PathwayGenome Databases and Software Tools Peter D Karp

Pathway/Genome Databases and Software Tools Peter D. Karp, Ph. D. Bioinformatics Research Group SRI International pkarp@ai. sri. com http: //ecocyc. Double. Twist. com/ecocyc/

SRI International Bioinformatics Overview l Overview of bioinformatics l Motivations for the Eco. Cyc project l Eco. Cyc demo l Description of Eco. Cyc database and Pathway Tools software l Underlying technologies l Ocelot object database l GKB Editor l X-windows to WWW translator

Definition of Bioinformatics l Computational SRI International Bioinformatics techniques for management and analysis of biological data and knowledge l Methods for disseminating, archiving, interpreting, and mining scientific information

SRI International Bioinformatics Motivations for Bioinformatics l Growth in molecular-biology knowledge l Industrialization l High-throughput of biological experimentation biology l Genome sequences l Gene and protein expression data l Protein-protein interaction data l Protein 3 -D structures l ….
SRI International Bioinformatics A E


Motivations for Eco. Cyc -E. coli Encyclopedia l Integrate SRI International Bioinformatics E. coli information dispersed in the literature l New paradigm of scientific publishing l Model the full metabolic network of an organism l Integrate l Develop l Provide genomic data with functional data algorithms for computing with function a challenging domain for computerscience research

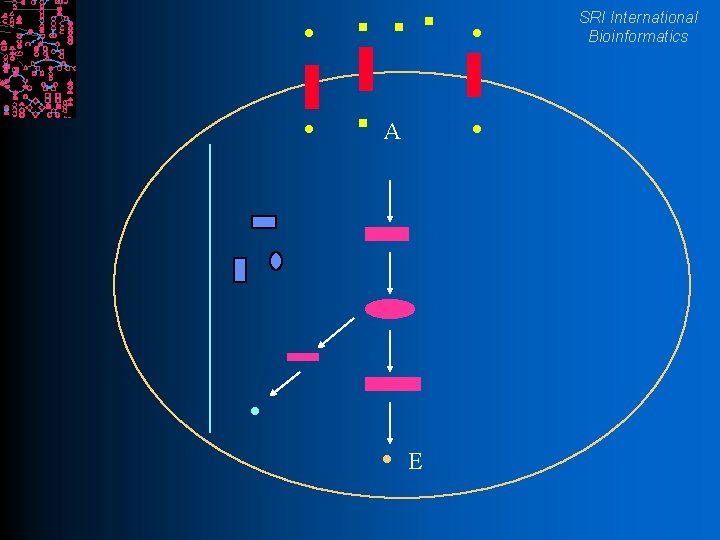
SRI International Bioinformatics Definitions l. A chemical reaction interconverts chemical compounds A+B=C+D l An enzyme is a protein that accelerates chemical reactions l. A pathway is a linked set of reactions A l. A C E conceptual unit of cell’s biochemical machine

Organism-Specific Pathway/Genome Databases l Layer SRI International Bioinformatics functional information above the genome l Rich ontology to encode biological information with high fidelity l Chromosomes, genes, operons, gene products, reactions, pathways l Curated by experts for that organism l Integrate literature and computational predictions

Pathway Tools Software SRI International Bioinformatics l Pathway/Genome Navigator l WWW publishing of PGDBs l Graphic depictions of pathways, chromosomes, operons l Pathway visualization of gene-expression data l Pathway/Genome Editors l Distributed curation of genome annotations l Distributed object database system l Interactive editing tools l Patho. Logic

Eco. Cyc = E. coli Dataset + Pathway/Genome Navigator SRI International Bioinformatics Metabolic Network Pathways: 158 Reactions: 1, 117 Compounds: 1, 887 Gene Products: 4, 393 Genes: 4, 393 Operons: 375 http: //ecocyc. Double. Twist. com/ecocyc/

