Peter J Russell CHAPTER 20 Transposable Elements edited

Peter J. Russell CHAPTER 20 Transposable Elements edited by Yue-Wen Wang Ph. D. Dept. of Agronomy, 台大農藝系 NTU 遺傳學 601 20000 1

General Features of Transposable Elements 1. Transposable elements are divided into two classes on the basis of their mechanism for movement: a. Some encode proteins that move the DNA directly to a new position or replicate the DNA to produce a new element that integrates elsewhere. This type is found in both prokaryotes and eukaryotes. b. Others are related to retroviruses, and encode reverse transcriptase for making DNA copies of their RNA transcripts, which then integrate at new sites. This type is found only in eukaryotes. 2. Transposition is nonhomologous recombination, with insertion into DNA that has no sequence homology with the transposon. a. In prokaryotes, transposition can be into the cell’s chromosome, a plasmid or a phage chromosome. b. In eukaryotes, insertion can be into the same or a different chromosome. 3. Transposable elements can cause genetic changes, and have been involved in the evolution of both prokaryotic and eukaryotic genomes. Transposons may: a. Insert into genes. b. Increase or decrease gene expression by insertion into regulatory sequences. c. Produce chromosomal mutations through the mechanics of transposition. 台大農藝系 遺傳學 601 20000 2

Transposable Elements in Prokaryotes 1. Prokaryotic examples include: a. Insertion sequence (IS) elements. b. Transposons (Tn). c. Bacteriophage Mu (replicated by transposition) 台大農藝系 遺傳學 601 20000 3

Insertion Sequences Animation: Insertion Sequences in Prokaryotes 1. IS elements are the simplest transposable elements found in prokaryotes, encoding only genes for mobilization and insertion of its DNA. IS elements are commonly found in bacterial chromosomes and plasmids. 2. IS elements were first identified in E. coli’s galactose operon, wheresome mutations’ were shown to result from insertion of a DNA sequence now called IS 1 (Figure 20. 1) 3. Prokaryotic IS elements range in size from 768 bp to over 5 kb. Known E. coli IS elements include: a. IS 1 is 768 bp long, and present in 4– 19 copies on the E. coli chromosome. b. IS 2 has 0– 12 copies on the chromosome, and 1 copy on the F plasmid. c. IS 10 is found in R plasmids. 4. The ends of all sequenced IS elements show inverted terminal repeats (IRs) of 9– 41 bp (e. g. , IS 1 has 23 bp of台大農藝系 nearly identical 遺傳學 601 sequence). 20000 4
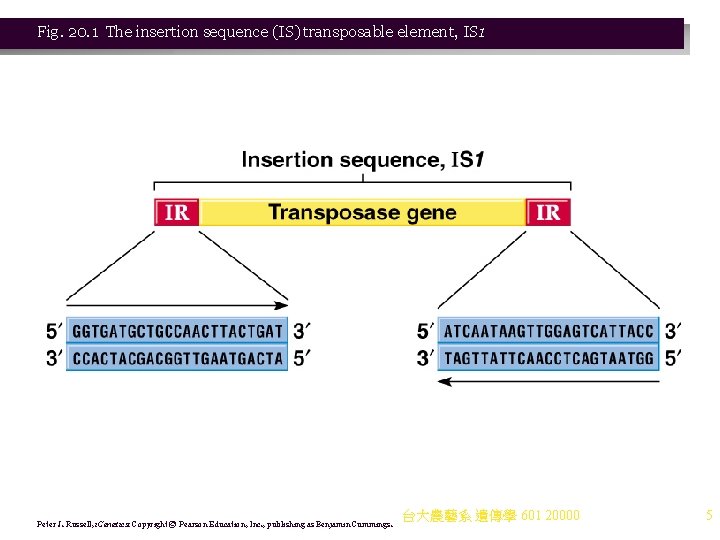
Fig. 20. 1 The insertion sequence (IS) transposable element, IS 1 Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 5

5. Integration of IS elements may: a. Disrupt coding sequences or regulatory regions. b. Alter expression of nearby genes by the action of IS element promoters. c. Cause deletions and inversions in adjacent DNA. d. Serve as a site for crossing-over between duplicated IS elements. 6. When an IS element transposes: a. The original copy stays in place, and a new copy inserts randomly into the chromosome. b. The IS element uses the host cell replication enzymes for precise replication. c. Transposition requires transposase, an enzyme encoded by the IS element. d. Transposase recognizes the IR sequences to initiate transposition. e. IS elements insert into the chromosome without sequence homology (illegitimate recombination) at target sites (Figure 20. 2). i. A staggered cut is made in the target site, and the IS element inserted. ii. DNA polymerase and ligase fill the gaps, producing small direct repeats of the target site flanking the IS element (target site duplications). f. Mutational analysis shows that IR sequences are the key 台大農藝系 遺傳學 601 20000 6
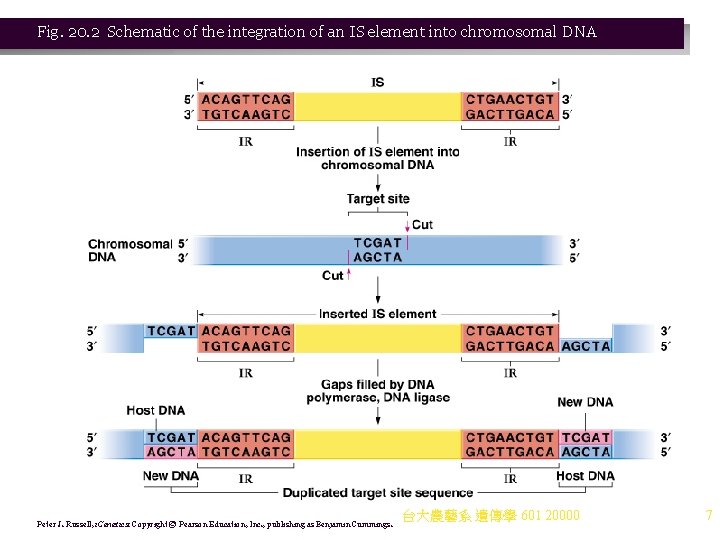
Fig. 20. 2 Schematic of the integration of an IS element into chromosomal DNA Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 7

