Passwort beta Sheet 20 1 Ziele der Vorlesung

Passwort: beta. Sheet 20 1

Ziele der Vorlesung • Faszination der Bioinformatik • Methoden und Techniken • Wissenschaftliches Arbeiten (Po. L= Problemorientiertes Lernen) 2

Introduction to Protein Structure Overview and Basic Principles Slides by Niko Beerenwinkel and Thomas Lengauer ©Thomas Lengauer, MPI Informatik 3

Roadmap l This lecture: l Protein structure l Building blocks l Basic structural elements l Motifs l Folds l Multi-domain topologies ©Thomas Lengauer, MPI Informatik 4

Material on Basic Biology l Literature l Stryer Biochemie: Berg, Tymoczko, Gatto Jr. , Stryer; Springer Spektrum, 2017 l Principles of Protein Structure: C. Branden, J. Tooze, 2 nd ed. , Garland, 1999 l An Introduction to Genetic Analysis: Griffiths et al. , Freeman, 2011 ©Thomas Lengauer, MPI Informatik 5
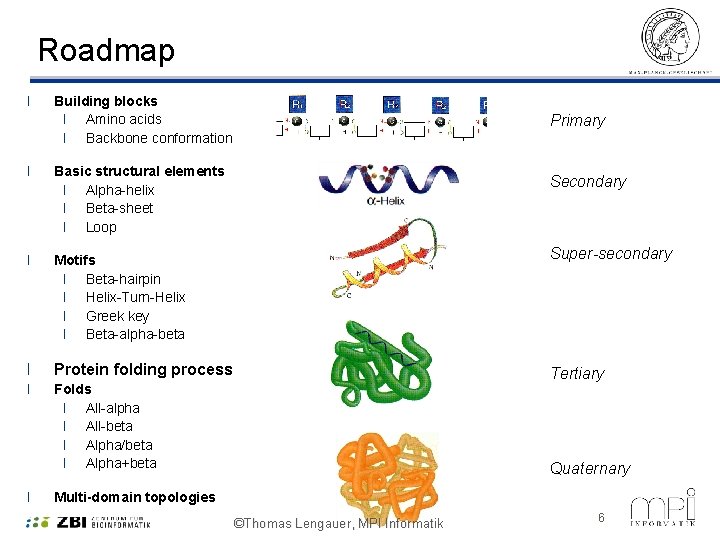
Roadmap l Building blocks l Amino acids l Backbone conformation l Basic structural elements l Alpha-helix l Beta-sheet l Loop l Motifs l Beta-hairpin l Helix-Turn-Helix l Greek key l Beta-alpha-beta l Protein folding process l Folds l All-alpha l All-beta l Alpha/beta l Alpha+beta l Primary Secondary Super-secondary Tertiary Quaternary Multi-domain topologies ©Thomas Lengauer, MPI Informatik 6
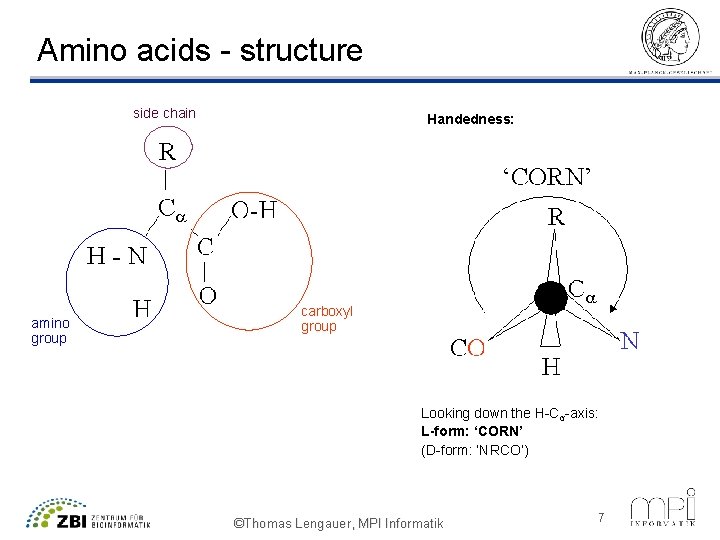
Amino acids - structure side chain amino group Handedness: carboxyl group Looking down the H-Cα-axis: L-form: ‘CORN’ (D-form: ‘NRCO’) ©Thomas Lengauer, MPI Informatik 7

