Pep Ar ML A modelfree resultcombining peptide identification

Pep. Ar. ML: A model-free, resultcombining peptide identification arbiter via machine learning Xue Wu, Chau-Wen Tseng, Nathan Edwards University of Maryland, College Park, and Georgetown University Medical Center

Comparison of Search Engines SEQUEST Mascot • No single score is comprehensive 14% • Search engines disagree 28% 14% 38% • Many spectra lack confident peptide assignment 1% 3% 2% • Many spectra lack any peptide assignment X! Tandem 2 Searle et al. JPR 7(1), 2008

Black-box Techniques • Significance re-estimation • Target-Decoy search • Bimodal distribution fit • Supervised machine learning • Train predictors on synthetic datasets • Select and/or create (many) good features • Result combiners • Incorrect peptide IDs unlikely to match • Significance re-estimation • Independence and/or supervised model 3

Pep. Ar. ML • Unified machine learning result combiner • Significance re-estimation too! • Model-free feature use and result combination • Use agreement and features if useful • Unsupervised training procedure • No loss of classification performance 4
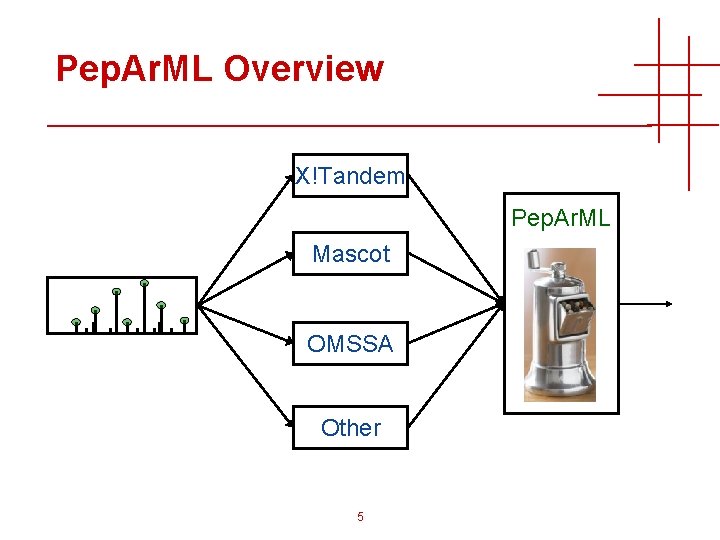
Pep. Ar. ML Overview X!Tandem Pep. Ar. ML Mascot OMSSA Other 5
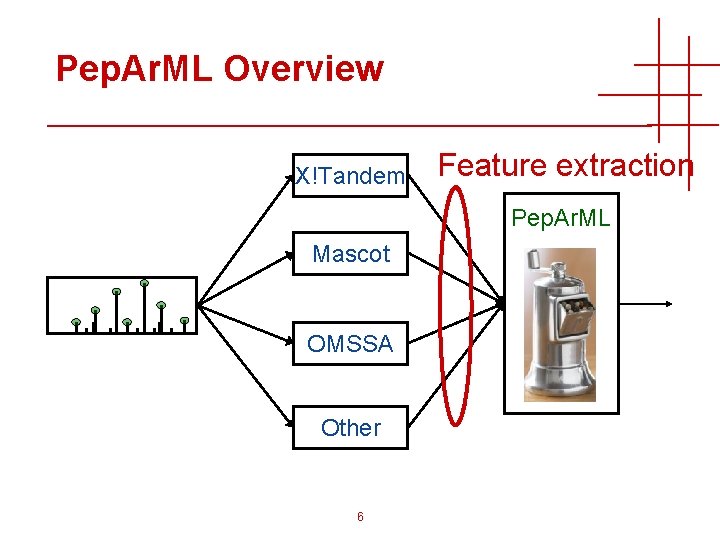
Pep. Ar. ML Overview X!Tandem Feature extraction Pep. Ar. ML Mascot OMSSA Other 6

Dataset Construction X!Tandem Mascot OMSSA T F T …… T 7