Transposons 1. Transposons are similar to IS elements, but carry additional genes, and have a more complex structure. There are two types of prokaryotic transposons: a. Composite transposons carry genes (e. g. , antibiotic resistance) flanked on both sides by IS elements (IS modules). i. The IS elements are of the same type, and called ISL (left) and ISR (right). ii. ISL and ISR may be in direct or inverted orientation to each other. iii. Tn 10 is an example of a composite transposon (Figure 20. 3). It is 9. 3 kb, and contains: (1) 6. 5 kb of central DNA with genes that include tetracycline resistance (a selectable marker). (2) 1. 4 kb IS elements (IS 10 L and IS 10 R) at each end, in an inverted orientation. iv. Transposition of composite transposons results from the IS elements, which supply transposase and its recognition signals, the IRs. (1) Tn 10’s transposition is rare, because transpose is produced at a rate of , 1 molecule/generation. (2) Transposons, like IS elements, produce target site duplications (e. g. , a 9 bp duplication for Tn 10). (Table 20. 1) 台大農藝系 遺傳學 601 20000 8
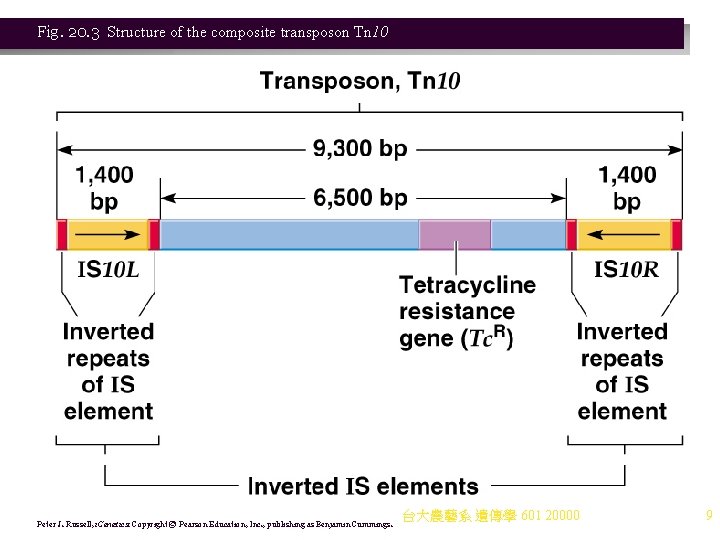
Fig. 20. 3 Structure of the composite transposon Tn 10 Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 9