The 20 natural amino acids ©Thomas Lengauer, MPI Informatik 8
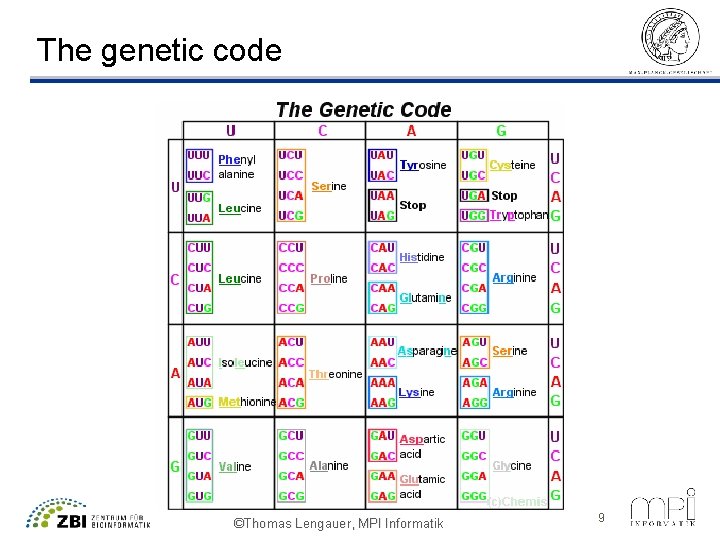
The genetic code ©Thomas Lengauer, MPI Informatik 9
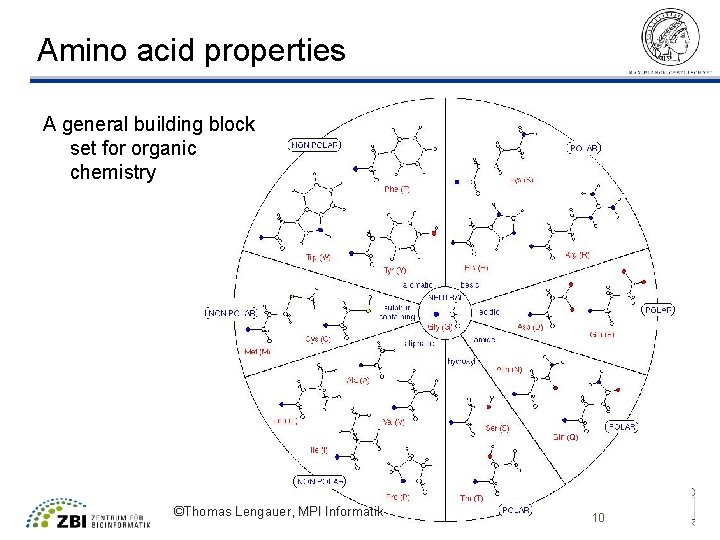
Amino acid properties A general building block set for organic chemistry ©Thomas Lengauer, MPI Informatik 10
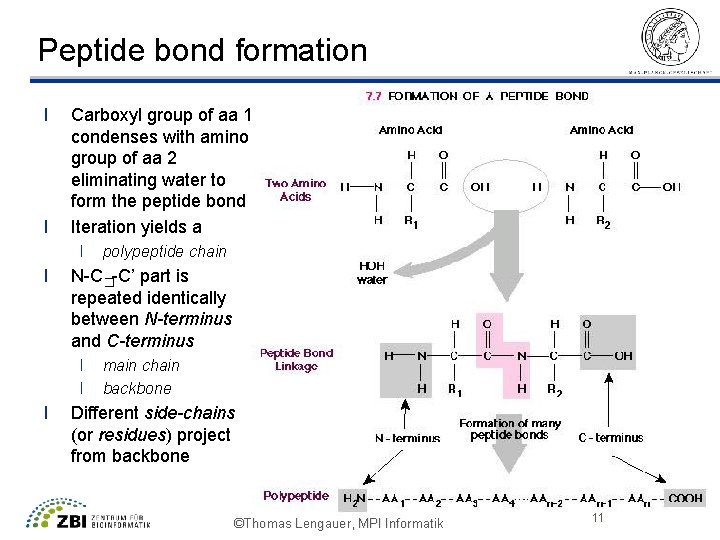
Peptide bond formation l l Carboxyl group of aa 1 condenses with amino group of aa 2 eliminating water to form the peptide bond Iteration yields a l l N-C�-C’ part is repeated identically between N-terminus and C-terminus l l l polypeptide chain main chain backbone Different side-chains (or residues) project from backbone ©Thomas Lengauer, MPI Informatik 11
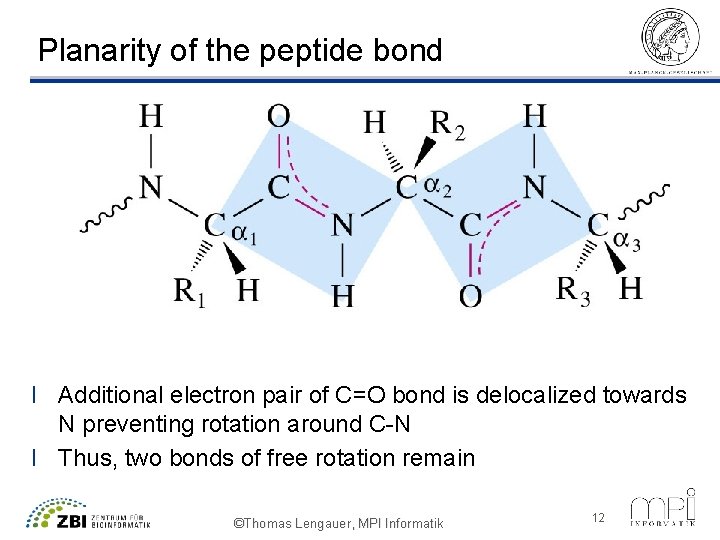
Planarity of the peptide bond l Additional electron pair of C=O bond is delocalized towards N preventing rotation around C-N l Thus, two bonds of free rotation remain ©Thomas Lengauer, MPI Informatik 12
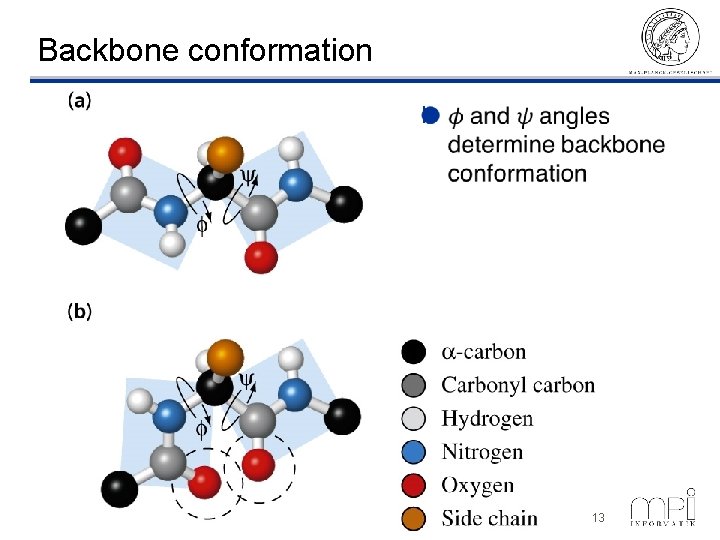
Backbone conformation l ©Thomas Lengauer, MPI Informatik 13
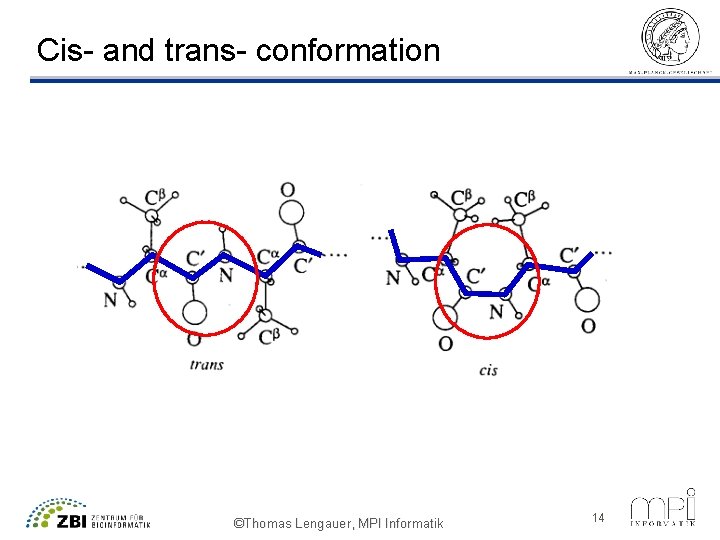
Cis- and trans- conformation ©Thomas Lengauer, MPI Informatik 14
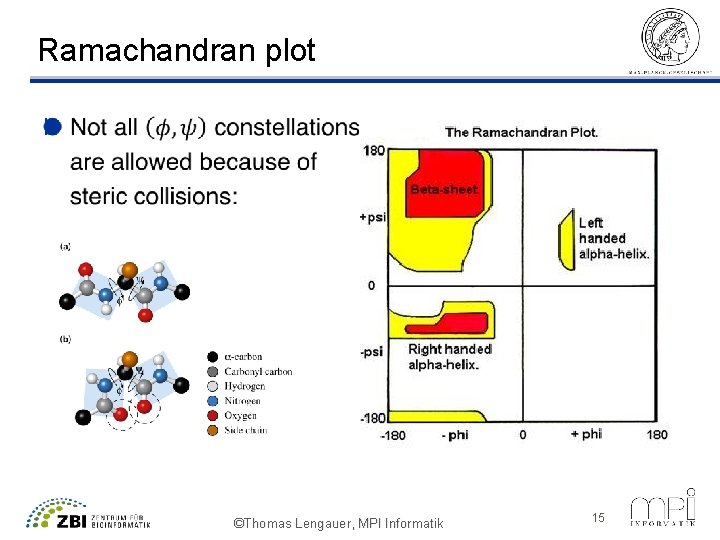
Ramachandran plot l ©Thomas Lengauer, MPI Informatik 15
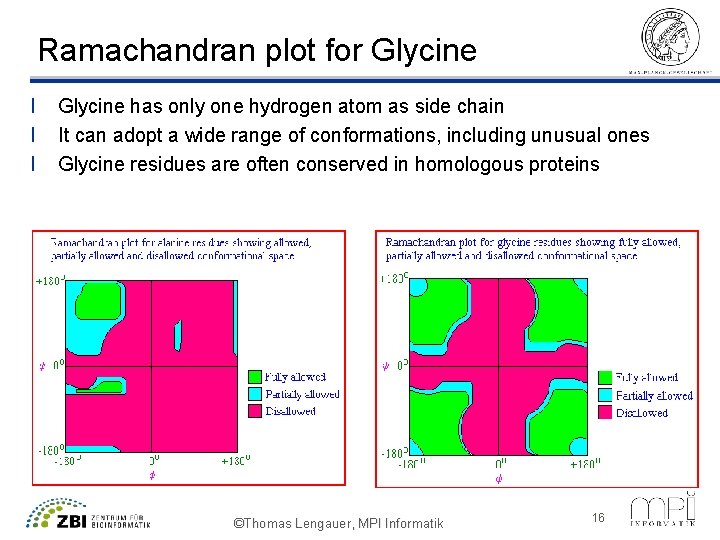