Dataset Construction • Calibrant 8 Protein Mix (C 8) • 4594 MS/MS spectra (LTQ) • 618 (11. 2%) true positives • Sashimi 17 mix_test 2 (S 17) • 1389 MS/MS spectra (Q-TOF) • 354 (25. 4%) true positives • AURUM 1. 0 (364 Proteins) • 7508 MS/MS spectra (MALDI-TOF) • 3775 (50. 3%) true positives 8

Pep. Ar. ML Machine Learning • Machine learning (generally) helps single search engines • Pep. Ar. ML result-combiner (C-TMO) improves on single search engines • Sometimes combining two search engines works as well, or better, than three 9
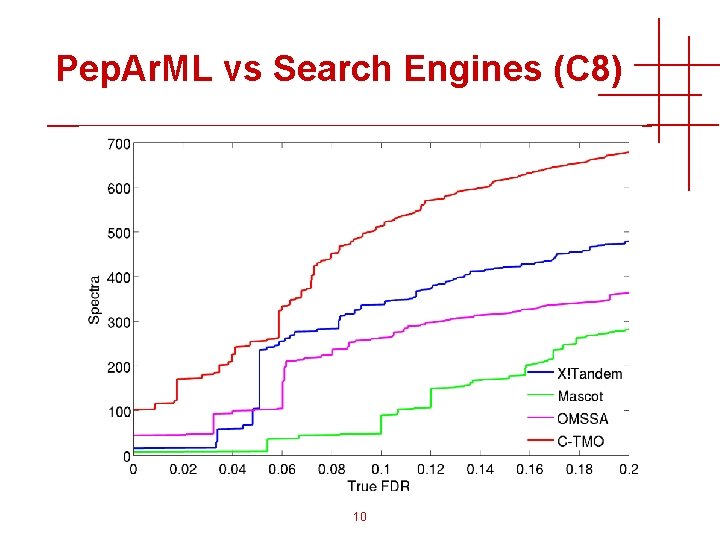
Pep. Ar. ML vs Search Engines (C 8) 10
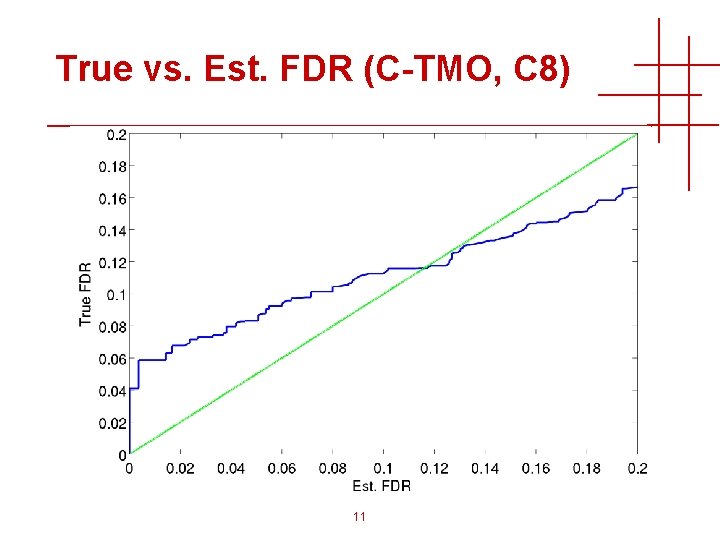
True vs. Est. FDR (C-TMO, C 8) 11
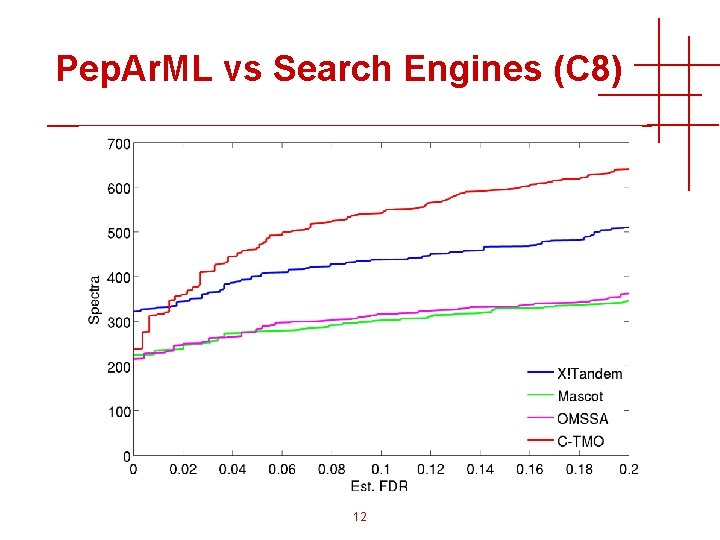
