Pathway Tools Bio Cyc Fundamentals Peter D Karp

Pathway Tools / Bio. Cyc Fundamentals Peter D. Karp, Ph. D. Bioinformatics Research Group SRI International pkarp@ai. sri. com Bio. Cyc. org Eco. Cyc. org, Meta. Cyc. org, Human. Cyc. org 1 SRI International Bioinformatics

Pathway Tools Capabilities l Create and maintain an organism database integrating genome, pathway, regulatory information l Computational inference tools l Interactive editing tools l Query and visualize that database l Use the database to interpret omics data l Metabolic network analysis tools l Comparative analysis tools l Export the metabolic network to SBML l Speed creation of flux-balance models by order of magnitude 2 SRI International Bioinformatics

Bio. Cyc l Hundreds of microbial genomes l Inferred operons and metabolic networks l Couples curated data with computational predictions l Supports analysis of omics data l Comparative analysis tools l Microbial emphasis. Exceptions: l Human. Cyc, Mouse. Cyc, Cattle. Cyc 3 SRI International Bioinformatics

Model Organism Databases / Organism Specific Databases 4 l DBs that describe the genome and other information about an organism l Every sequenced organism with an active experimental community requires a MOD l Integrate genome data with information about the biochemical and genetic network of the organism l Integrate literature-based information with computational predictions l Curated by experts for that organism l No one group can curate all the world’s genomes l Distribute workload across a community of experts to create a community resource SRI International Bioinformatics

Rationale for MODs 5 l Each “complete” genome is incomplete in several respects: l 40%-60% of genes have no assigned function l Roughly 7% of those assigned functions are incorrect l Many assigned functions are non-specific l Need continuous updating of annotations with respect to new experimental data and computational predictions l MODs are platforms for global analyses of an organism l Interpret omics data in a pathway context l In silico prediction of essential genes l Characterize systems properties of metabolic and genetic networks SRI International Bioinformatics

What is Curation? l l l l l 6 Ongoing updating and refinement of a PGDB Correcting false-positive and false-negative predictions Incorporating information from experimental literature Authoring of comments and citations Updating database fields Gene positions, names, synonyms Protein functions, activators, inhibitors Addition of new pathways, modification of existing pathways Defining TF binding sites, promoters, regulation of transcription initiation and other processes SRI International Bioinformatics
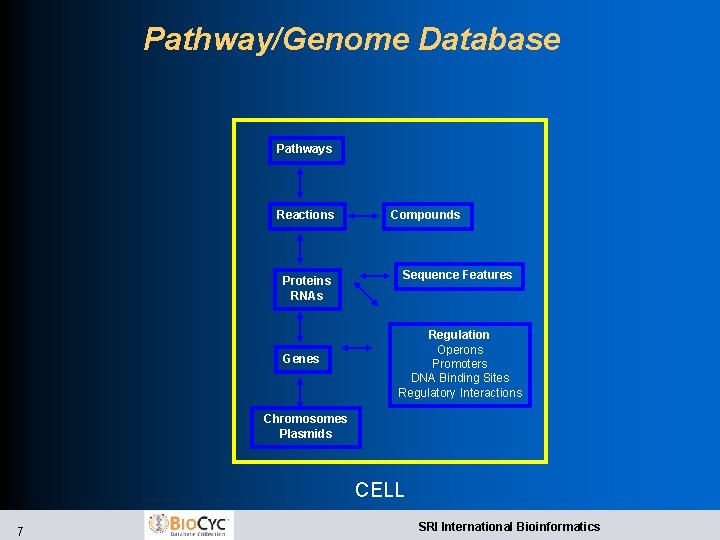
Pathway/Genome Database Pathways Reactions Proteins RNAs Genes Compounds Sequence Features Regulation Operons Promoters DNA Binding Sites Regulatory Interactions Chromosomes Plasmids CELL 7 SRI International Bioinformatics

Bio. Cyc Collection of 507 Pathway/Genome Databases l. Pathway/Genome Database (PGDB) – combines information about l Pathways, reactions, substrates l Enzymes, transporters l Genes, replicons l Transcription factors/sites, promoters, operons l. Tier 1: Literature-Derived PGDBs l Meta. Cyc l Eco. Cyc -- Escherichia coli K-12 l. Tier 2: Computationally-derived DBs, Some Curation -- 24 PGDBs l Human. Cyc l Mycobacterium tuberculosis l. Tier 3: Computationally-derived DBs, No Curation -- 481 DBs 8 SRI International Bioinformatics
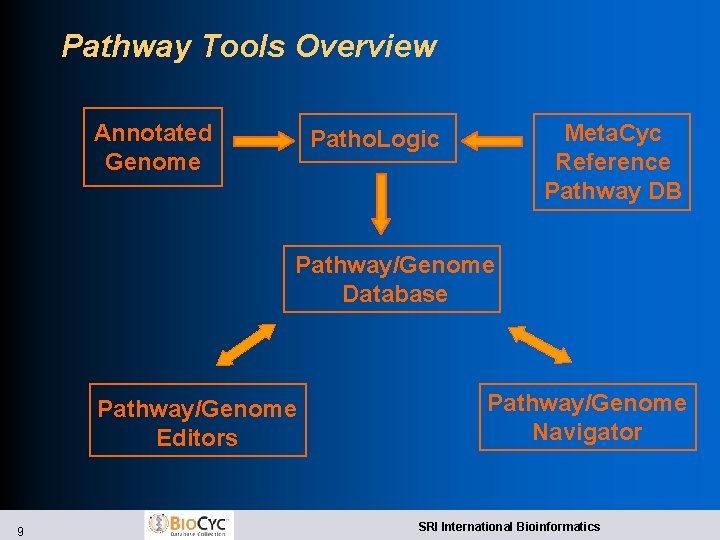
Pathway Tools Overview Annotated Genome Meta. Cyc Reference Pathway DB Patho. Logic Pathway/Genome Database Pathway/Genome Editors 9 Pathway/Genome Navigator SRI International Bioinformatics