Ramachandran plot for Glycine l l l Glycine has only one hydrogen atom as side chain It can adopt a wide range of conformations, including unusual ones Glycine residues are often conserved in homologous proteins ©Thomas Lengauer, MPI Informatik 16
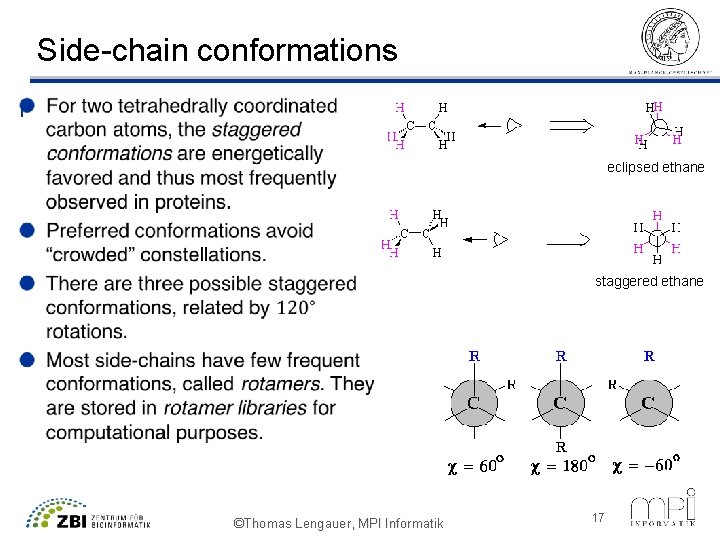
Side-chain conformations l eclipsed ethane staggered ethane ©Thomas Lengauer, MPI Informatik 17
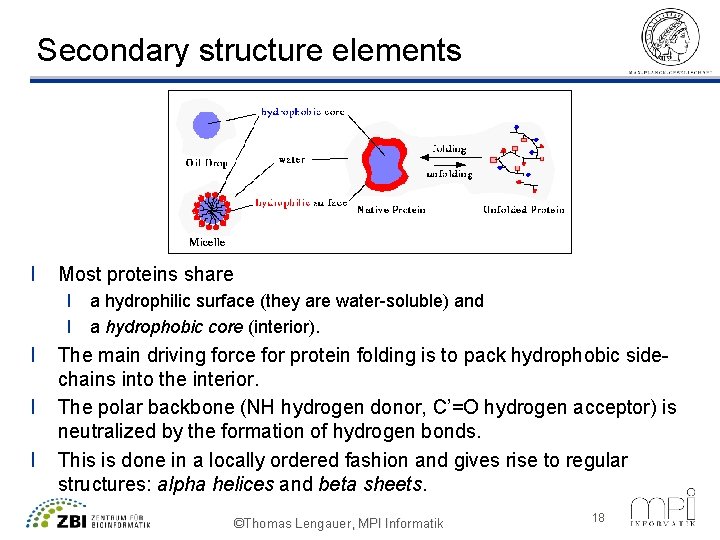
Secondary structure elements l Most proteins share l a hydrophilic surface (they are water-soluble) and l a hydrophobic core (interior). l l l The main driving force for protein folding is to pack hydrophobic sidechains into the interior. The polar backbone (NH hydrogen donor, C’=O hydrogen acceptor) is neutralized by the formation of hydrogen bonds. This is done in a locally ordered fashion and gives rise to regular structures: alpha helices and beta sheets. ©Thomas Lengauer, MPI Informatik 18
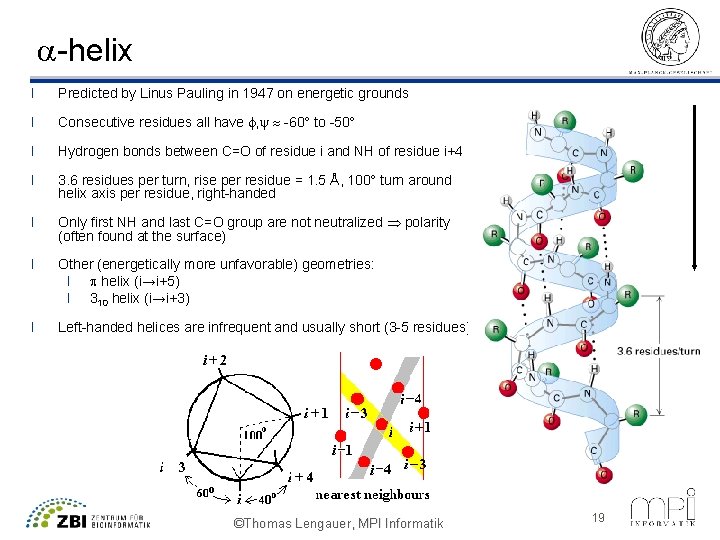
a-helix l Predicted by Linus Pauling in 1947 on energetic grounds l Consecutive residues all have , -60 to -50 l Hydrogen bonds between C=O of residue i and NH of residue i+4 l 3. 6 residues per turn, rise per residue = 1. 5 Å, 100 turn around helix axis per residue, right-handed l Only first NH and last C=O group are not neutralized polarity (often found at the surface) l Other (energetically more unfavorable) geometries: l helix (i→i+5) l 310 helix (i→i+3) l Left-handed helices are infrequent and usually short (3 -5 residues) ©Thomas Lengauer, MPI Informatik 19
