PCR assay of the molecular alternations at cinnabar

PCR assay of the molecular alternations at cinnabar and vestigial genes of Drosophila melanogaster after γ-rays and neutron action • Aleksievich Olga • Sheresh Irina • Lebogang Sepini • Igor D. Alexandrov, Ph. D. , Dr. Sci. (Biology), chief. Sci. res. • Margarita V. Alexandrova, Ph. D. , senior sci. res.

GOAL OF PROJECT TO STUDY THE MOLECULAR GENETIC ACTION OF GAMMARAYS AND NEUTRONS ON THE CINNABAR AND VESTIGIAL GENES IN DROSOPHILA MELANOGASTER GERM CELLS

Potential genetic risks Nuclear explosion Technogenic catastrophes Sun radiation

Why Drosophila? • Well studied example, gene structure known • Has common principal DNA structure with humans • Short life cycle (~15 days) • Permits the study of heritable gene mutation • Low cost
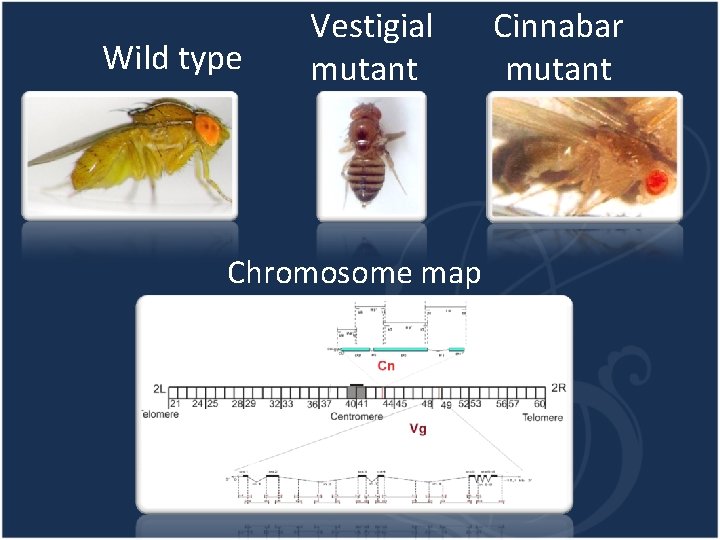
Wild type Vestigial mutant Chromosome map Cinnabar mutant
Main steps • Isolation of DNA • Polymerase chain reaction • Electrophoresis

Isolation of DNAs were isolated from imago of wild type (as control) and cinnabar and vestigial mutants using procedure below: • Cell lysis • DNA absorption from the nucleus surface with silica solution (Nucleo. S™). • Washing DNA with solution buffer • DNA extraction from silica solution with Extra. Gene™.
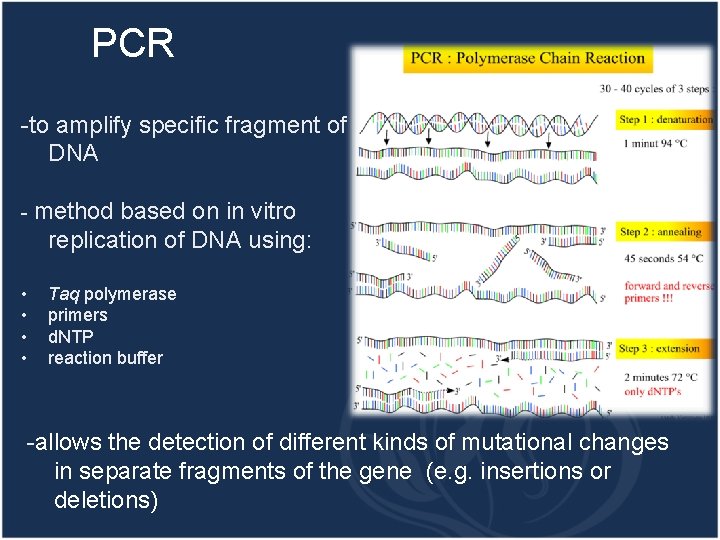
PCR -to amplify specific fragment of DNA - method based on in vitro replication of DNA using: • • Taq polymerase primers d. NTP reaction buffer -allows the detection of different kinds of mutational changes in separate fragments of the gene (e. g. insertions or deletions)