Pathway Tools Software: Patho. Logic l Computational creation of new Pathway/Genome Databases l Transforms genome into Pathway Tools schema and layers inferred information above the genome l Predicts operons l Predicts metabolic network l Predicts which genes code for missing enzymes in metabolic pathways l Infers transport reactions from transporter names Bioinformatics 18: S 225 2002 10 SRI International Bioinformatics

Pathway Tools Software: Pathway/Genome Editors l Interactively update PGDBs with graphical editors l Support geographically distributed teams of curators with object database system l Gene editor Protein editor Reaction editor Compound editor Pathway editor Operon editor Publication editor l l l 11 SRI International Bioinformatics
Pathway Tools Software: Pathway/Genome Navigator l Querying and visualization of: l Pathways l Reactions l Metabolites l Proteins l Genes l Chromosomes l Two modes of operation: l Web mode l Desktop mode l Most functionality shared, but each has unique functionality 12 SRI International Bioinformatics

Pathway Tools Software: PGDBs Created Outside SRI l 1, 700+ licensees: 75+ groups applying software to 300+ organisms l. Saccharomyces cerevisiae, SGD project, Stanford University l 135 pathways / 565 publications l. Candida albicans, CGD project, Stanford University ldicty. Base, Northwestern University l. Mouse, MGD, Jackson Laboratory l. Under development: l Drosophila, Fly. Base l C. elegans, Worm. Base l. Arabidopsis thaliana, TAIR, Carnegie Institution of Washington l 288 pathways / 2282 publications l. Plant. Cyc, Carnegie Institution of Washington l. Six Solanaceae species, Cornell University l. Gramene. DB, Cold Spring Harbor Laboratory l. Medicago truncatula, Samuel Roberts Noble Foundation 13 SRI International Bioinformatics

Pathway Tools Software: PGDBs Created Outside SRI l. NIAID BRCs for Biodefense pathogens: l Bio. Health. Base -- Mycobacterium tuberculosis, Francisella tuleremia l Pathema -- 80+ PGDBs l PATRIC – Brucella suis, Coxiella burnetii, Rickettsia typhi l Eu. Path. DB – Cryptosporidium, Plasmodium l. G. Xie, Los Alamos Lab, Dental pathogens l. F. Brinkman, Simon Fraser Univ, Pseudomonas aeruginosa l. V. Schachter, Genoscope, Acinetobacter l. M. Bibb, John Innes Centre, Streptomyces coelicolor l. G. Church, Harvard, Prochlorococcus marinus, multiple strains l. E. Uberbacher, ORNL and G. Serres, MBL, Shewanella onedensis l. R. J. S. Baerends, University of Groningen, Lactococcus lactis IL 1403, Lactococcus lactis MG 1363, Streptococcus pneumoniae TIGR 4, Bacillus subtilis 168, Bacillus cereus ATCC 14579 l. Matthew Berriman, Sanger Centre, Trypanosoma brucei, Leishmania major l. Sergio Encarnacion, UNAM, Sinorhizobium meliloti l. Mark van der Giezen, University of London, Entamoeba histolytica, Giardia intestinalis l. Michael Gottfert, Technische Universitat Dresden, Bradyrhizobium japonicum l. Artiva Maria Goudel, Universidade Federal de Santa Catarina, Brazil, Chromobacterium violaceum ATCC 12472 14 SRI International Bioinformatics

Pathway Tools Software: PGDBs Created Outside SRI l Large scale users: l C. Medigue, Genoscope, 200+ PGDBs l G. Sutton, J. Craig Venter Institute, 80+ PGDBs l G. Burger, U Montreal, 60+ PGDBs l Bart Weimer, Utah State University, Lactococcus lactis, Brevibacterium linens, Lactobacillus acidophilus, Lactobacillus plantarum, Lactobacillus johnsonii, Listeria monocytogenes l Partial 15 listing of outside PGDBs at Bio. Cyc. org SRI International Bioinformatics

Obtaining a PGDB for Organism of Interest l Find existing curated PGDB l Find existing PGDB in Bio. Cyc l Create 16 your own SRI International Bioinformatics

Eco. Cyc Project – Eco. Cyc. org l E. coli Encyclopedia l Review-level Model-Organism Database for E. coli l Tracks evolving annotation of the E. coli genome and cellular networks l The two paradigms of Eco. Cyc l “Multi-dimensional annotation of the E. coli K-12 genome” l Positions of genes; functions of gene products – 76% / 66% exp l Gene Ontology terms; Multi. Fun terms l Gene product summaries and literature citations l Evidence codes l Multimeric complexes l Metabolic pathways Karp, Gunsalus, Collado-Vides, Paulsen l Cellular regulation Nuc. Acids Res. 35: 7577 2007 17 ASM News 70: 25 2004 Science 293: 2040 SRI International Bioinformatics

URL: Eco. Cyc. org Eco. Cyc = E. coli Dataset + Pathway/Genome Navigator Pathways: 246 Eco. Cyc v 13. 6 Reactions: Metabolic: 1394 Transport: 246 Compounds: 1, 830 Citations: 19, 000 Proteins: 4, 479 Complexes: 895 RNAs: 285 Genes: 4, 492 18 Gene Regulation: Operons: 3, 369 Trans Factors: 196 Promoters: 1, 796 TF Binding Sites: 2, 205 SRI International Bioinformatics

Paradigm 1: Eco. Cyc as Textual Review Article l All gene products for which experimental literature exists are curated with a minireview summary l Found on protein and RNA pages, not gene pages! l 3257 gene products contain summaries l Summaries cover function, interactions, mutant phenotypes, crystal structures, regulation, and more l Additional summaries found in pages for operons, pathways l Eco. Cyc 19 cites 17, 300 publications SRI International Bioinformatics