Pep. Ar. ML vs Search Engines (C 8) 12
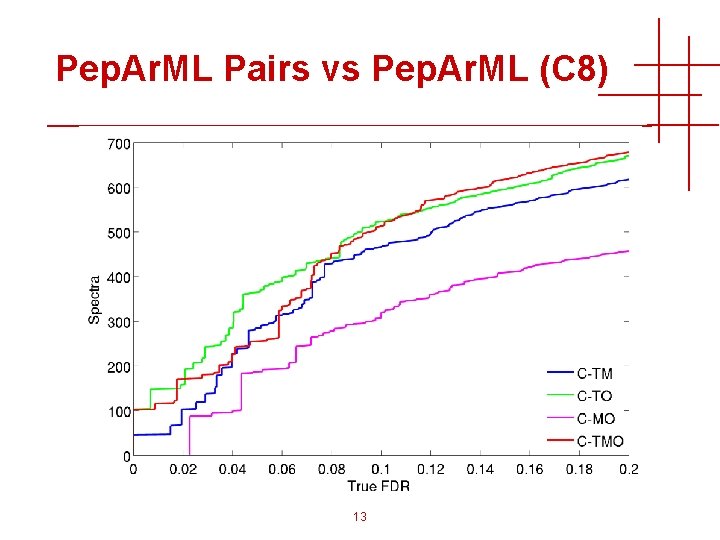
Pep. Ar. ML Pairs vs Pep. Ar. ML (C 8) 13
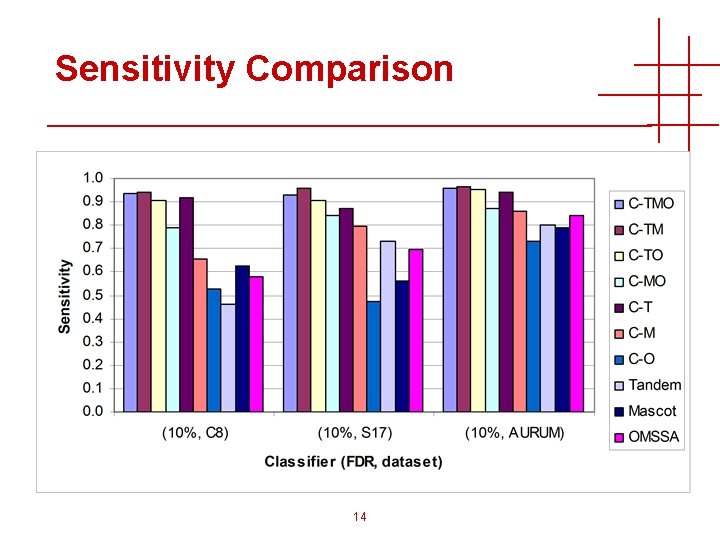
Sensitivity Comparison 14
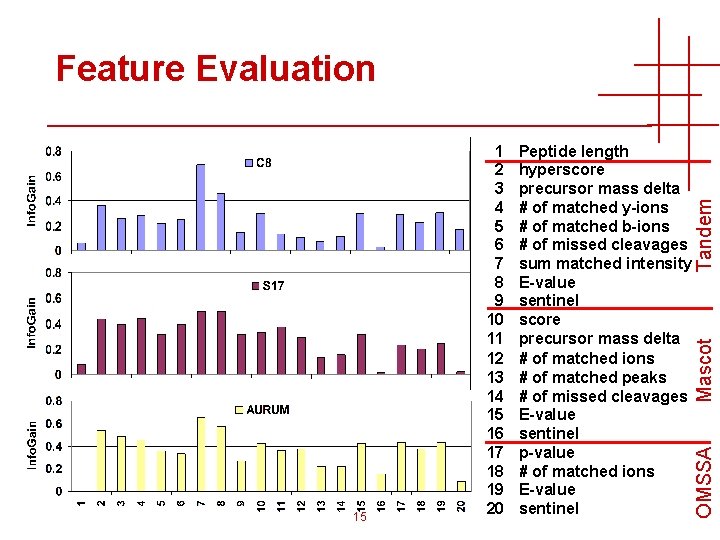
Mascot Peptide length hyperscore precursor mass delta # of matched y-ions # of matched b-ions # of missed cleavages sum matched intensity E-value sentinel score precursor mass delta # of matched ions # of matched peaks # of missed cleavages E-value sentinel p-value # of matched ions E-value sentinel OMSSA 15 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 Tandem Feature Evaluation