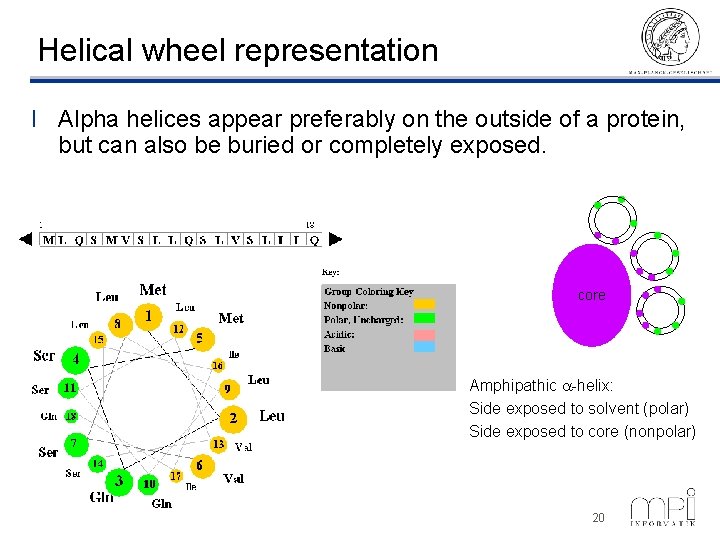
Helical wheel representation l Alpha helices appear preferably on the outside of a protein, but can also be buried or completely exposed. core Amphipathic a-helix: Side exposed to solvent (polar) Side exposed to core (nonpolar) ©Thomas Lengauer, MPI Informatik 20
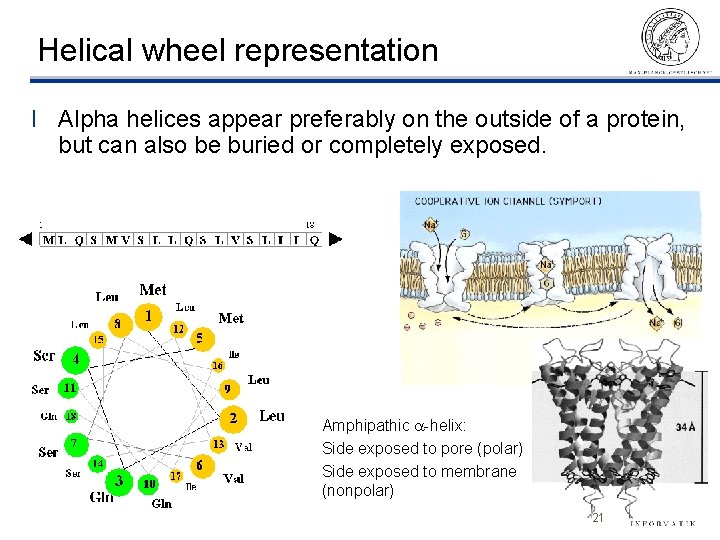
Helical wheel representation l Alpha helices appear preferably on the outside of a protein, but can also be buried or completely exposed. Amphipathic a-helix: Side exposed to pore (polar) Side exposed to membrane (nonpolar) ©Thomas Lengauer, MPI Informatik 21
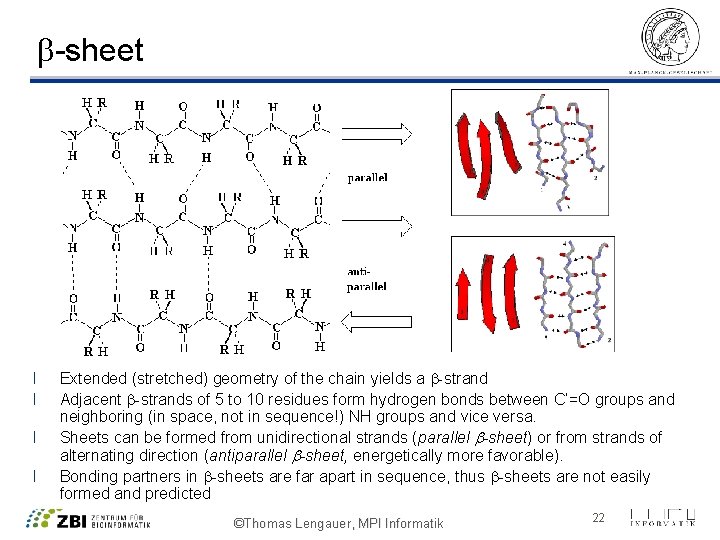
b-sheet l l Extended (stretched) geometry of the chain yields a b-strand Adjacent b-strands of 5 to 10 residues form hydrogen bonds between C’=O groups and neighboring (in space, not in sequence!) NH groups and vice versa. Sheets can be formed from unidirectional strands (parallel b-sheet) or from strands of alternating direction (antiparallel b-sheet, energetically more favorable). Bonding partners in b-sheets are far apart in sequence, thus b-sheets are not easily formed and predicted ©Thomas Lengauer, MPI Informatik 22
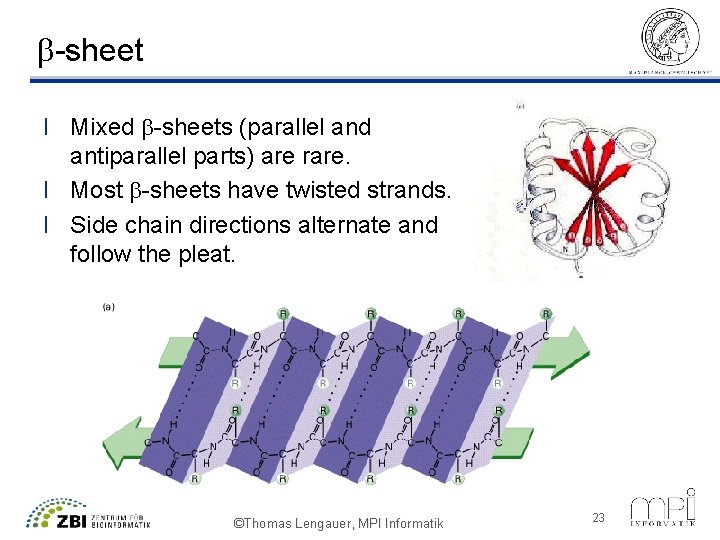
b-sheet l Mixed b-sheets (parallel and antiparallel parts) are rare. l Most b-sheets have twisted strands. l Side chain directions alternate and follow the pleat. ©Thomas Lengauer, MPI Informatik 23