Amplificator
Gel electrophoresis
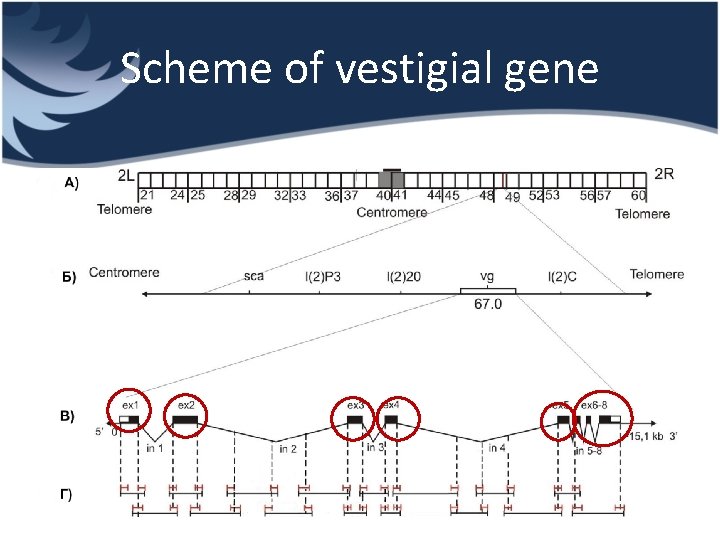
Scheme of vestigial gene
Ex 6 -8 Ex 3 Ex 5 Ex 4 Ex 1 Ex 2
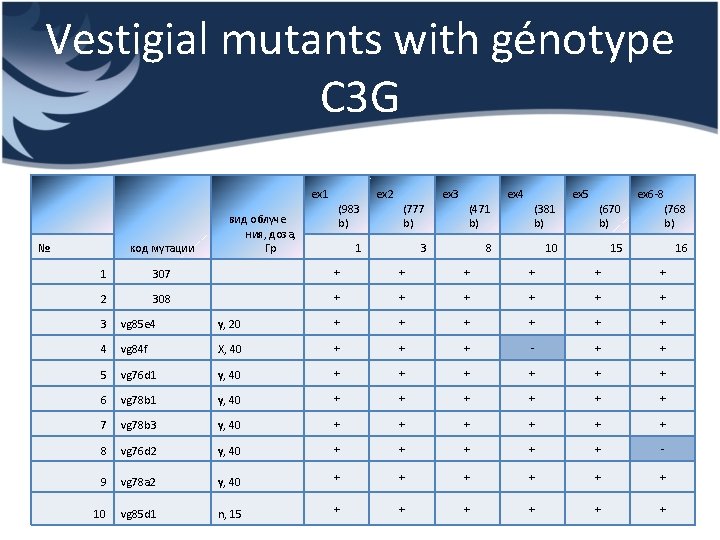
Vestigial mutants with génotype C 3 G ex 1 № код мутации вид облуче ния, доза, Гр (983 b) ex 2 (777 b) 1 ex 3 (471 b) 3 ex 4 (381 b) 8 ex 5 (670 b) ex 6 -8 (768 b) 15 16 10 1 307 + + + 2 308 + + + 3 vg 85 e 4 γ, 20 + + + 4 vg 84 f X, 40 + + + - + + 5 vg 76 d 1 γ, 40 + + + 6 vg 78 b 1 γ, 40 + + + 7 vg 78 b 3 γ, 40 + + + 8 vg 76 d 2 γ, 40 + + + - 9 vg 78 a 2 γ, 40 + + + 10 vg 85 d 1 n, 15 + + +

Results • 2 control lines and 8 mutants (γ-irradiated) were examined • 6 fragments of vestigial gene were analyzed • More than 60 polymerase chain reactions were carried out • As you can see from the results of PCR only 2 mutants have one deletion of different fragments each