Paradigm 2: Eco. Cyc as Computational Symbolic Theory l Highly structured, high-fidelity knowledge representation provides computable information l Each molecular species defined as a DB object l Genes, proteins, small molecules l Each molecular interaction defined as a DB object l Metabolic reactions l Transport reactions l Transcriptional regulation of gene expression l 220 database fields capture extensive properties and relationships 20 SRI International Bioinformatics

Eco. Cyc Procedures l DB updates performed by 5 staff curators l Information gathered from biomedical literature u u u l 21 Enter data into structured database fields Author extensive summaries Update evidence codes Corrections submitted by E. coli researchers l Four releases per year l Quality assurance of data and software l Evaluate database consistency constraints l Perform element balancing of reactions l Run other checking programs SRI International Bioinformatics

Eco. Cyc Accelerates Science l l l 22 Experimentalists l E. coli experimentalists l Experimentalists working with other microbes l Analysis of expression data Computational biologists l Biological research using computational methods l Genome annotation l Study connectivity of E. coli metabolic network l Study phylogentic extent of metabolic pathways and enzymes in all domains of life Bioinformaticists l Training and validation of new bioinformatics algorithms – predict operons, promoters, protein functional linkages, protein-protein interactions, Metabolic engineers l “Design of organisms for the production of organic acids, amino acids, ethanol, hydrogen, and solvents “ Educators SRI International Bioinformatics

Meta. Cyc: Metabolic Encyclopedia l l l Describe a representative sample of every experimentally determined metabolic pathway Describe properties of metabolic enzymes Literature-based DB with extensive references and commentary Pathways, reactions, enzymes, substrates Jointly developed by l P. Karp, R. Caspi, C. Fulcher, SRI International l L. Mueller, A. Pujar, Boyce Thompson Institute l S. Rhee, P. Zhang, Carnegie Institution Nucleic Acids Research 2008 23 SRI International Bioinformatics

Applications of Meta. Cyc l Reference source on metabolic pathways l Metabolic engineering l Find enzymes with desired activities, regulatory properties l Determine cofactor requirements l Predict pathways from genomes l Systematic studies of metabolism l Computer-aided 24 education SRI International Bioinformatics

Meta. Cyc Data -- Version 13. 6 25 Pathways 1, 436 Reactions 8, 200 Enzymes 6, 060 Small Molecules 8, 400 Organisms 1, 800 Citations 21, 700 SRI International Bioinformatics
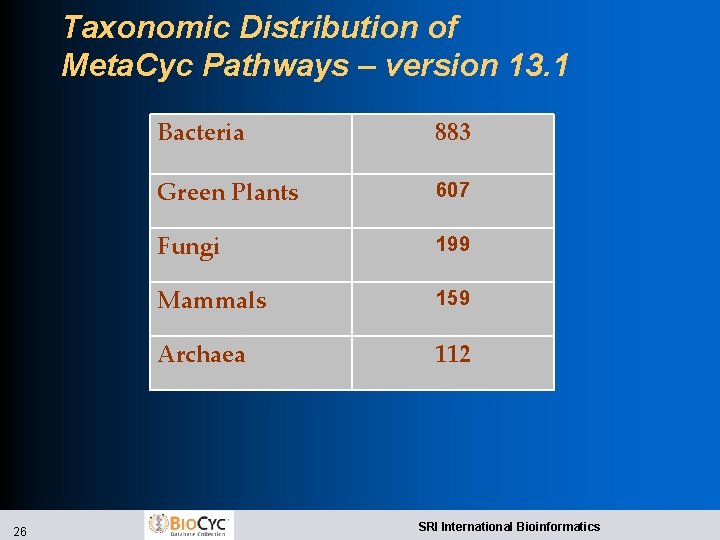
Taxonomic Distribution of Meta. Cyc Pathways – version 13. 1 26 Bacteria 883 Green Plants 607 Fungi 199 Mammals 159 Archaea 112 SRI International Bioinformatics

Enzyme Data Available in Meta. Cyc l Reaction(s) catalyzed l Alternative substrates l Activators, inhibitors, cofactors, prosthetic groups l Subunit structure l Genes l Features on protein sequence l Cellular location l p. I, molecular weight, Km, Vmax l Gene Ontology terms l Links to other bioinformatics databases 30 SRI International Bioinformatics

Meta. Cyc Pathway Variants l Pathways that accomplish similar biochemical functions using different biochemical routes l Alanine biosynthesis I – E. coli l Alanine biosynthesis II – H. sapiens l Pathways that accomplish similar biochemical functions using similar sets of reactions l Several variants of TCA Cycle 31 SRI International Bioinformatics

Meta. Cyc Super-Pathways l l l 32 Groups of pathways linked by common substrates Example: Super-pathway containing l Chorismate biosynthesis l Tryptophan biosynthesis l Phenylalanine biosynthesis l Tyrosine biosynthesis Super-pathways defined by listing their component pathways Multiple levels of super-pathways can be defined Pathway layout algorithms accommodate super-pathways SRI International Bioinformatics

Comparison of Bio. Cyc to KEGG 34 l KEGG approach: Static collection of reference pathway diagrams are color-coded to produce organism-specific views l KEGG vs Meta. Cyc: Resource on literature-derived pathways l KEGG maps are not pathways Nuc Acids Res 34: 3687 2006 l KEGG maps contain multiple biological pathways l KEGG maps are composites of pathways in many organisms -- do not identify what specific pathways elucidated in what organisms l KEGG has no literature citations, no comments, less enzyme detail l KEGG vs Bio. Cyc organism-specific PGDBs l KEGG does not curate or customize pathway networks for each organism l Highly curated PGDBs now exist for important organisms such as E. coli, yeast, mouse, Arabidopsis SRI International l KEGG re-annotates entire genome for each Bioinformatics organism