Application to Real Data • How well do these models generalize? • Different instruments • Spectral characteristics change scores • Search parameters • Different parameters change score values • Supervised learning requires • (Synthetic) experimental data from every instrument • Search results from available search engines • Training/models for all parameters x search engine sets x instruments 16
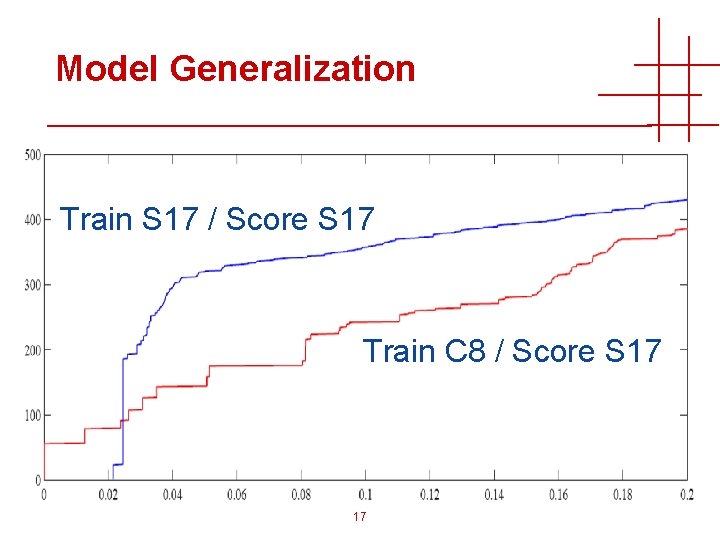
Model Generalization Train S 17 / Score S 17 Train C 8 / Score S 17 17

Rescuing Machine Learning • Train a new machine learning model for every dataset! • Generalization not required • No predetermined search engines, parameters, instruments, features • Perhaps we can “guess” the true proteins • Most proteins not in doubt • Machine learning can tolerate imperfect labels 18