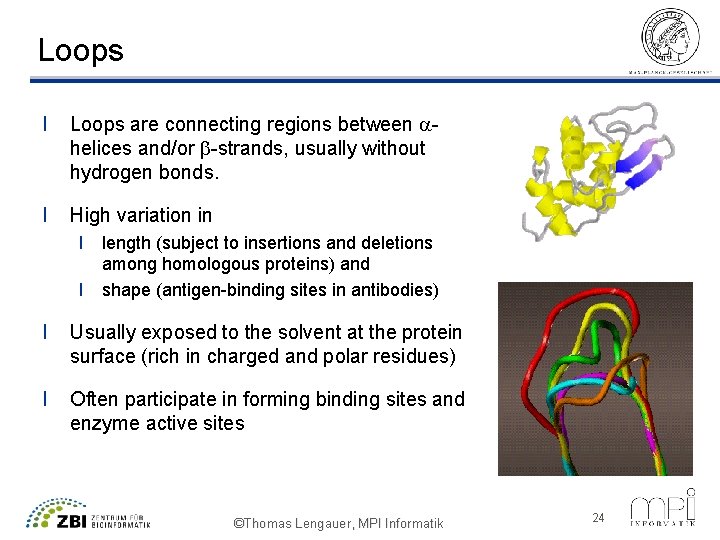
Loops l Loops are connecting regions between ahelices and/or b-strands, usually without hydrogen bonds. l High variation in l length (subject to insertions and deletions among homologous proteins) and l shape (antigen-binding sites in antibodies) l Usually exposed to the solvent at the protein surface (rich in charged and polar residues) l Often participate in forming binding sites and enzyme active sites ©Thomas Lengauer, MPI Informatik 24
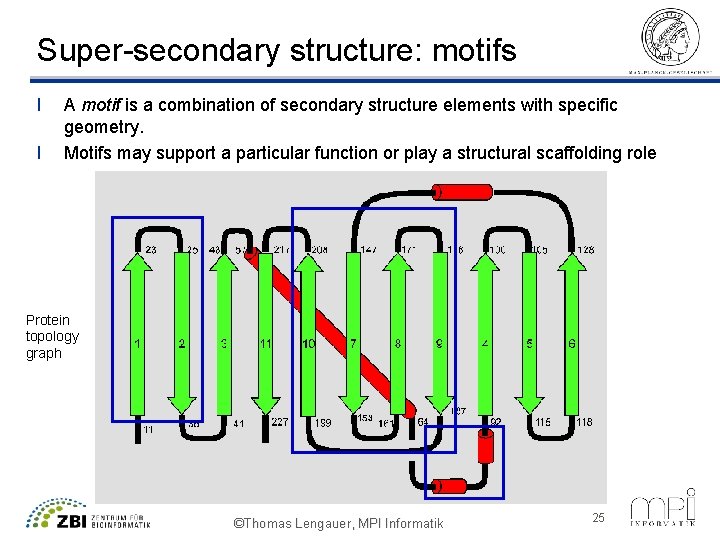
Super-secondary structure: motifs l l A motif is a combination of secondary structure elements with specific geometry. Motifs may support a particular function or play a structural scaffolding role Protein topology graph ©Thomas Lengauer, MPI Informatik 25
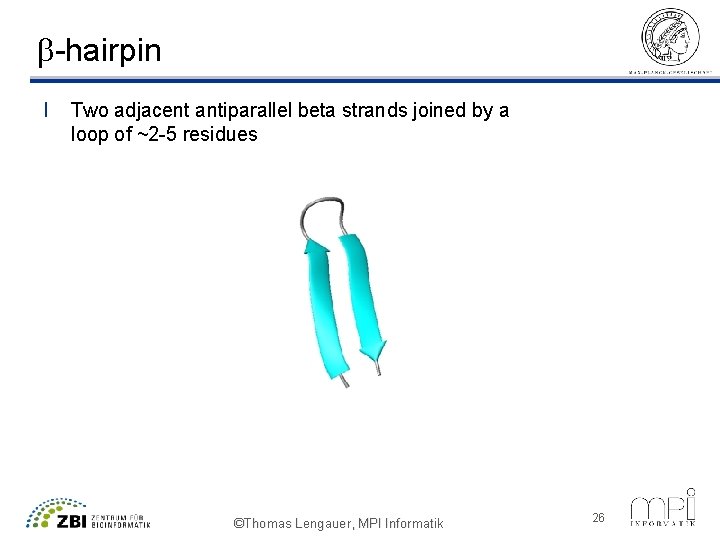
b-hairpin l Two adjacent antiparallel beta strands joined by a loop of ~2 -5 residues ©Thomas Lengauer, MPI Informatik 26
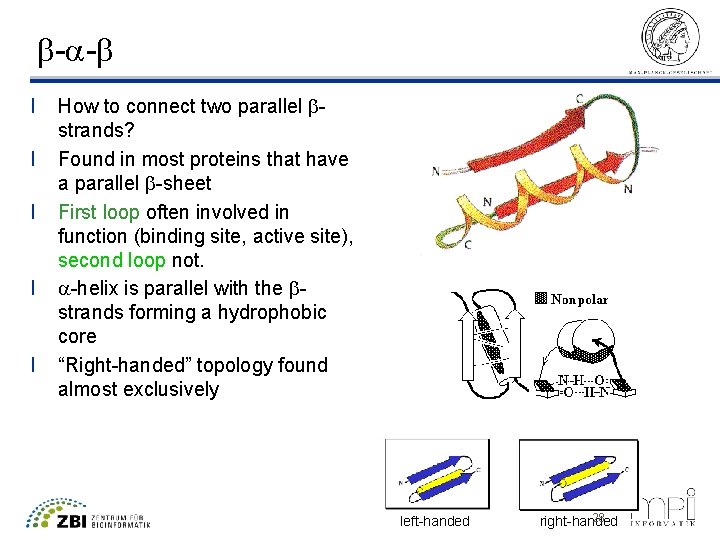
b-a-b l l l How to connect two parallel bstrands? Found in most proteins that have a parallel b-sheet First loop often involved in function (binding site, active site), second loop not. a-helix is parallel with the bstrands forming a hydrophobic core “Right-handed” topology found almost exclusively left-handed 28 right-handed
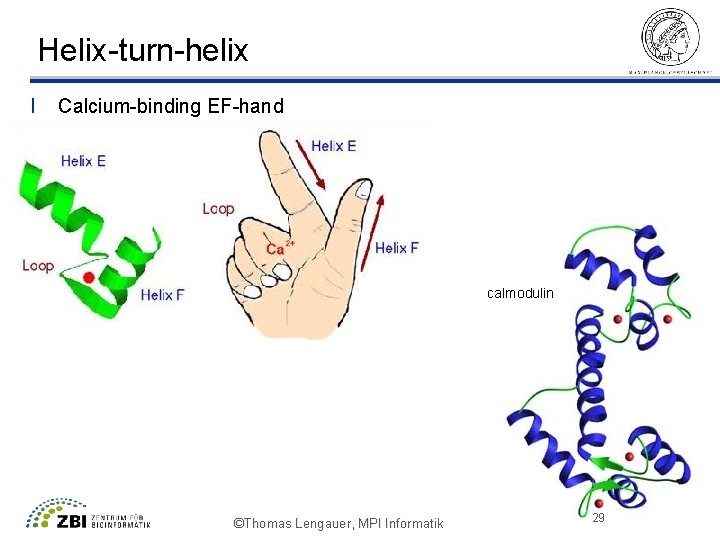
Helix-turn-helix l Calcium-binding EF-hand calmodulin ©Thomas Lengauer, MPI Informatik 29