Comparison of Pathway Tools to KEGG l Inference tools l KEGG does not predict presence or absence of pathways l KEGG lacks pathway hole filler, operon predictor l Curation tools l KEGG does not distribute curation tools l No ability to customize pathways to the organism l Pathway Tools schema much more comprehensive l Visualization and analysis l KEGG does not perform automatic pathway layout l KEGG metabolic-map diagram extremely limited l No comparative pathway analysis 35 SRI International Bioinformatics

Pathway Tools Implementation Details 36 l Platforms: l Macintosh, PC/Linux, and PC/Windows platforms l Same binary can run as desktop app or Web server l Production-quality software l Version control l Two regular releases per year l Extensive quality assurance l Extensive documentation l Auto-patch l Automatic DB-upgrade l 420, 000 lines of Lisp code SRI International Bioinformatics

l Ptools-support@ai. sri. com 37 SRI International Bioinformatics
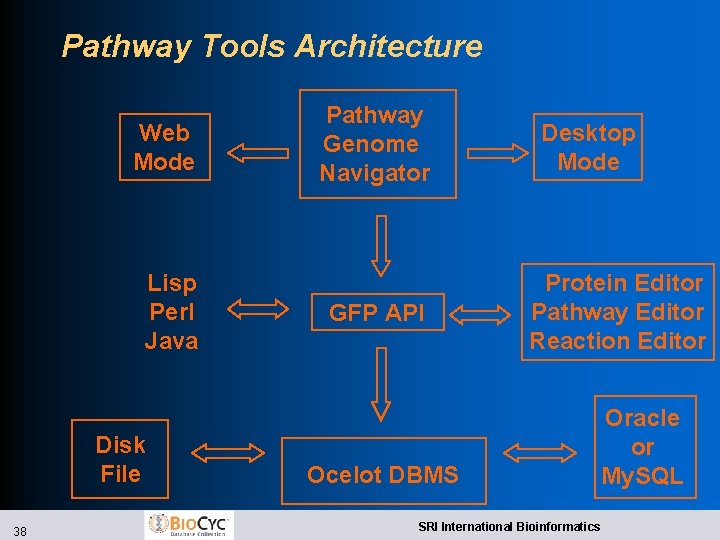
Pathway Tools Architecture Web Mode Lisp Perl Java Disk File 38 Pathway Genome Navigator GFP API Desktop Mode Protein Editor Pathway Editor Reaction Editor Ocelot DBMS SRI International Bioinformatics Oracle or My. SQL

Ocelot Knowledge Server Architecture l Frame data model l Minimizes size of schema relative to semantic complexity l Schema is stored within the DB Schema is self documenting Slot units define metadata about slots l Domain, range, inverse l Collection type, number of values, value constraints l Comment l l l 39 Schema evolution facilitated by l Easy addition/removal of slots, or alteration of slot datatypes l Flexible data formats that do not require dumping/reloading of data SRI International Bioinformatics

Ocelot Storage System Architecture l Persistent storage via disk files or Oracle or My. SQL l Concurrent development: Oracle or My. SQL l Single-user development: disk files l Oracle/My. SQL DBMS storage l DBMS is submerged within Ocelot, invisible to users l Frames transferred from DBMS to Ocelot u u l 40 On demand By background prefetcher Memory cache Persistent disk cache to speed performance via Internet Transaction logging facility SRI International Bioinformatics

Why Do We Code in Common Lisp? l Gatt studied Lisp and Java implementation of 16 programs by 14 programmers (Intelligence 11: 21 2000) l The average Lisp program ran 33 times faster than the average Java program l The average Lisp program was written 5 times faster than the average Java program l Roberts compared Java and Lisp implementations of a Domain Name Server (DNS) resolver 41 l http: //www. findinglisp. com/papers/case_study_java_lisp_dns. html l The Lisp version had ½ as many lines as code SRI International Bioinformatics

Common Lisp Programming Environment l Interpreted and/or compiled execution l Fabulous debugging environment l High-level language l Interactive data exploration l Extensive built-in libraries l Dynamic redefinition l Find out more! l See ALU. org or l http: //www. international-lisp-conference. org/ 42 SRI International Bioinformatics

Patho. Logic Processing 1. 2. 3. 4. 5. 6. 43 Translate source genome to PGDB form Predict operons Predict metabolic pathways Predict pathway hole fillers Transport inference parser Build metabolic overview diagram SRI International Bioinformatics
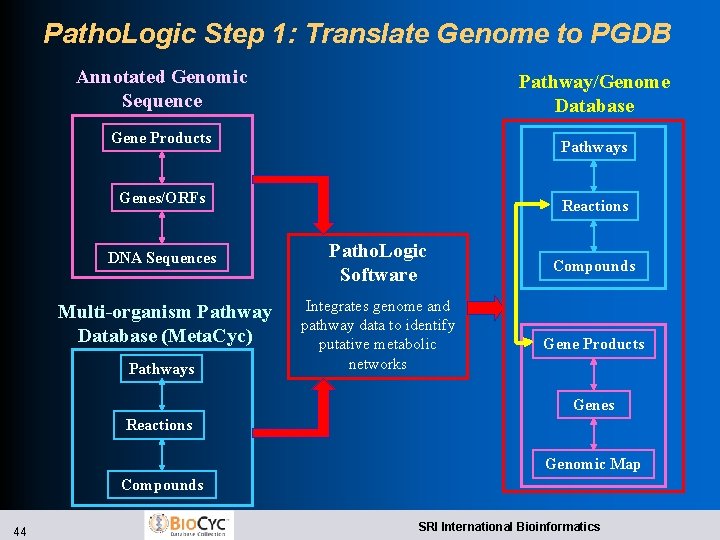
Patho. Logic Step 1: Translate Genome to PGDB Annotated Genomic Sequence Pathway/Genome Database Gene Products Pathways Genes/ORFs DNA Sequences Multi-organism Pathway Database (Meta. Cyc) Pathways Reactions Patho. Logic Software Integrates genome and pathway data to identify putative metabolic networks Compounds Gene Products Genes Reactions Genomic Map Compounds 44 SRI International Bioinformatics