b. Noncomposite transposons also carry genes (e. g. , drug resistance) but do not terminate with IS elements. i. Transposition proteins are encoded in the central region. ii. The ends are repeated sequences (but not IS elements). iii. Noncomposite transposons cause target site duplications (like composite transposons). iv. An example is Tn 3. (1) Tn 3’s length is about 5 kb, with 38 -bp inverted terminal repeats. (2) It has three genes in its central region: (a) bla encodes β-lactamase, which breaks down ampiciliin. (b) tnp. A encodes transposase, needed for insertion into a new site. (c) tnp. B encodes resolvase, involved in recombinational events needed for transposition (not found in all transposons). (3) Tn 3 produces a 5 -bp duplication upon insertion (Figure 20. 5). 台大農藝系 遺傳學 601 20000 10
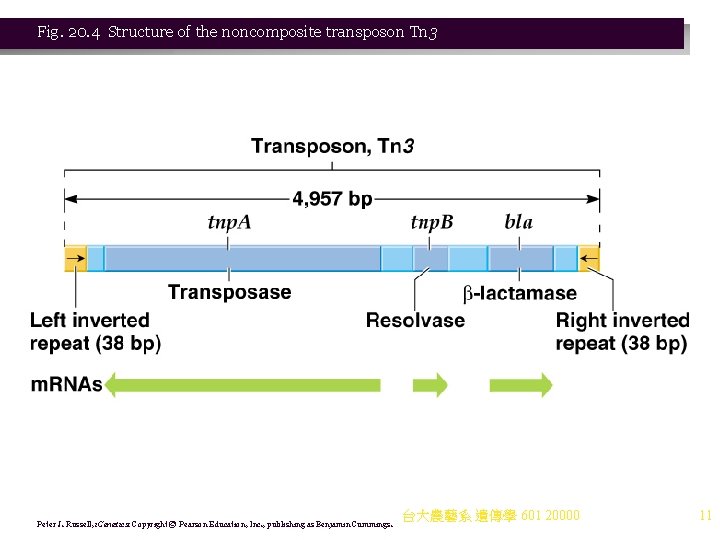
Fig. 20. 4 Structure of the noncomposite transposon Tn 3 Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 11
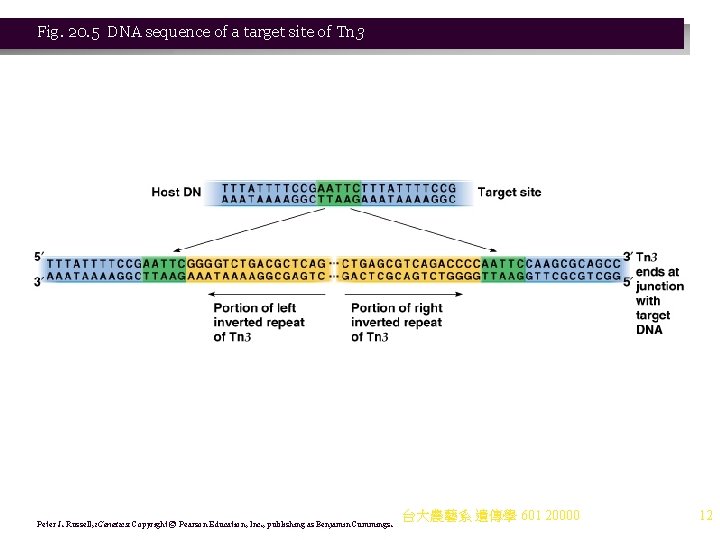
Fig. 20. 5 DNA sequence of a target site of Tn 3 Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 12

2. Models have been generated for transposition: a. Cointegration is an example of the replicative transposition that occurs with Tn 3 and its relatives (Figure 20. 6). i. Donor DNA containing the Tn fuses with recipient DNA. ii. The Tn is duplicated, with one copy at each donor-recipient DNA junction, producing a cointegrate. iii. The cointegrate is resolved into two products, each with one copy of the Tn. b. Conservative (nonreplicative) transposition is used by Tn 10, for example. The Tn is lost from its original position when it transposes. 3. Transposons cause the same sorts of mutations caused by IS elements: a. Insertion into a gene disrupts it. b. Gene expression is changed by adjacent Tn promoters. c. Deletions and insertions occur. d. Crossing-over results from duplicated Tn sequences in the genome. 台大農藝系 遺傳學 601 20000 13
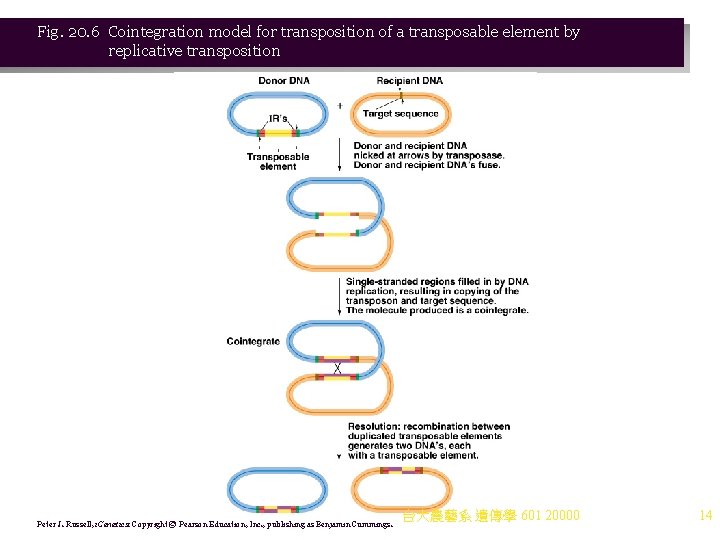
Fig. 20. 6 Cointegration model for transposition of a transposable element by replicative transposition Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 14