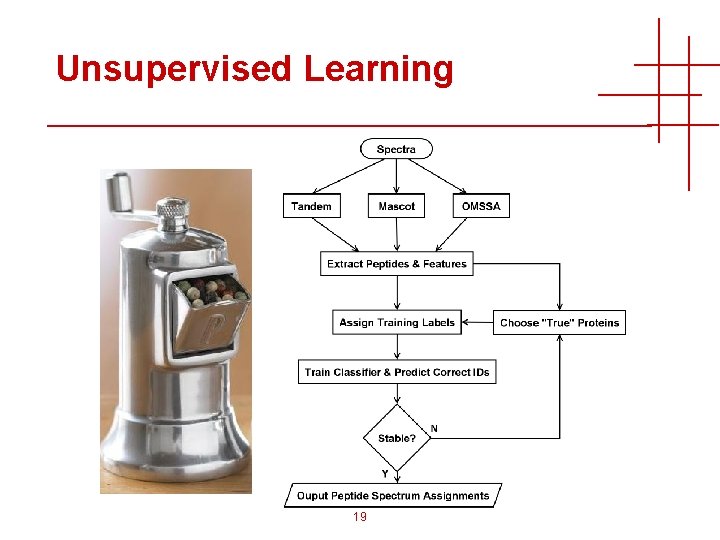
Unsupervised Learning 19
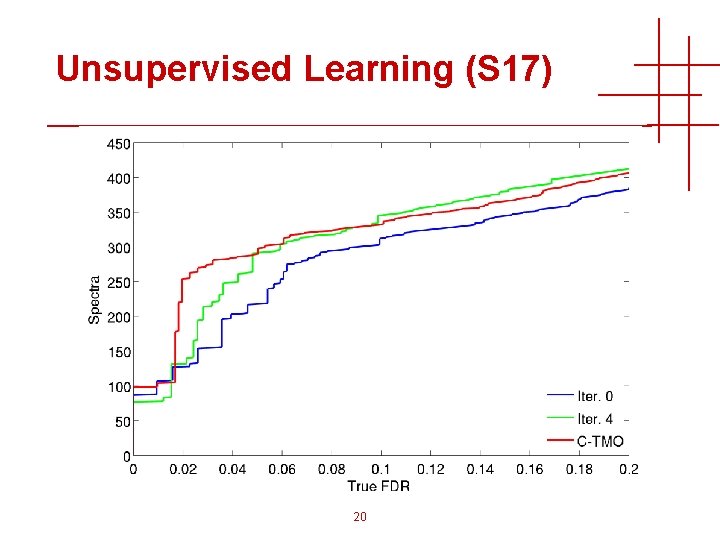
Unsupervised Learning (S 17) 20
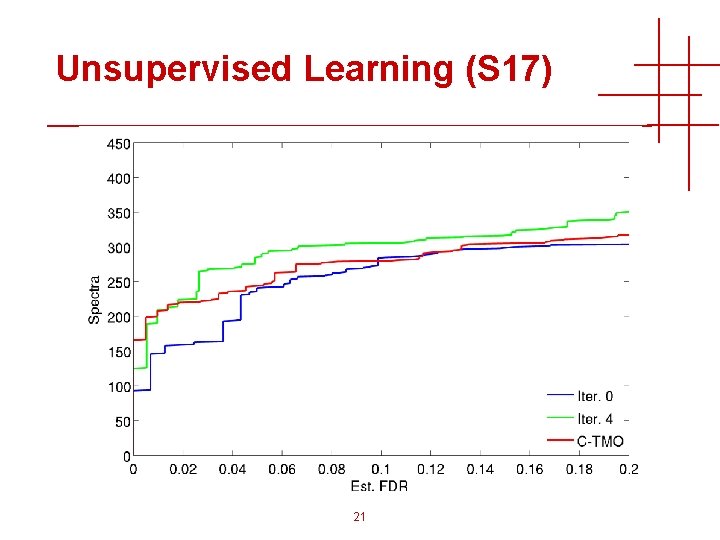
Unsupervised Learning (S 17) 21

Protein Selection Heuristic • Modeled on typical protein identification criteria • High confidence peptide IDs • At least 2 non-overlapping peptides • At least 10% sequence coverage • Robust, fast convergence • Easily enforce additional constraints 22

What about real data? Dr. Rado Goldman (LCCC, GUMC) • Proteolytic serum peptides from clinical hepatocellular carcinoma samples • ~ 200 MALDI MS/MS Spectra (TOF-TOF) Pep. Ar. ML for non-specific search of IPI-Human • Increase in confidence & sensitivity • Observation of “ragged” proteolytic trimming 23