Patho. Logic Step 3: Prediction of Metabolic Pathways l Infer reaction complement of organism l Match enzymes in source genome to Meta. Cyc reactions they catalyze l Match enzyme names and EC numbers to Meta. Cyc l Support user in manually matching additional enzymes l Computationally predict which Meta. Cyc metabolic pathways are present l For each Meta. Cyc pathway, evaluate which of its reactions are catalyzed by the organism 47 SRI International Bioinformatics
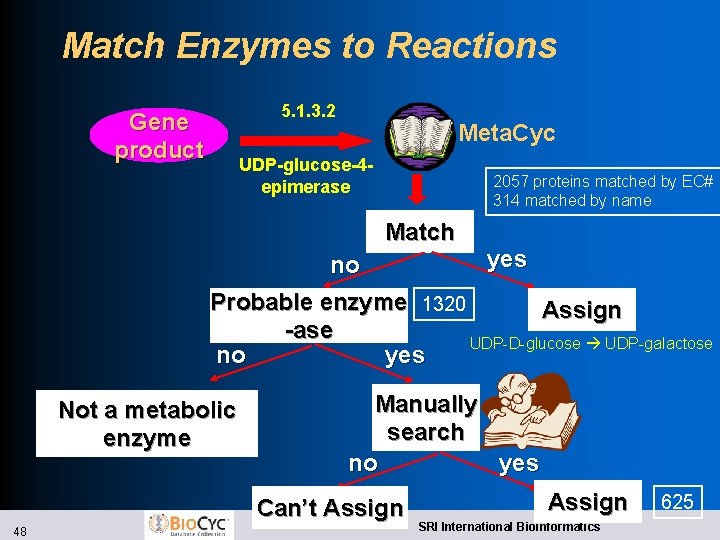
Match Enzymes to Reactions 5. 1. 3. 2 Gene product Meta. Cyc UDP-glucose-4 epimerase 2057 proteins matched by EC# 314 matched by name Match yes no Probable enzyme 1320 Assign -ase UDP-D-glucose UDP-galactose no yes Not a metabolic enzyme Manually search no yes Can’t Assign 48 Assign SRI International Bioinformatics 625
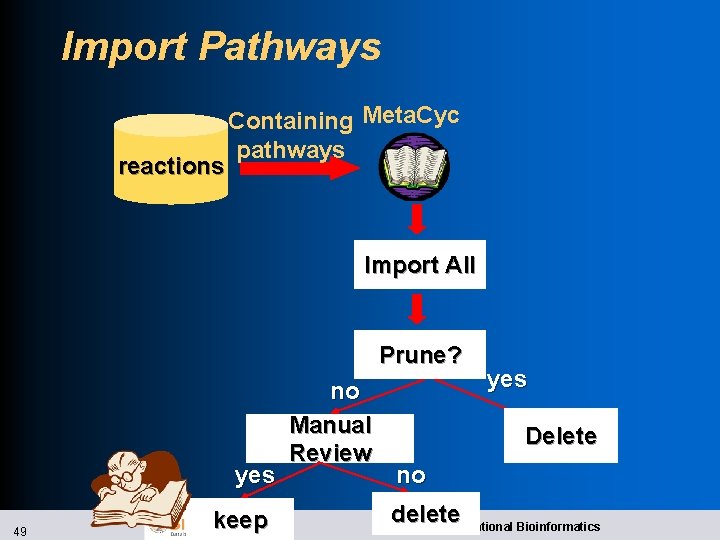
Import Pathways reactions Containing Meta. Cyc pathways Import All Prune? yes 49 keep no Manual Review yes Delete no delete SRI International Bioinformatics

Pathway Prediction l Prediction is hard because l Enzyme naming is irregular l Some reactions present in multiple pathways l Pathway variants share many reactions in common l Meta. Cyc now has many pathways 50 SRI International Bioinformatics

Pathway Scoring Criteria l Imported pathways must satisfy: l Pathways outside their taxonomic range must have enzymes for all reactions l If any reactions in a pathway are designated as “key, ” an enzyme must be present for at least one l Pathway P is imported if any conditions satisfied: l One unique enzyme present for P l P missing at most one reaction l More reactions present than absent for P l P is not a superset of another pathway with the same number of enzymes present 51 SRI International Bioinformatics
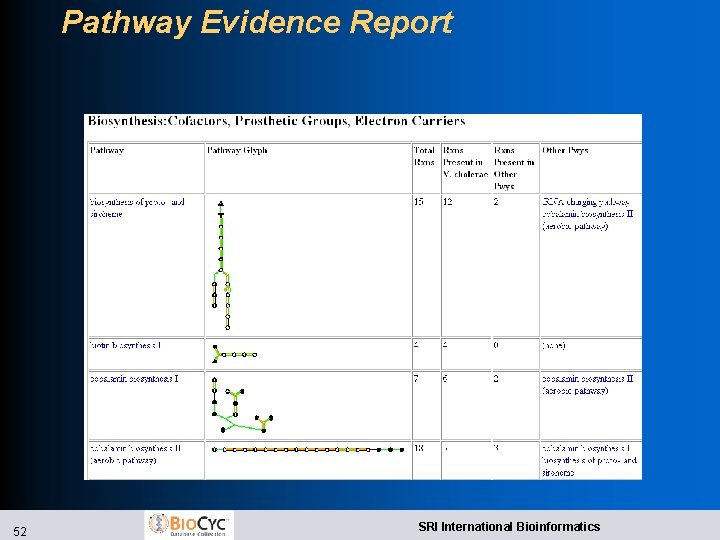
Pathway Evidence Report 52 SRI International Bioinformatics