Conclusions • Such a little quantity of PCR-detected deletions could be caused by specific action of γ-rays. Because the type of radiation mentioned above induce point damage (as a rule) that cannot be detected by PCR • In our research work only exones were examined. But there is a probability to discover deletions in intrones. So it can be the next step in analysis of this mutants
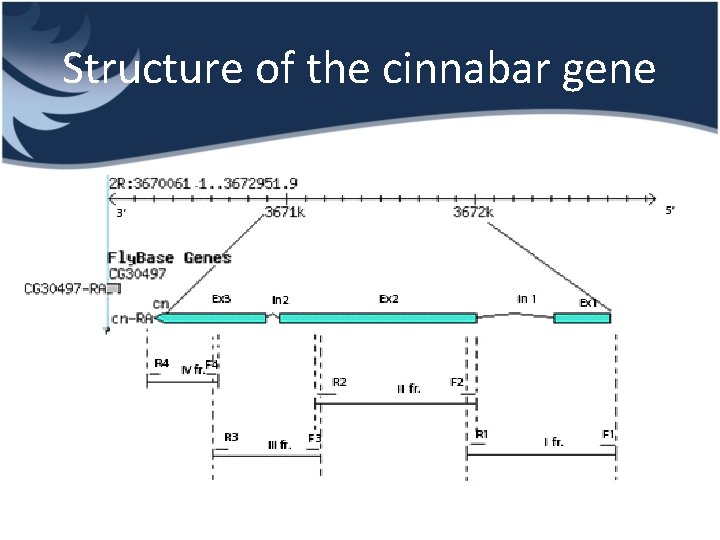
Structure of the cinnabar gene

Visualization of Electrophoresis Using UV Light
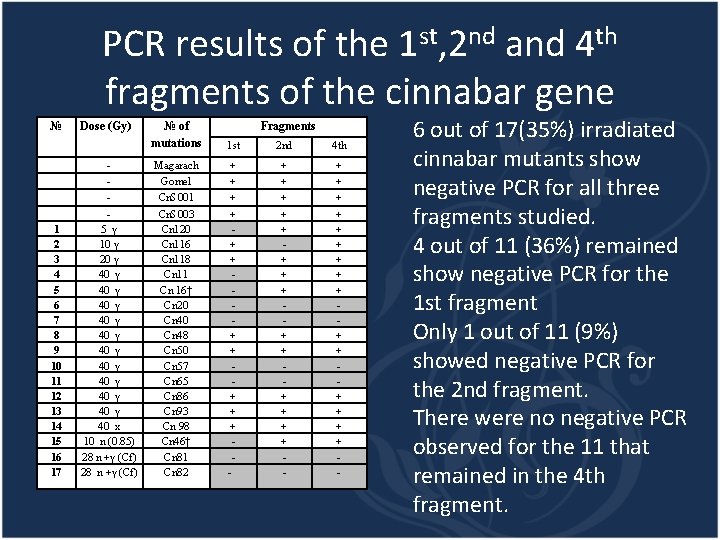
PCR results of the 1 st, 2 nd and 4 th fragments of the cinnabar gene № Dose (Gy) - 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 5 γ 10 γ 20 γ 40 γ 40 γ 40 x 10 n (0. 85) 28 n +γ (Cf) № of mutations Fragments 1 st 2 nd 4 th Magarach Gomel Cn. S 001 Cn. S 003 Cn 120 Cn 116 Cn 118 Cn 11 Cn 16† Cn 20 Cn 48 Cn 50 Cn 57 Cn 65 Cn 86 Cn 93 Cn 98 Cn 46† Cn 81 Cn 82 + + + + + + - + + + + + - 6 out of 17(35%) irradiated cinnabar mutants show negative PCR for all three fragments studied. 4 out of 11 (36%) remained show negative PCR for the 1 st fragment Only 1 out of 11 (9%) showed negative PCR for the 2 nd fragment. There were no negative PCR observed for the 11 that remained in the 4 th fragment.

Conclusions High frequency of the loss of the entire gene could be determined by the size and position of the cinnabar gene on the chromosome The beginning of the gene could be a hot spot for the irradiation for both gamma- and neutron radiation

General Conclusions • We have studied the molecular alterations induced by ionizing radiation at 2 Drosophila genes with different sizes, structure and position on the chromosomes. • Two different genes-targets react in different ways on the action of radiation. They have different picture of radiomutability. • It is suggested that this differences could be connected with different position of 2 genes on the chromosome.

Thanks for your attention!
- Slides: 21