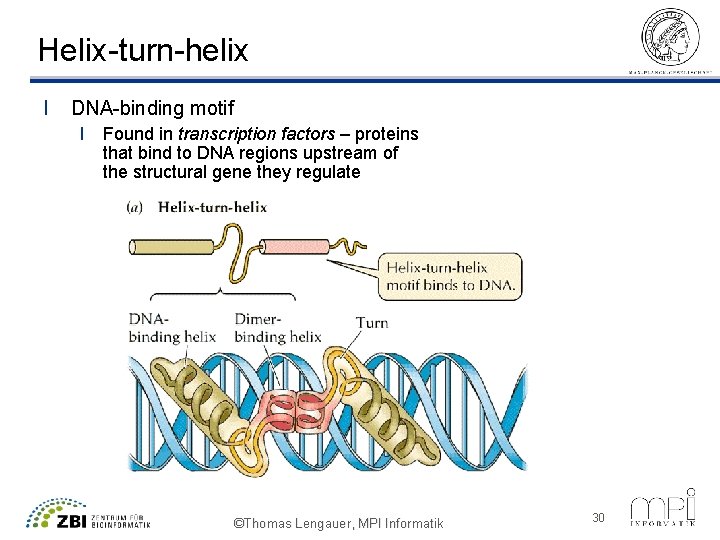
Helix-turn-helix l DNA-binding motif l Found in transcription factors – proteins that bind to DNA regions upstream of the structural gene they regulate ©Thomas Lengauer, MPI Informatik 30
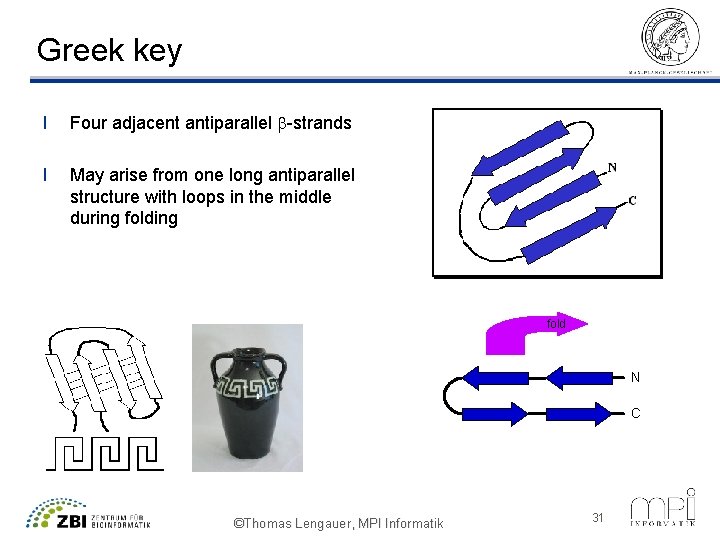
Greek key l Four adjacent antiparallel b-strands l May arise from one long antiparallel structure with loops in the middle during fold N C ©Thomas Lengauer, MPI Informatik 31
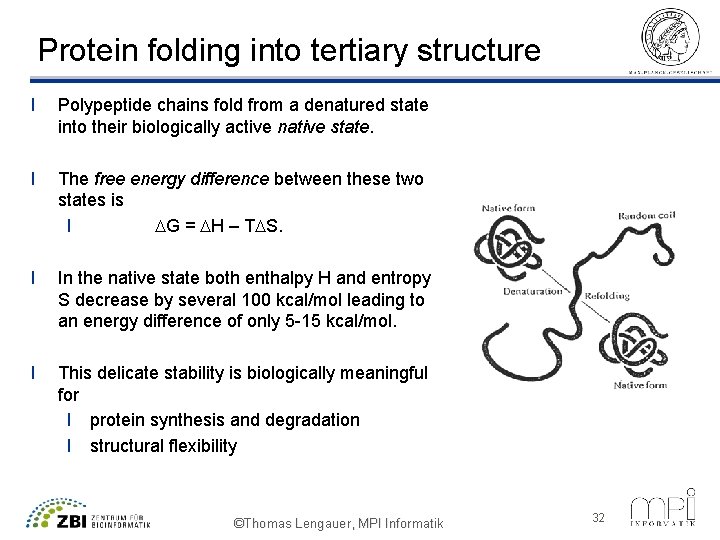
Protein folding into tertiary structure l Polypeptide chains fold from a denatured state into their biologically active native state. l The free energy difference between these two states is l G = H – T S. l In the native state both enthalpy H and entropy S decrease by several 100 kcal/mol leading to an energy difference of only 5 -15 kcal/mol. l This delicate stability is biologically meaningful for l protein synthesis and degradation l structural flexibility ©Thomas Lengauer, MPI Informatik 32
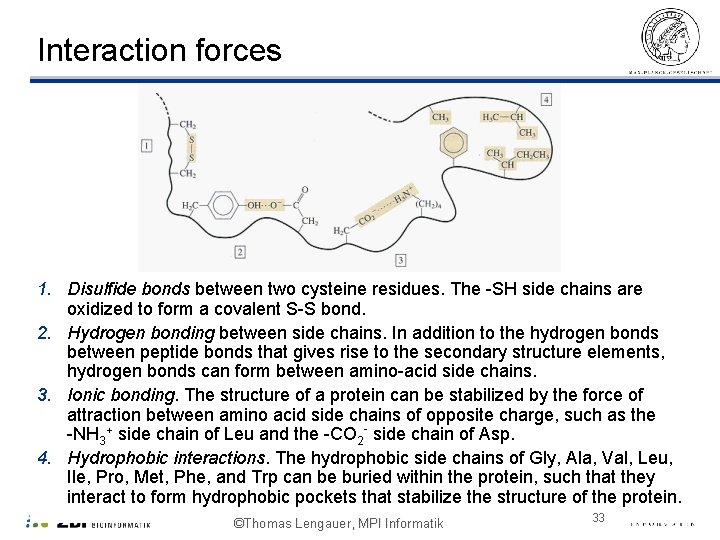
Interaction forces 1. Disulfide bonds between two cysteine residues. The -SH side chains are oxidized to form a covalent S-S bond. 2. Hydrogen bonding between side chains. In addition to the hydrogen bonds between peptide bonds that gives rise to the secondary structure elements, hydrogen bonds can form between amino-acid side chains. 