Patho. Logic Step 4: Pathway Hole Filler l. Definition: Pathway Holes are reactions in metabolic pathways for which no enzyme is identified L-aspartate 1. 4. 3. - iminoaspartate quinolinate synthetase nad. A quinolinate holes NAD+ synthetase, NH 3 dependent CC 3619 deamido-NAD n. n. pyrophosphorylase nad. C 2. 7. 7. 18 NAD 53 nicotinate nucleotide 6. 3. 5. 1 SRI International Bioinformatics
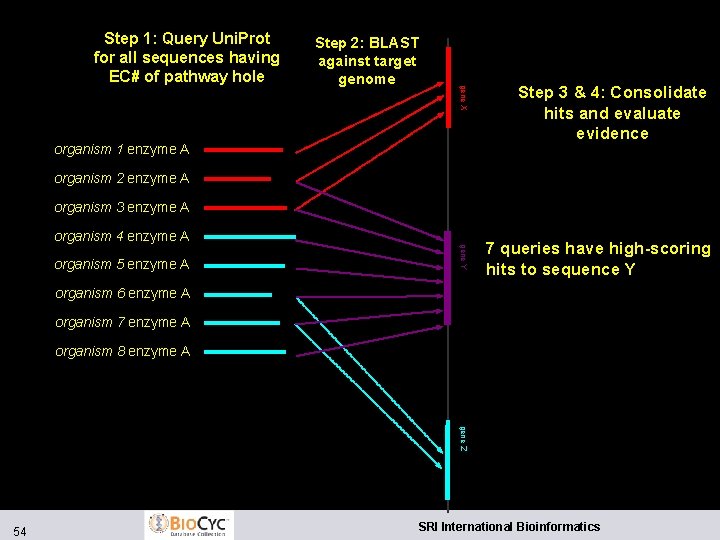
Step 2: BLAST against target genome gene X Step 1: Query Uni. Prot for all sequences having EC# of pathway hole organism 1 enzyme A Step 3 & 4: Consolidate hits and evaluate evidence organism 2 enzyme A organism 3 enzyme A organism 4 enzyme A gene Y organism 5 enzyme A 7 queries have high-scoring hits to sequence Y organism 6 enzyme A organism 7 enzyme A organism 8 enzyme A gene Z 54 SRI International Bioinformatics

Pathway Hole Filler l Why should hole filler find things beyond the original genome annotation? l Reverse BLAST searches more sensitive l Reverse BLAST searches find second domains l Integration of multiple evidence types 56 SRI International Bioinformatics

Caulobacter crescentus Pathway Holes l l 130 pathways containing 582 reactions 92 pathways contain 236 pathway holes Caulobacter holes filled: l 77 holes filled at P >0. 9 l Previous functions of candidate hole fillers: l No predicted function l Correctly assigned single function l Incorrectly assigned function l Imprecise functional assignment BMC Bioinformatics 5: 76 2004 57 SRI International Bioinformatics
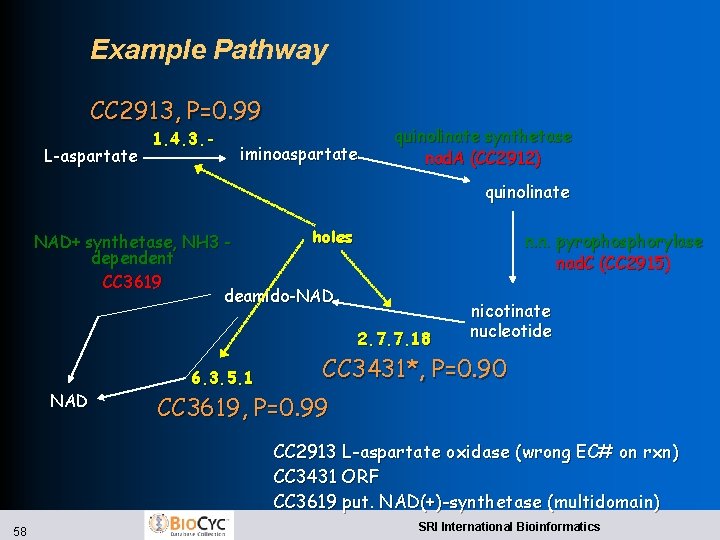
Example Pathway CC 2913, P=0. 99 L-aspartate 1. 4. 3. - iminoaspartate quinolinate synthetase nad. A (CC 2912) quinolinate holes NAD+ synthetase, NH 3 dependent CC 3619 deamido-NAD n. n. pyrophosphorylase nad. C (CC 2915) 2. 7. 7. 18 NAD 6. 3. 5. 1 nicotinate nucleotide CC 3431*, P=0. 90 CC 3619, P=0. 99 CC 2913 L-aspartate oxidase (wrong EC# on rxn) CC 3431 ORF CC 3619 put. NAD(+)-synthetase (multidomain) 58 SRI International Bioinformatics

Patho. Logic Step 5: Transport Inference Parser l. Problem: Write a program to query a genome annotation to compute the substrates an organism can transport l. Typical l l 59 genome annotations for transporters: ATP transporter for ribose ABC transporter D-ribose ATP transporter ABC transporter, membrane spanning protein [ribose] ABC transporter, membrane spanning protein [D-ribose] SRI International Bioinformatics