IS Elements and Transposons in Plasmids 1. Bacterial plasmids are extrachromosomal DNA capable of self-replication. Some are episomes, able to integrate into the bacterial chromosome. The E. coli F plasmid is an example (Figure 20. 7): a. Important genetic elements of the F plasmid are: i. tra genes for conjugal transfer of DNA from donor to recipient. ii. Genes for plasmid replication. iii. 4 IS elements: 2 copies of IS 3, 1 of IS 2, and 1 of γδ (gammadelta). All have homology with IS elements itt the E. coli chromosome. b. The F factor integrates by homologous recombination between IS elements, mediated by the tra genes. 2. R plasmids have medical significance, because they carry genes for resistance to antibiotics, and transfer them between bacteria (Figure 20. 7). a. Genetic features of R plasmids include: i. The resistance transfer factor region (RTF), needed for conjugal transfer. It includes a DNA region homologous to an F plasmid region, and genes for plasmid-specific DNA replication. ii. Differing sets of genes, such as those for resistance to antibiotics or heavy metals. The resistance genes are transposons, flanked by IS module-like sequences, and can replicate and insert into the bacterial chromosome. b. R plasmids are clinically significant, because they disseminate drug resistance genes between bacteria. 台大農藝系 遺傳學 601 20000 15
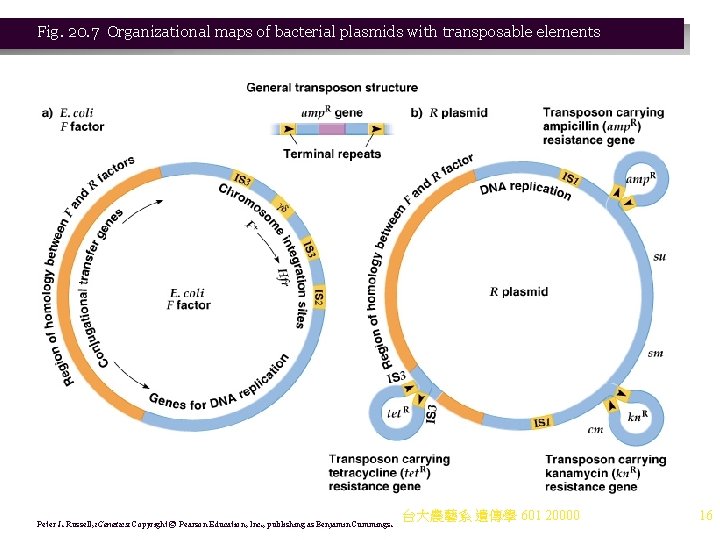
Fig. 20. 7 Organizational maps of bacterial plasmids with transposable elements Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 16

Bacteriophage Mu 1. Temperate bacteriophage Mu (mutator) can cause mutations when it transposes. Its structure includes: a. A 37 kb linear DNA in the phage particle that has central phage DNA and unequal lengths of host DNA at the ends (Figure 20. 8). b. The DNA’s G segment can invert, and is found in both orientations in viral DNA. 2. Following infection, Mu integrates into the host chromosome by conservative (non-replicative) transposition. a. Integration produces prophage DNA flanked by 5 bp target site direct repeats. b. Flanking DNA from the previous host is lost during integration. c. The Mu prophage now replicates only when the E. coli chromosome replicates, due to a phage-encocled repressor that prevents most Mu gene expression. 3. Mu prophage stays integrated during the lytic cycle, and replication of Mu’s genome is by replicative transposition. 4. Mu causes insertions, deletions, inversions and translocations (Figure 20. 9). 台大農藝系 遺傳學 601 20000 17
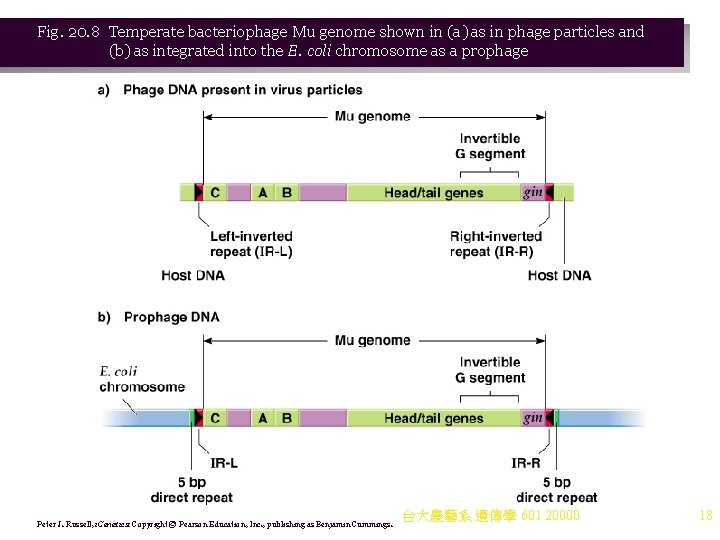
Fig. 20. 8 Temperate bacteriophage Mu genome shown in (a) as in phage particles and (b) as integrated into the E. coli chromosome as a prophage Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 18
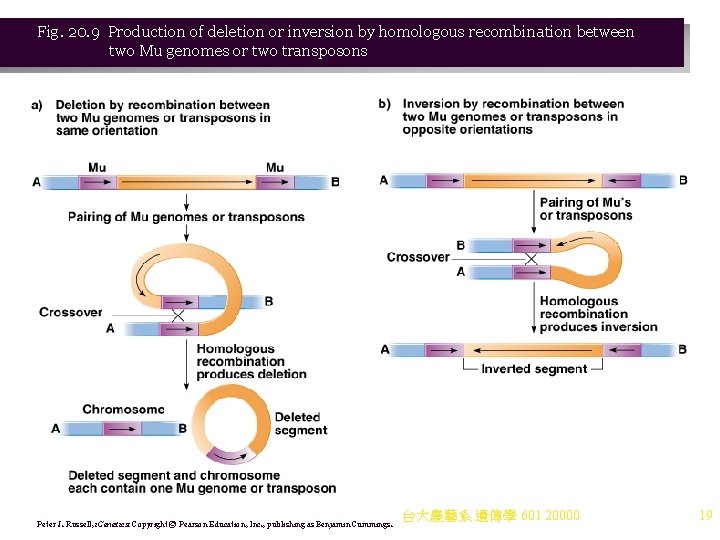
Fig. 20. 9 Production of deletion or inversion by homologous recombination between two Mu genomes or two transposons Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 19

