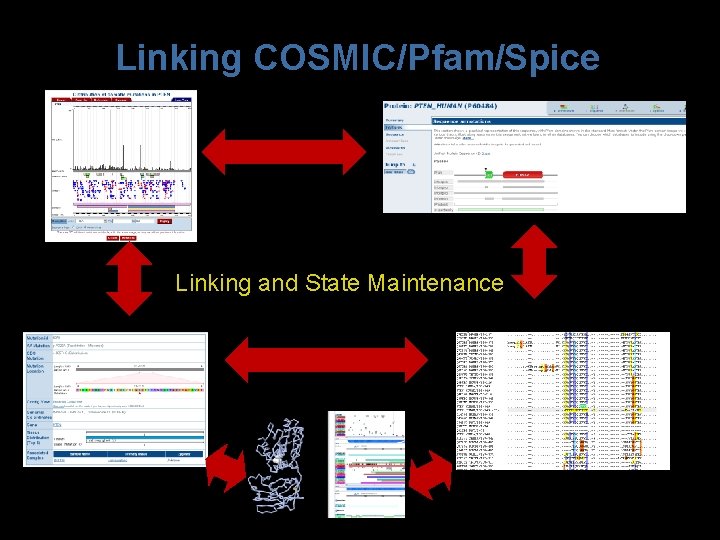
Linking COSMIC/Pfam/Spice Linking and State Maintenance

Acknowledgements • Pfam – Prasad Gunasekaran – John Tate – Alex Bateman – Penny Coggill – Jaina Mistry • COSMIC – Jon Teague – Cosmic team…… Questions?

- Slides: 19