Transport Inference Parser l. Input: “ATP transporter of phosphonate” l. Output: Structured description of transport activity l. Locates most transporters in genome annotation using keyword analysis l. Parse l l l product name using a series of rules to identify: Transported substrate, co-substrate Influx/efflux Energy coupling mechanism l. Creates transport reaction object: phosphonate[periplasm] + H 2 O + ATP = phosphonate + Pi + ADP 60 SRI International Bioinformatics
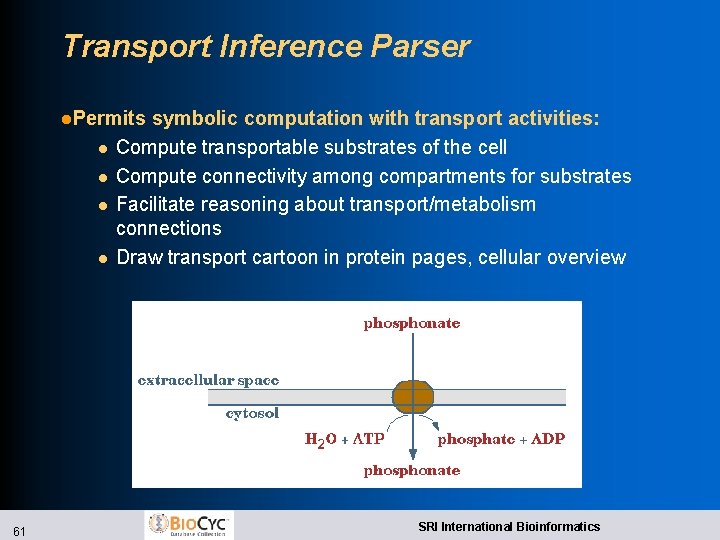
Transport Inference Parser l. Permits l l 61 symbolic computation with transport activities: Compute transportable substrates of the cell Compute connectivity among compartments for substrates Facilitate reasoning about transport/metabolism connections Draw transport cartoon in protein pages, cellular overview SRI International Bioinformatics
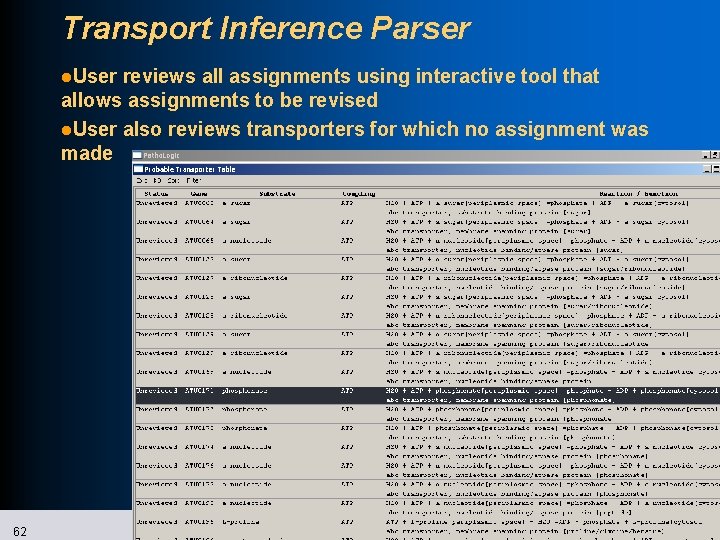
Transport Inference Parser l. User reviews all assignments using interactive tool that allows assignments to be revised l. User also reviews transporters for which no assignment was made 62 SRI International Bioinformatics

Regulation 63 SRI International Bioinformatics

Regulation in Pathway Tools l Substrate-level regulation of enzyme activity l Binding to proteins or small molecules (phosphorylation) l Regulation of transcription initiation l Attenuation of transcription l Regulation of translation by proteins and by small RNAs 65 SRI International Bioinformatics

Regulation in Pathway Tools l Editing tools l Transcription factor display window l Transcription unit display window l Regulatory 67 Overview / Omics Viewer SRI International Bioinformatics
Regulatory Interaction Editor 68 SRI International Bioinformatics
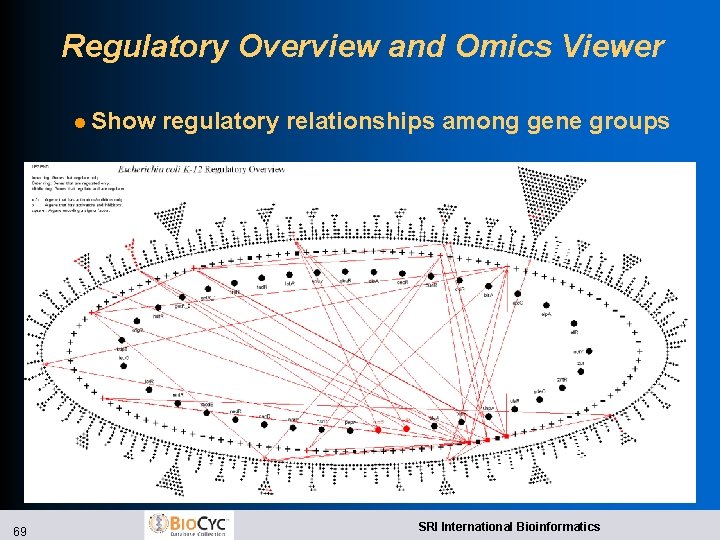
Regulatory Overview and Omics Viewer l Show 69 regulatory relationships among gene groups SRI International Bioinformatics

Comparative Analysis l Via Cellular Overview l Comparative genome browser l Comparative pathway table l Comparative analysis reports l Compare reaction complements l Compare pathway complements l Compare transporter complements 71 SRI International Bioinformatics

Information Sources 73 l Pathway Tools User’s Guide l aic-export/pathwaytools/ptools/13. 0/doc/manuals/userguide. pdf l NOTE: Location of the aic-export directory can vary across different computers l Pathway Tools Web Site l http: //bioinformatics. ai. sri. com/ptools/ l Publications, FAQ, programming examples, etc. l Slides from this tutorial l http: //www. ai. sri. com/pkarp/talks/ l Bio. Cyc Webinars l http: //biocyc. org/webinar. shtml SRI International Bioinformatics