Transposable Elements in Eukaryotes 1. Rhoades (1930 s) working with sweet corn, observed interactions between two genes: a. A gene for purple seed color, the Al locus. Homozygous mutants (a/a) have colorless seeds. b. A gene on a different chromosome, Dt (dotted) that causes seeds with genotype a/a Dt/-- to have purple dots. i. Dt appears to mutate the a allele back to the Al wild-type in regions of the seed, producing a dotted phenotype. ii. The effect of the Dt allele is dose dependent. (1) One dose gave an average of 7. 2 dots per seed. (2) Two doses gave an average of 22. 2 dots/seed. (3) Three doses gave an average of 121. 9 dots/seed. c. Rhoades interpreted Dt as a mutator gene. 2. Mc. Clintock (1940 s-50 s), working with corn (Zea mays) proposed the existence of “controlling elements” that regulate other genes and are mobile in the genome. 3. The genes studied by both Rhoades and Mc. Clintock have turned out to be transposable elements, and many others have been identified in various eukaryotes. a. Most studied are transposons of yeast, Drosophila, corn and humans. b. Their structure is very similar to that of prokaryotic transposable elements. c. Eukaryotic transposable elements have genes for transposition and integration at a number of sites, as well as a variety of other genes. d. Random insertion results from non-homologous recombination, and means that any 台大農藝系 遺傳學 601 20000 chromosomal gene may be regulated by a transposon. 20

Transposons in Plants Animation: Transposable Elements in Plants 1. Plant transposons also have IR sequences, and generate short direct target site repeats. 2. The result of transposon insertion into a plant chromosome will depend on the properties of the transposon, with possible effects including: a. Activation or repression of adjacent genes by disrupting a cellular promoter, or by action of transposon promoters. b. Chromosome mutations such as duplications, deletions, inversions, translocations or breakage. c. Disruption of genes to produce a null mutation (gene is nonfunctional). 3. Several families of transposons have been identified in corn, each with characteristic numbers, types and locations. a. Each family has two forms of transposon. Either can insert into a gene and produce a mutant allele. i. Autonomous elements, which can transpose by themselves. Alleles produced by an autonomous element are mutable alleles, creating mutations that revert when the transposon is excised from the gene. ii. Nonautonomous elements, which lack a transposition gene and rely on the presence of another transposon to supply the missing function. Mutation by these elements is stable (except when an autonomous element from the family is also present). 台大農藝系 遺傳學 601 20000 21

4. Multiple genes control corn color, and classical genetics indicates that a mutation in any of these genes leads to a colorless kernel. Mc. Clintock studied the unstable mutation that produces spots of purple pigment on white kernels (Figure 20. 10). a. She concluded that spots do not result from a conventional mutation, but from a controlling element (now Tn). b. A corn plant with genotype c/c will have white kernels, while C/-- will result in purple ones. i. If a reversion of c to C occurs in a cell, that cell will produce purple pigment, and hence a spot. ii. The earlier in development the reversion occurs, the larger the spot. 台大農藝系 遺傳學 601 20000 22

iii. Mc. Clintock concluded that the c allele resulted from insertion of a “mobile controlling element” into the C allele. (1) The element is Ds (dissociation), now known to be a nonautonomous transposon. (2) Its transposition is controlled by Ac (activator), an autonomous transposon (Figure 20. 11). c. Mc. Clintock’s evidence of transposable elements did not fit the prevailing model of a static genome. More recent studies have confirmed and characterized the elements involved. i. The Ac-Ds system involves an autonomous element (Ac) whose insertions are unstable, and a nonautonomous element (Ds) whose insertions are stable if only Ds is present. ii. Mc. Clintock (1950 s) showed that some Ds elements derive from Ac elements. 台大農藝系 遺傳學 601 20000 23
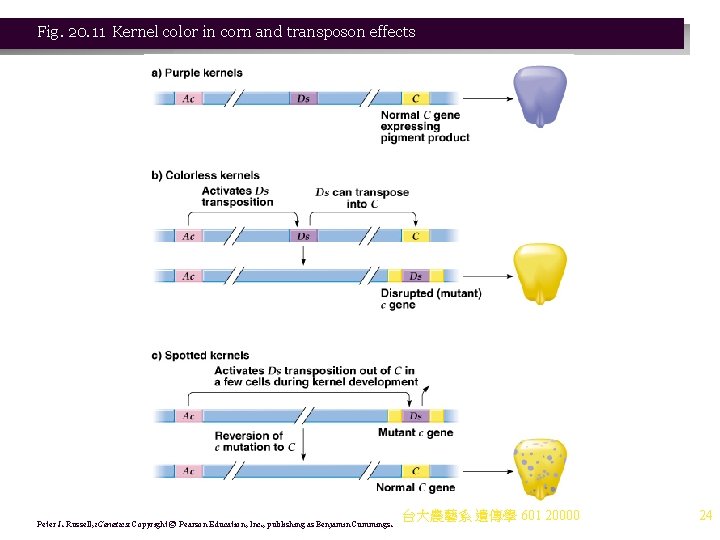
Fig. 20. 11 Kernel color in corn and transposon effects Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 24