Eco. Cyc l Collaborative SRI International Bioinformatics development via internet l Karp -- Bioinformatics architect l Riley -- Metabolic pathways, signal transduction l Saier and Paulsen -- Transport l Collado -- Regulation of gene expression l Ontology of 1000 biological classes l 14, 000 instances l Over 2, 600 registered users
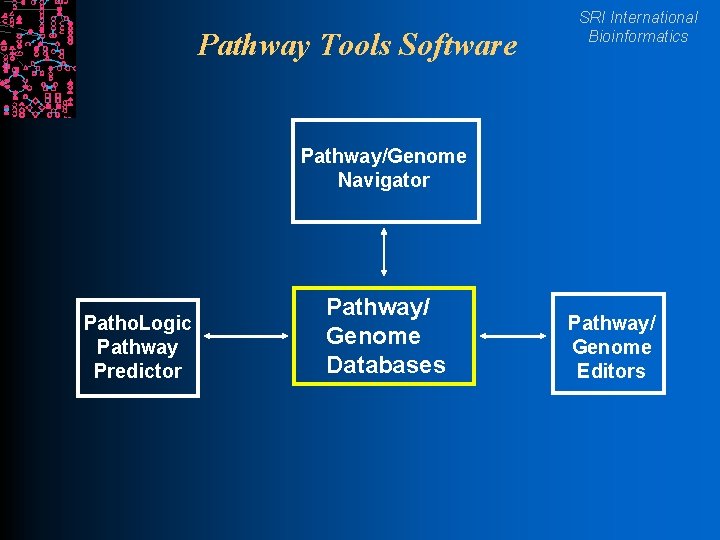
Pathway Tools Software SRI International Bioinformatics Pathway/Genome Navigator Patho. Logic Pathway Predictor Pathway/ Genome Databases Pathway/ Genome Editors




Creation of the Overview Graph SRI International Bioinformatics l Run layout algorithms on individual pathway graphs l Automatically determine topology of pathway graph l Apply associated layout algorithm (linear, circular, tidy tree) l Use superpathways to create hierarchical layouts l Treat each individual pathway as a single node l Pathway connections are edges l Run appropriate layout algorithm
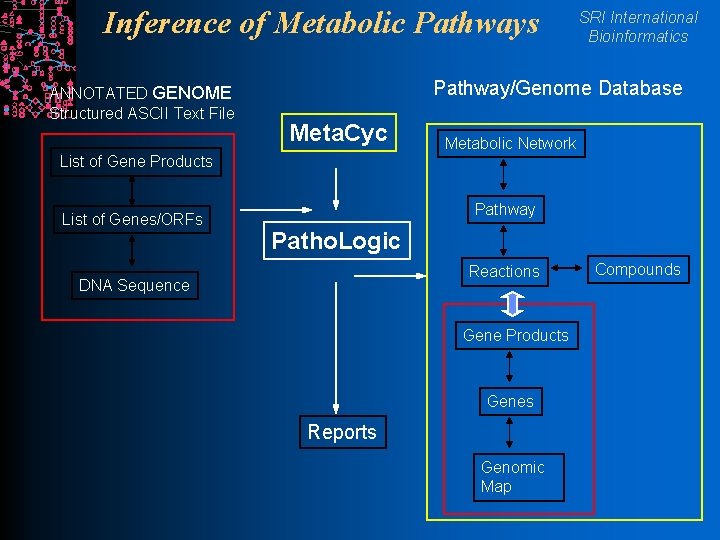
Inference of Metabolic Pathways ANNOTATED GENOME Structured ASCII Text File SRI International Bioinformatics Pathway/Genome Database Meta. Cyc Metabolic Network List of Gene Products List of Genes/ORFs Pathway Patho. Logic Reactions DNA Sequence Gene Products Genes Reports Genomic Map Compounds

Summary of H. pylori Analysis SRI International Bioinformatics l For 121 E. coli pathways, what is the evidence that each pathway occurs in H. pylori? l Strong evidence: 41 l Medium evidence: 29 l Little or no evidence: 51 l 31 reactions catalyzed by H. pylori but not by E. coli l H. pylori has partial abilities to synthesize cofactors and amino-acids, extremely limited carbohydrate catabolism, some amino acid utilization, and a reductive citric-acid pathway

SRI International Bioinformatics Microbial Pathway/ Genome DBs Literature-based Datasets: Patho. Logic-based Datasets: l. Meta. Cyc l. Bacillus l. Escherichia coli subtilis l. Mycobacterium tuberculosis l. Helicobacter pylori l. Haemophilus influenzae l. Mycoplasma pneumonia l. Treponema pallidum l. Chlamydia trachomatis l. Saccharomyces cerevisiae