Bio. Cyc and Pathway Tools Availability l Bio. Cyc. org Web site and database files freely available to all l Pathway Tools freely available to non-profits l Macintosh, PC/Windows, PC/Linux 74 SRI International Bioinformatics

Acknowledgements l. SRI l l. Funding Suzanne Paley, Ron Caspi, Ingrid Keseler, Carol Fulcher, Markus Krummenacker, Alex Shearer, Tomer Altman, Joe Dale, Fred Gilham, Pallavi Kaipa l. Eco. Cyc l Collaborators Julio Collado-Vides, Robert Gunsalus, Ian Paulsen l. Meta. Cyc l l l sources: NIH National Center for Research Resources NIH National Institute of General Medical Sciences NIH National Human Genome Research Institute Bio. Cyc. org Collaborators l Sue Rhee, Peifen Zhang, Kate Bio. Cyc Dreherwebinars: biocyc. org/webinar. shtml Learn more from l Lukas Mueller, Anuradha SRI International Bioinformatics 75

Symbolic Systems Biology Definition: Global analyses of biological systems using symbolic computing 76 SRI International Bioinformatics

Symbolic Systems Biology 77 l “Symbolic computing is concerned with the representation and manipulation of information in symbolic form. It is often contrasted with numeric representation. ” -- R. Cameron l Examples of symbolic computation: l Symbolic algebra programs, e. g. , Mathematica, Graphing Calculator l Compilers and interpreters for programming languages l Database query languages l Text analysis programs, e. g. , Google l String matching for DNA and protein sequences l Artificial Intelligence methods, e. g. , expert systems, symbolic logic, machine learning, natural language understanding SRI International Bioinformatics

Symbolic Systems Biology l Concerned with different questions than quantitative systems biology l Symbolic analyses can in many cases produce answers when quantitative approaches fail because of lack of parameters or intractable mathematics l Symbolic computation is intimately dependent on the use of structured ontologies 78 SRI International Bioinformatics

Pathway Tools Ontology l 1064 classes l Main classes such as: u l 79 Pathways, Reactions, Compounds, Macromolecules, Proteins, Replicons, DNA-Segments (Genes, Operons, Promoters) Taxonomies for Pathways, Reactions, Compounds l 205 slots l Meta-data: Creator, Creation-Date l Comment, Citations, Common-Name, Synonyms l Attributes: Molecular-Weight, DNA-Footprint-Size l Relationships: Catalyzes, Component-Of, Product l Classes, instances, slots all stored side by side in DBMS SRI International Bioinformatics

Symbolic Computation on PGDBs: Complex Query l Show metabolic enzymes regulated by a specified transcription factor l For transcription factor F: l Find all promoters F regulates l Find all genes in the operons controlled by those promoters u 80 Find their protein products – Find the reactions they catalyze » Highlight them in the diagram SRI International Bioinformatics

Critiquing the Parts List 81 Slide thanks to Hirotada Mori (minus the banana!) SRI International Bioinformatics

Dead End Metabolites l. A small molecule C is a dead-end if: l Def 1 does not depend on accurate reaction directions; l Def 2 more accurate l Definition 1: l C is a substrate in only one reaction of the set of SMM reactions occurring in Compartment AND l No reactions exist containing parent classes of C AND l No transporter acts on C in Compartment, nor on parent classes of C l Definition 2: l C is produced only by SMM reactions in Compartment, and no transporter acts on C in Compartment OR 82 SRI International Bioinformatics

Dead End Metabolites l Not yet an official part of Pathway Tools l Contact us if you’d like to use it 83 SRI International Bioinformatics

Reachability Analysis of Metabolic Networks l l l Given: l A PGDB for an organism l A set of initial metabolites Infer: l What set of products can be synthesized by the smallmolecule metabolism of the organism Motivations: l Quality control for PGDBs u Verify that a known growth medium yields known essential compounds Experiment with other growth media l Experiment with reaction knock-outs Limitations l Cannot properly handle compounds required for their own synthesis l Nutrients needed for reachability may be a superset of those required for growth l l Romero and Karp, Pacific Symposium on Biocomputing, 2001 84 SRI International Bioinformatics

Algorithm: Forward Propagation Through Production System l l Each reaction becomes a production rule Each of the 21 metabolites in the nutrient set becomes an axiom Nutrient set Tr an Products sp o rt A+B C 85 Metabolite pool PGDB reaction set “Fire” reactions Reactants SRI International Bioinformatics

Nutrients: A, B, C, E, F A+B W C+D X E+F Y W+Y Z Produced Compounds: W, Y, Z 86 SRI International Bioinformatics

Initial Metabolite Nutrient Set (Total: 21 compounds) 87 SRI International Bioinformatics

Essential Compounds E. coli Total: 41 compounds l Proteins (20) l Amino acids l Nucleic acids (DNA & RNA) (8) l Nucleosides l Cell membrane (3) l Phospholipids l Cell wall (10) l Peptidoglycan precursors l Outer cell wall precursors (Lipid-A, oligosaccharides) 88 SRI International Bioinformatics
89 SRI International Bioinformatics

l http: //brg. ai. sri. com/ptools 09/slides/Tuesday/grow th-experiment-Markus-Krummenacker. txt 90 SRI International Bioinformatics

Flux Balance Modeling l l l 91 Generate, store, and update metabolic model within Pathway Tools l Fast, accurate generation of metabolic model l Close coupling to genome and regulatory information l Extensive schema l Extensive query and visualization tools Debug/validate model using Pathway Tools Export to SBML and import to constraint solver for model execution Visualize reaction flux and omics data using overviews Copy/update multiple PGDBs to reflect alternative strains SRI International Bioinformatics
- Slides: 81