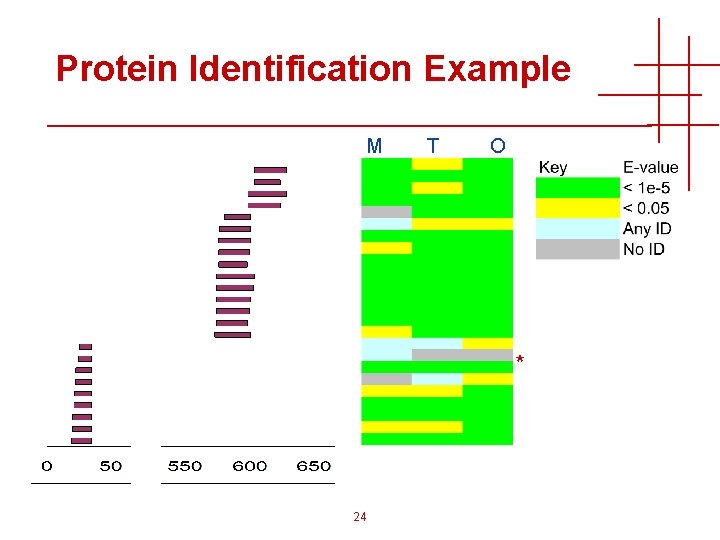
Protein Identification Example M T O * 24

Future Directions • Apply to more experimental datasets • Integrate • novel features • new search engines, spectral matching • multiple searches with varied parameters, sequence databases • Construct meta-search engine • FDR by bimodal fit instead of decoys • Release as open source • http: //peparml. sourceforge. org 25
http: //Pep. Ar. ML. Source. Forge. Net 26

Acknowledgements • Xue Wu* & Dr. Chau-Wen Tseng, • Computer Science University of Maryland, College Park • Dr. Brian Balgley, Dr. Paul Rudnick • Calibrant Biosystems & NIST • Dr. Rado Goldman, Dr. Yanming An • Department