3. Ionic bonding. The structure of a protein can be stabilized by the force of attraction between amino acid side chains of opposite charge, such as the -NH 3+ side chain of Leu and the -CO 2 - side chain of Asp. 4. Hydrophobic interactions. The hydrophobic side chains of Gly, Ala, Val, Leu, Ile, Pro, Met, Phe, and Trp can be buried within the protein, such that they interact to form hydrophobic pockets that stabilize the structure of the protein. ©Thomas Lengauer, MPI Informatik 33
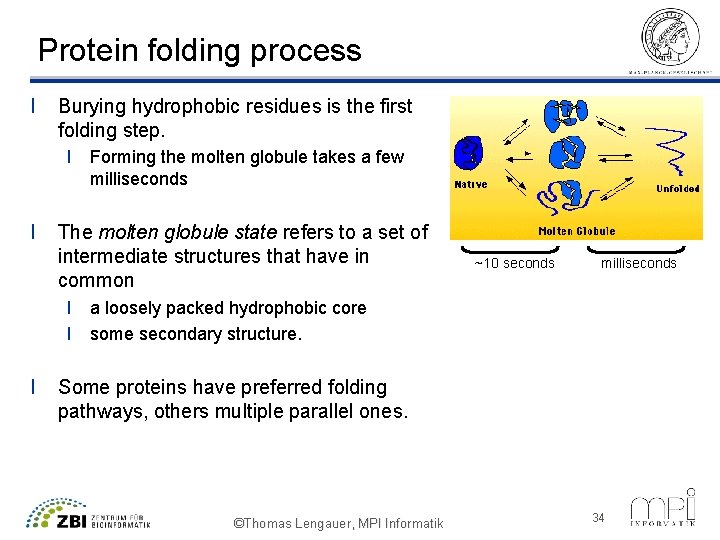
Protein folding process l Burying hydrophobic residues is the first folding step. l Forming the molten globule takes a few milliseconds l The molten globule state refers to a set of intermediate structures that have in common ~10 seconds milliseconds l a loosely packed hydrophobic core l some secondary structure. l Some proteins have preferred folding pathways, others multiple parallel ones. ©Thomas Lengauer, MPI Informatik 34
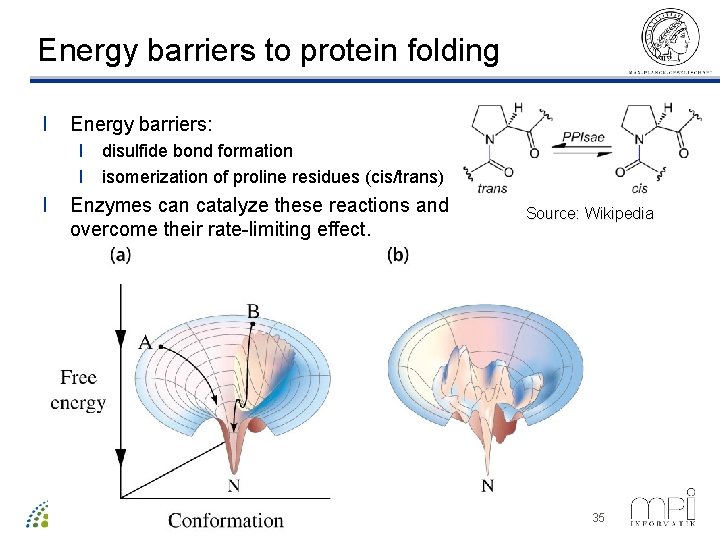
Energy barriers to protein folding l Energy barriers: l disulfide bond formation l isomerization of proline residues (cis/trans) l Enzymes can catalyze these reactions and overcome their rate-limiting effect. ©Thomas Lengauer, MPI Informatik Source: Wikipedia 35
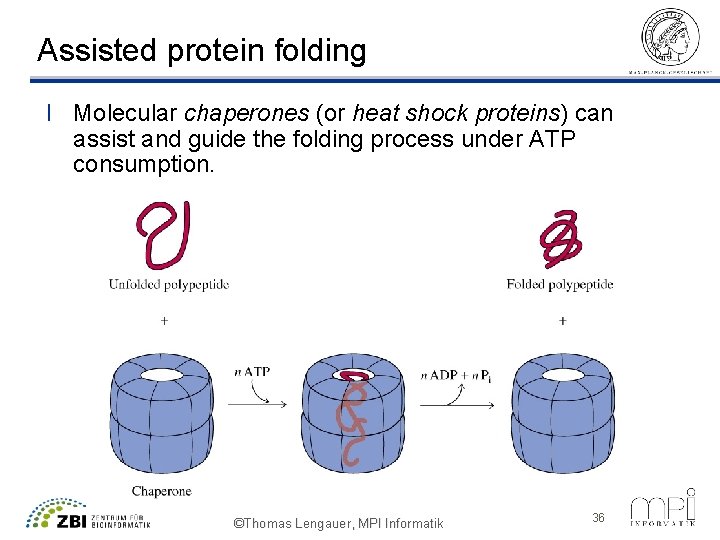
Assisted protein folding l Molecular chaperones (or heat shock proteins) can assist and guide the folding process under ATP consumption. ©Thomas Lengauer, MPI Informatik 36