SRI International Bioinformatics Pathway Tools Software Architecture l Implemented in Common Lisp l WWW server runs as a single Unix process with a separate thread to service each query l Grasper-CL l Ocelot graph manager object database l GKB Editor schema-driven editor
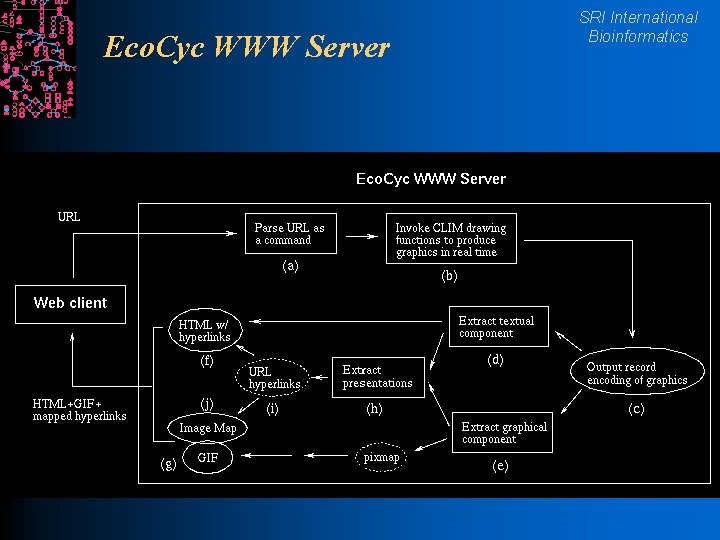
Eco. Cyc WWW Server SRI International Bioinformatics
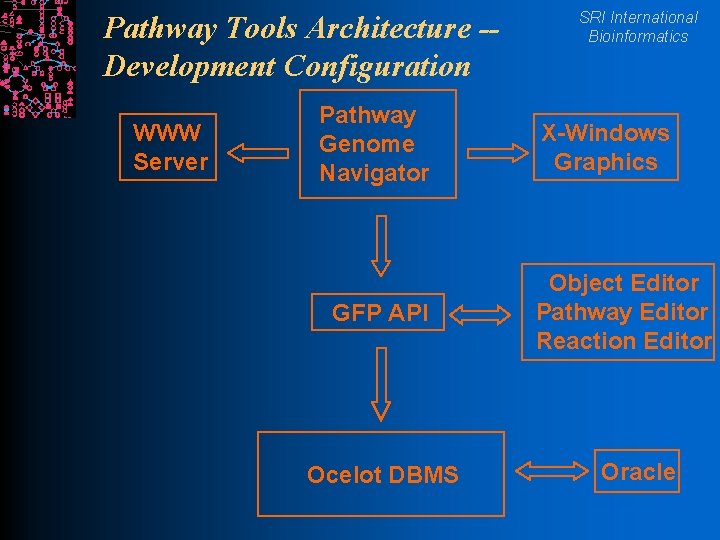
Pathway Tools Architecture -Development Configuration WWW Server Pathway Genome Navigator GFP API Ocelot DBMS SRI International Bioinformatics X-Windows Graphics Object Editor Pathway Editor Reaction Editor Oracle

Ocelot Database System SRI International Bioinformatics l Object Database Manager l Persistence via filesystem or relational DBMS l Demand background faulting of objects from RDBMS l Two-level object caching l Extensive bioinformatics schema l Stored transaction history l Inspect object history

Ocelot Knowledge Server Architecture l Frame SRI International Bioinformatics data model l Persistent storage via l Disk files l Oracle DBMS l Optimistic concurrency-control protocol l Schema evolution l Logging facility

The Frame Data Model l Frames SRI International Bioinformatics are of two types: classes, instances l Frames have slots that define their properties, attributes, relationships l. A slot has one or more values l Each value can be any Lisp datatype l Slotunits define metadata about slots: l Domain, range, inverse l Collection type, number of values, value

Inference Capabilities l Inheritance l Slot SRI International Bioinformatics of defaults values computed via attached procedures l Maintenance l Constraint of inverse relationships system l Deferred evaluation l Tolerant of nonconformant data