iii. Ac is 4, 563 bp, with 1 1 -bp imperfect terminal IRs and 1 transcription unit producing a 3. 5 kb m. RNA encoding an 807 amino acid transposase. Insertion generates an 8 -bp target site duplication (Figure 20. 12). iv. Ac activates Ds to transpose or break the chromosome where it is inserted. v. Ds elements vary in length and sequence, but all have the same terminal IRs as Ac, and many are deleted or rearranged versions of Ac. vi. Unique to corn transposons, timing and frequency of transposition and gene rearrangements are developmentally regulated. vii. Ac transposes only during chromosome replication, and does not leave a copy behind. There are two possible results of Ac transposition, depending on whether the target DNA has replicated or not (Figure 20. 13). (1) If Ac transposes during replication into a replicated target site, its chromatid’s donor site will be empty since that copy of Ac has inserted elsewhere. In the homologous donor site on the other chromatid, a copy will remain. There is no net increase in copies of Ac. (2) Transposition to an unreplicated chromosome site also leaves one donor site empty (and the other with a copy of Ac). The DNA into which Ac inserts will then be replicated, resulting in a net gain of one copy of Ac. viii. Replication of Ds is the same, except that the transposition protein is supplied by an integrated Ac element. 台大農藝系 遺傳學 601 20000 25

Fig. 20. 12 The structure of the Ac autonomous transposable element of corn and of several Ds nonautonomous elements derived from Ac Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 26
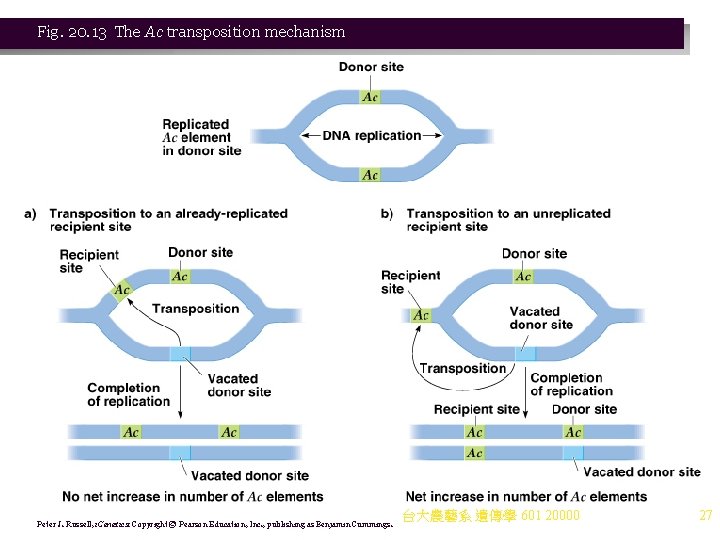
Fig. 20. 13 The Ac transposition mechanism Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 27

5. In Mendel’s wild-type (SS) peas the starch grains are large and simple, while in wrinlded peas (ss) they are small and fissured. a. SS seeds contain more starch and less sucrose than ss seeds. b. The sucrose difference makes ss seeds larger, with higher water content, so that when dried they are wrinided. c. One type of starch-branching enzyme (SBEI) is missing in ss plants, reducing their starch content. d. The SBEI gene corresponding to the s allele has a 0. 8 kb transposon similar to the Ax/Ds family inserted into the wildtype S allele. 台大農藝系 遺傳學 601 20000 28

Ty Elements in Yeast 1. Ty elements share characteristics with bacterial transposons: a. Terminal repeated sequences. b. Integration at non-homologous sites. c. Generation of a target site duplication (5 bp). 2. Ty element is diagrammed in Figure 20. 14: a. It is 5. 9 kb including 2 terminal direct repeats of 334 bp, the long terminal repeats (LTR) or deltas (δ). b. Each delta contains a promoter and transposase recognition sequences. c. Ty elements encode one 5. 7 kb m. RNA beginning at the delta 5’ promoter (Figure 20. 14). d. There are two ORFs in the m. RNA, designated Ty. A and Ty. B, encoding two different proteins. e. Ty copy number varies between yeast strains, with an average of about 35. 3. Ty elements also share similarities with retroviruses, ss. RNA viruses that replicate via ds. DNA intermediates. a. Ty elements transpose by making an RNA copy of the integrated DNA sequence, them making DNA using reverse transcriptase. This DNA can integrate at a new chromosomal site. Evidence for this includes: i. An experimentally introduced intron in the Ty element (which normally lacks introns) was monitored through transposition. The intron was removed, indicating an RNA intermediate. ii. Ty elements encode a reverse transcriptase. iii. Virus-like particles containing Ty RNA and reverse transcriptase activity occur. 台大農藝系 遺傳學 601 20000 29 b. Ty elements are referred to as retrotransposons.
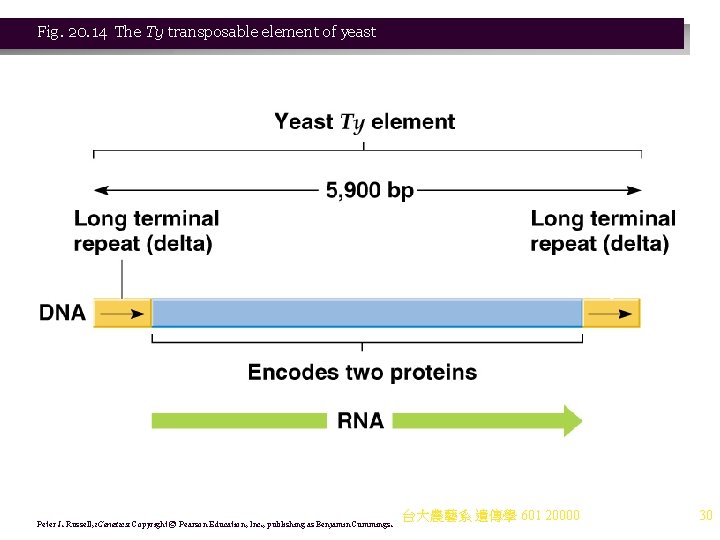
Fig. 20. 14 The Ty transposable element of yeast Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 30