of Oncology Georgetown University Medical Center • Kam Ho To • Biochemistry Masters student Georgetown University • Funding: NIH/NCI CPTAC 27

28
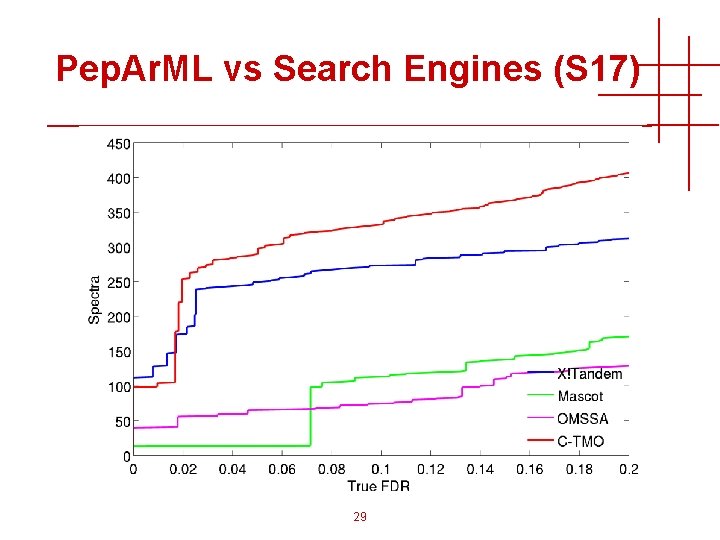
Pep. Ar. ML vs Search Engines (S 17) 29
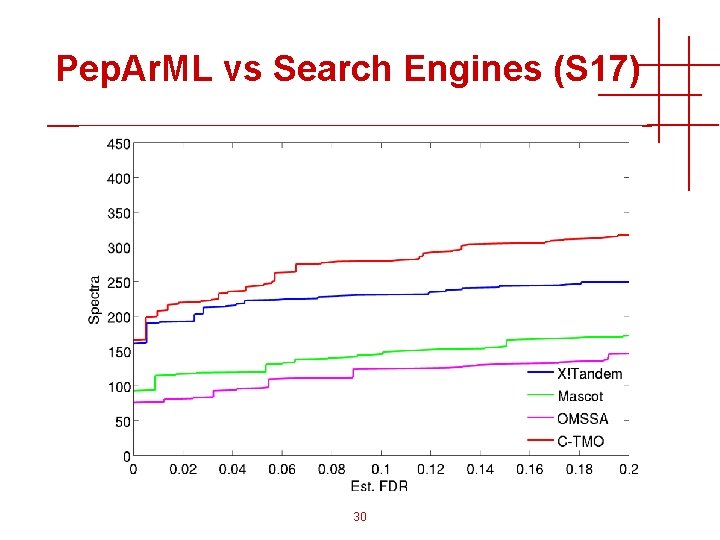
Pep. Ar. ML vs Search Engines (S 17) 30
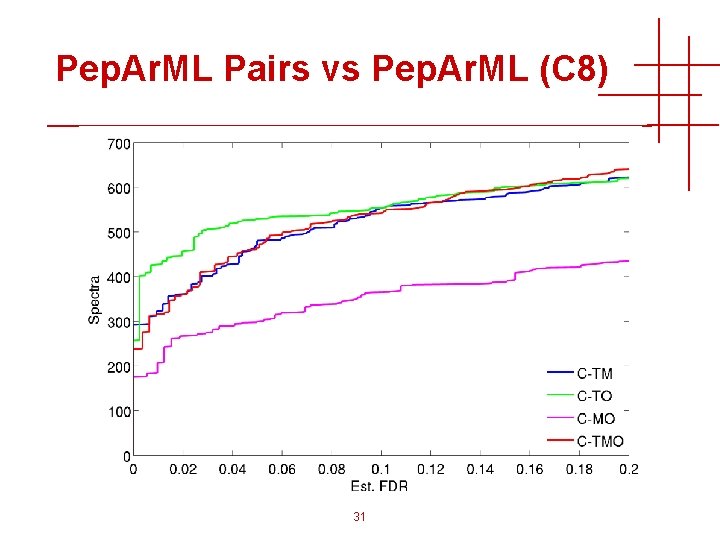
Pep. Ar. ML Pairs vs Pep. Ar. ML (C 8) 31
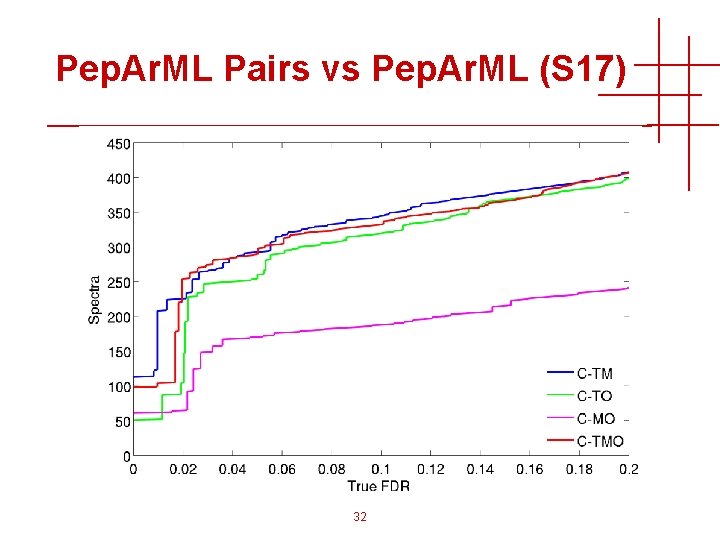
Pep. Ar. ML Pairs vs Pep. Ar. ML (S 17) 32
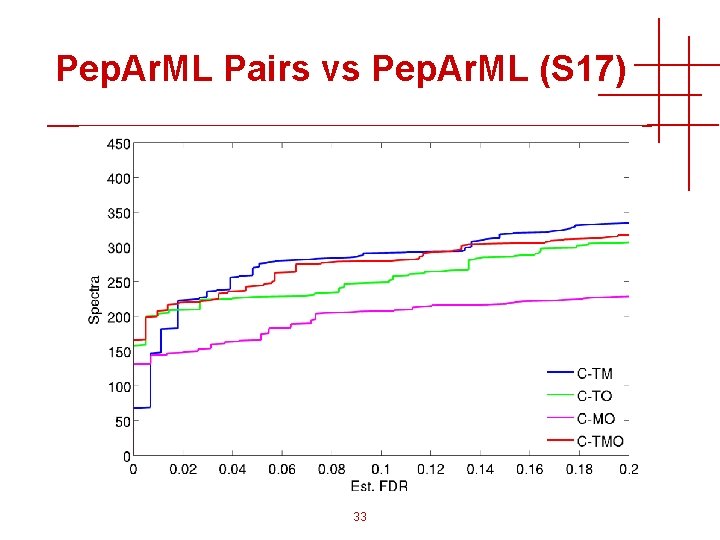
Pep. Ar. ML Pairs vs Pep. Ar. ML (S 17) 33
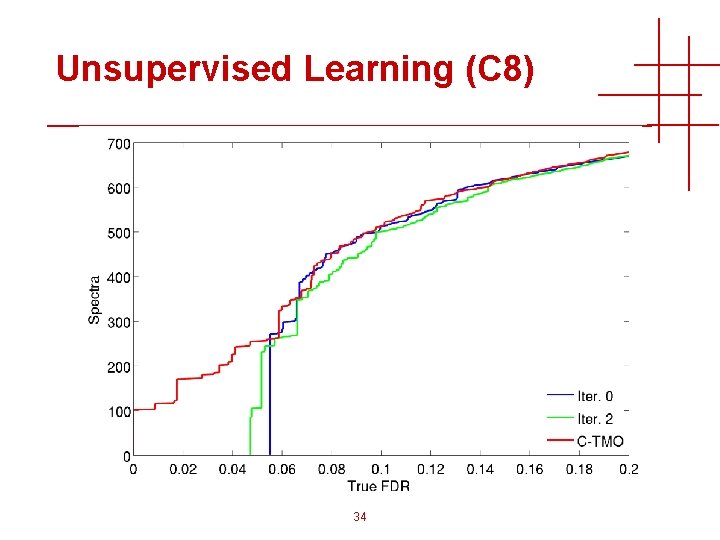
Unsupervised Learning (C 8) 34
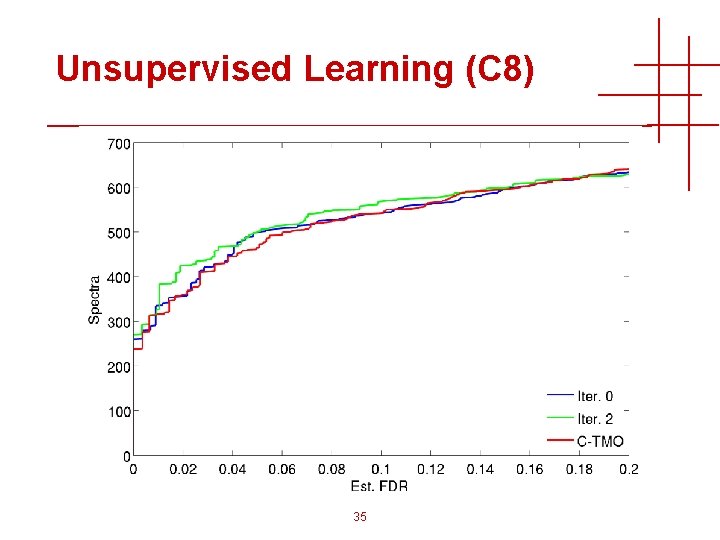
Unsupervised Learning (C 8) 35
- Slides: 35