Storage System Architecture l Oracle SRI International Bioinformatics KBs l DBMS is submerged within FRS l Relational schema is domain independent, supports multiple KBs simultaneously l Frames transferred from DBMS to Ocelot l On demand l By background prefetcher l Memory cache l Persistent disk cache to speed performance via Internet

Frame Faulting SRI International Bioinformatics (get-slot-value gene ‘map-position) l Gene present in in-memory object cache? l Gene present in cache on local disk? l Query Oracle DBMS

Logging l Oracle SRI International Bioinformatics DBMS stores: l The latest version of each frame l A history of all OKBC operations applied to KB l Reconstruct earlier versions of KB l View history of changes to an object l Update replicates l Concurrency control

Schema Management SRI International Bioinformatics l FRSs store and process class and instance information similarly l Applications can query schema information as easily as they can query instances

GKB Editor l Browser l Four SRI International Bioinformatics and editor for KBs and ontologies editing tools l GKB Editor reusable with multiple FRSs l All database queries via OKBC/GFP API l Interoperability achieved with Ocelot, LOOM, Ontolingua l All operations are schema driven l http: //www. ai. sri. com/~gkb/overview. html

SRI International Bioinformatics Editors l Taxonomy l Frame editor l Relationships l Spreadsheet editor



SRI International Bioinformatics Results l Ocelot in use in the Eco. Cyc project for 5 years l Supports collaborative development of Eco. Cyc by four groups in North America l Distributed architecture l GKB Editor in active use l Supports development of 8 Pathway/Genome Databases

SRI International Bioinformatics Summary l Pathway/Genome l Pathway Databases Tools software l Extract pathways from genomes l Distributed curation tools l Query, visualization, WWW publishing l Analysis algorithms

Computer Science Results SRI International Bioinformatics l Extend scalability and multiuser access for knowledge representation systems l Reusable, schema-driven KB editor l Hierarchical graph layout algorithms l Dynamic translation from X-windows to HTML+GIF l Importance of ontologies and of content: l Discovery = Algorithm + Database
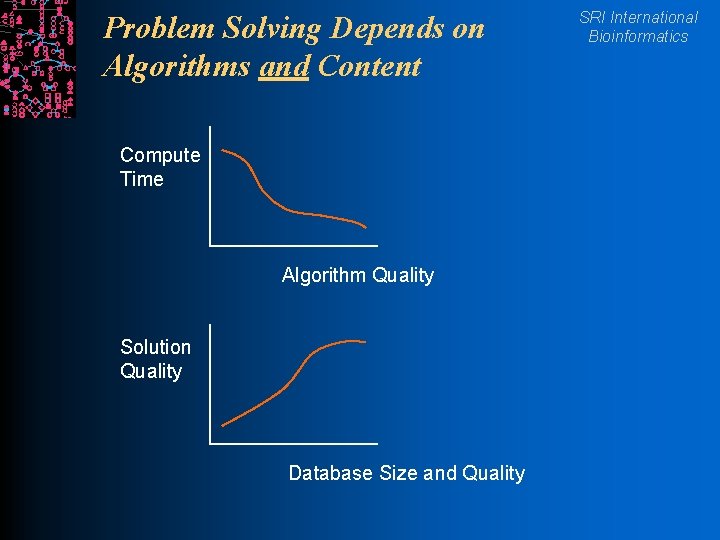
Problem Solving Depends on Algorithms and Content Compute Time Algorithm Quality Solution Quality Database Size and Quality SRI International Bioinformatics

Bioinformatics Results: Content SRI International Bioinformatics l The Eco. Cyc database describes the full metabolic map of an organism l The Meta. Cyc database describes over 300 metabolic pathways l Ontology spans genome to pathway information

Bioinformatics Results: Algorithms SRI International Bioinformatics l Software environment for genome and pathway information l Query and visualization l Distributed database development l Patho. Logic algorithm predicts the metabolic network of an organism from its genome l Algorithms under development for qualitative modeling of the cell

Acknowledgements SRI International Bioinformatics l Funding sources: l NIH National Center for Research Resources l Collaborators: Monica Riley, Marine Biological Laboratory l Milton Saier, UC San Diego l Julio Collado, UNAM l Christos Ouzounis, European Bioinformatics Institute l Peter D. Karp, Ph. D. http: //www. ai. sri. com/pkarp/
- Slides: 42