Drosophila transposons 1. It is estimated that 15% of the Drosophila genome is mobile! These transposons fall into different classes: a. The copia retrotransposons include several families, each highly conserved and present in 5 -100 widely scattered copies per genome (Figure 20. 15). i. All copia elements in Drosophila can transpose, and there are differences in number and distribution between fly strains. ii. Structurally, copia elements are similar to yeast Ty elements: (1) Direct LTRs of 276 bp flank a 5 kb DNA segment. (2) The end of each LTR has 17 bp inverted repeats. (3) An RNA intermediate and reverse transcriptase are used for transposition. (4) Virus-like particles (VLPs) occur with copia. (5) Integration results in target site duplication (3 -6 bp). 台大農藝系 遺傳學 601 20000 31

Fig. 20. 15 Structure of the transposable element copia, a retrotransposon found in Drosophila melanogaster Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 32

b. P elements cause hybrid dysgenesis, a series of defects (mutations, chromosomal aberrations and sterility) that result from crossing certain Drosophila strains (Figure 20. 16). i. A mutant lab strain female (M) crossed with a wildtype male (P) will result in hybrid dysgenesis. ii. A mutant lab strain male (M) crossed with a wildtype (P) female (reciprocal cross) will have normal offspring. iii. Thus, hybrid dysgenesis results when chromosomes of the P male parent enter cytoplasm of an M type oocyte, but cytoplasm from P oocytes does not induce hybrid dysgenesis. 台大農藝系 遺傳學 601 20000 33
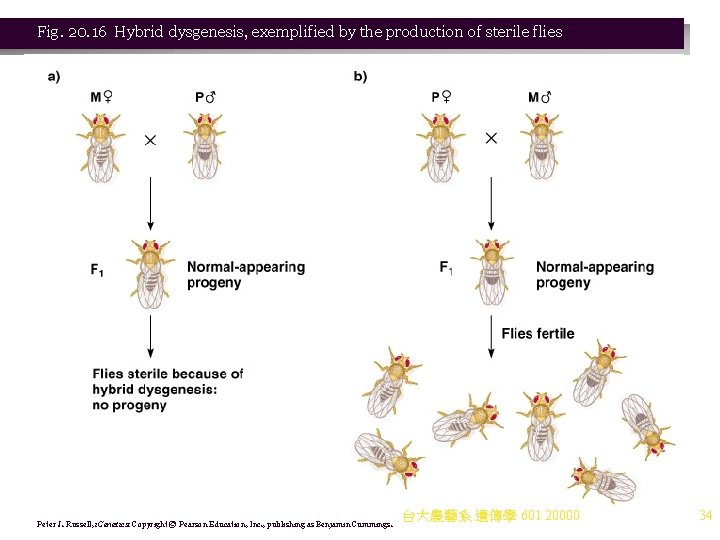
Fig. 20. 16 Hybrid dysgenesis, exemplified by the production of sterile flies Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 34

iv. The model is based on the observation that the M strain has no P elements, while the haploid genome of the P male has about 40 copies. (1) P elements vary from full-length autonomous elements through shorter versions resulting from a variety of internal deletions. (2) P element transposition is activated only in the germ line. (3) The F 1 of an M female crossed with a P male have P elements inserted at new sites, flanked by target site repeats. (4) P elements are thought to encode a repressor protein that prevents transposase gene expression, preventing transposition. (5) Cytoplasm in an M oocyte lacks the repressor, and so when fertilized with P-bearing chromosomes, transposition occurs into the maternal chromosomes, leading to hybrid dysgenesis. v. P elements are used experimentally to transfer genes into the germ line of Drosophila embryos. For example (Figure 20. 18): (1) The wild-type rosy (ry) gene was inserted into a P element, cloned in a plasmid and microinjected into a mutant ry/ry strain. (2) Insertion of the recombinant P element into the recipient chromosome introduced the ry allele, and produced wild-type flies. 台大農藝系 遺傳學 601 20000 35
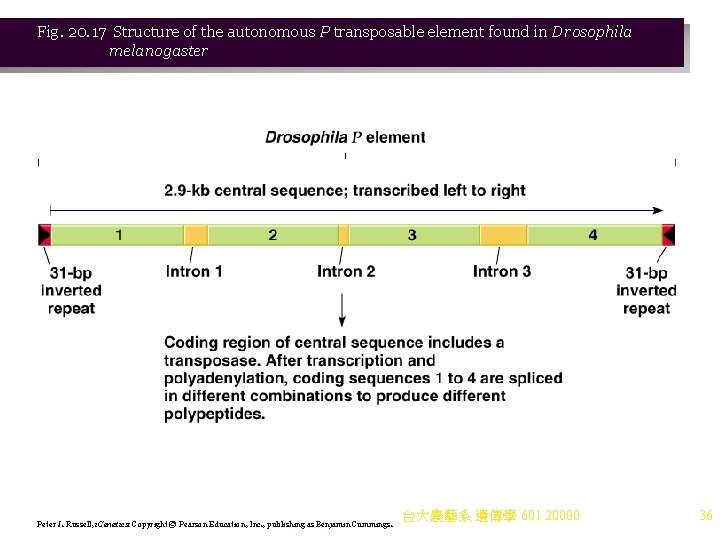
Fig. 20. 17 Structure of the autonomous P transposable element found in Drosophila melanogaster Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 36
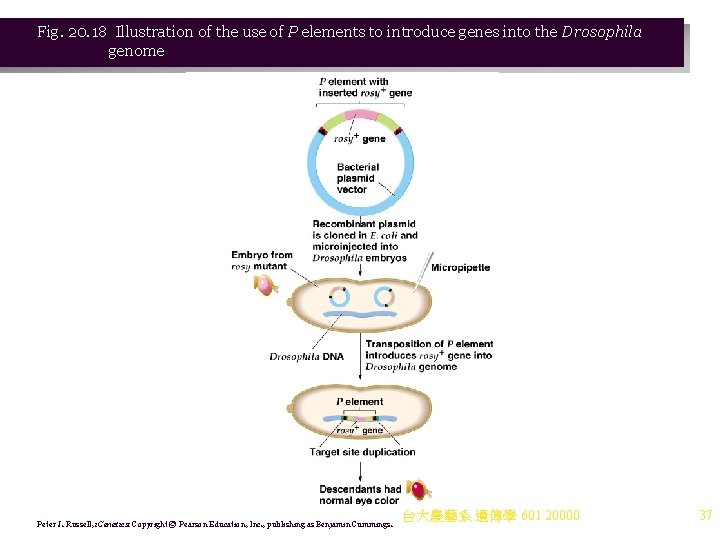
Fig. 20. 18 Illustration of the use of P elements to introduce genes into the Drosophila genome Peter J. Russell, i. Genetics: Copyright © Pearson Education, Inc. , publishing as Benjamin Cummings. 台大農藝系 遺傳學 601 20000 37

Human Retrotansposons 1. Retrotransposons also appear to be present in mammals. For example, a very abundant human SINE repeat (short interspersed sequence) is the Mu family, named for the Alu. I restriction site in its sequence. a. Mu sequences are about 300 bp, repeated 300, 000 -500, 000 times in the human genome (up to 3% of total human DNA). b. Sequences are divergent, related but not identical. c. Each Mu sequence is flanked by 7 -20 bp direct repeats. d. At least a few Mu sequences can be transcribed, and the model is that transcriptionally active Mu sequences are retrotransposons that move via an RNA intermediate. e. A human case of a genetic disease, neurofibromatosis, provides some evidence. i. Neurofibromas (tumorlike growths on the body) result from an autosomal dominant mutation. ii. In a patient’s DNA, an unusual Mu sequence was detected in one of the introns of the neurofibromatosis gene. iii. The resulting longer transcript is incorrectly proessed, removing an exon from the m. RNA and producing a nonfunctional protein. iv. Neither parent had this Mu sequence in the neurofibromatosis gene. v. Divergent Mu sequences made it possible to track this particular version to an insertion event in the germ line of the patient’s father. f. It is not clear how the functions needed for Mu retrotransposition are provided. 台大農藝系 遺傳學 601 20000 38

2. A mammalian LINEs family, LINEs-i (Li elements) is also thought to be retrotransposons. a. Humans have 50, 000 -100, 000 copies of the Li element, comprising about 5% of the genome. b. The full-length element (6. 5 kb) is not abundant, and most Li elements are deleted versions. c. The full-length Li element contains a large ORF with homolegy to known reverse transcriptases. Experimentally, the Li ORF can substitute for the yeast Ty reverse transcriptase gene. d. Li elements are thought to be retrotransposons, but do not have LTRs. e. Clinically, cases of hemophilia have been shown to result from newly transposed Li insertions into the factor VIII gene. (Factor VIII is required for normal blood clotting. ) 台大農藝系 遺傳學 601 20000 39
- Slides: 39