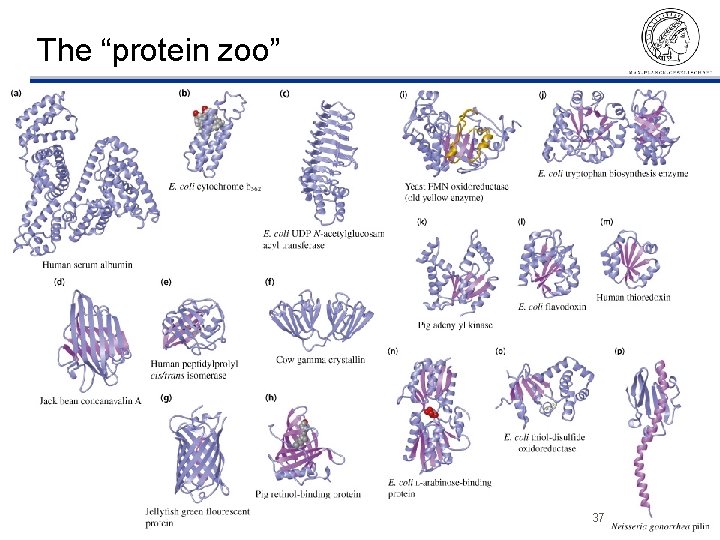
The “protein zoo” ©Thomas Lengauer, MPI Informatik 37
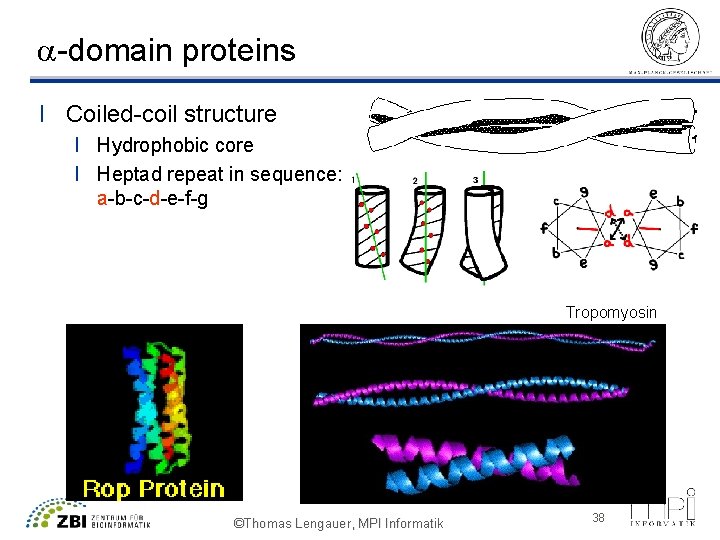
a-domain proteins l Coiled-coil structure l Hydrophobic core l Heptad repeat in sequence: a-b-c-d-e-f-g Tropomyosin ©Thomas Lengauer, MPI Informatik 38
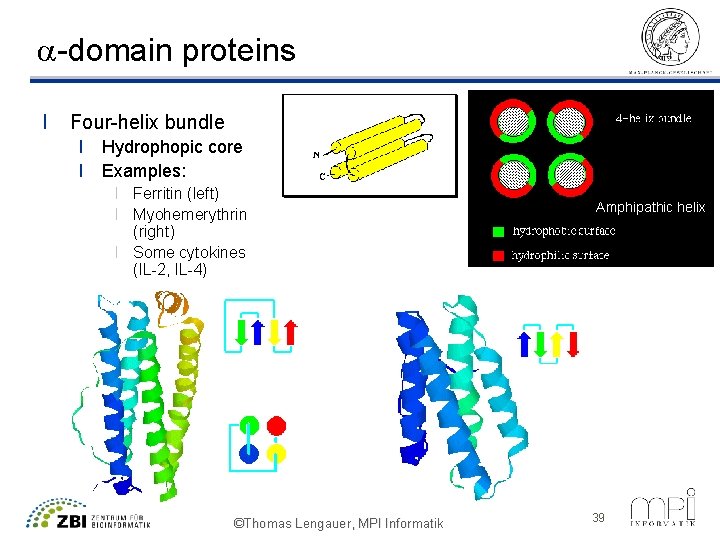
a-domain proteins l Four-helix bundle l Hydrophopic core l Examples: l Ferritin (left) l Myohemerythrin (right) l Some cytokines (IL-2, IL-4) ©Thomas Lengauer, MPI Informatik Amphipathic helix 39
a-domain proteins Amphipathic helix 40
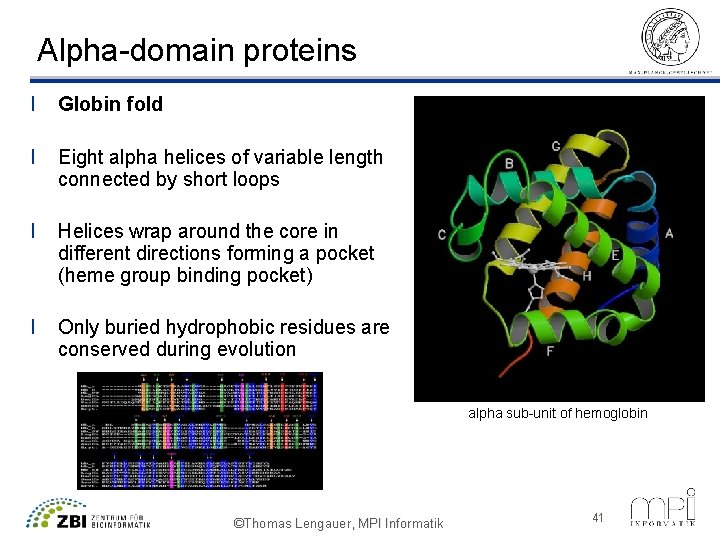
Alpha-domain proteins l Globin fold l Eight alpha helices of variable length connected by short loops l Helices wrap around the core in different directions forming a pocket (heme group binding pocket) l Only buried hydrophobic residues are conserved during evolution alpha sub-unit of hemoglobin ©Thomas Lengauer, MPI Informatik 41
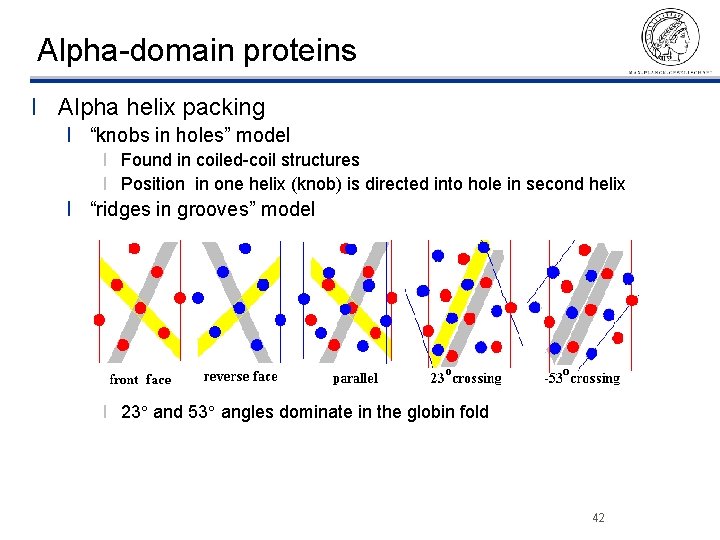
Alpha-domain proteins l Alpha helix packing l “knobs in holes” model l Found in coiled-coil structures l Position in one helix (knob) is directed into hole in second helix l “ridges in grooves” model l 23 and 53 angles dominate in the globin fold ©Thomas Lengauer, MPI Informatik 42
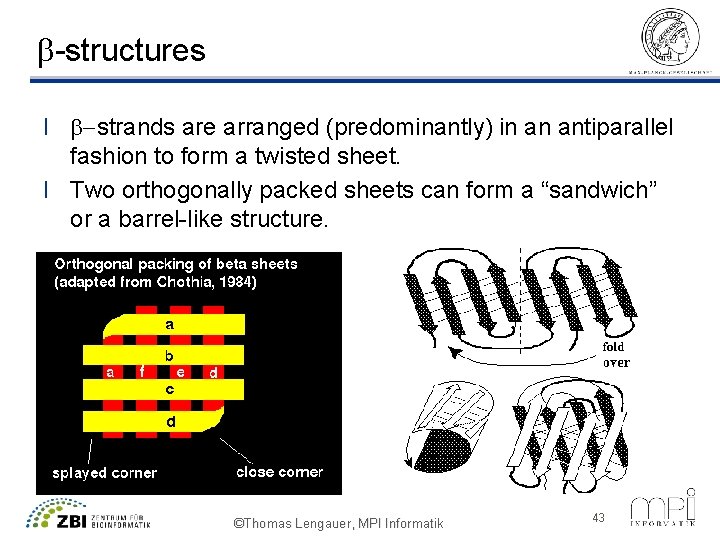
b-structures l b-strands are arranged (predominantly) in an antiparallel fashion to form a twisted sheet. l Two orthogonally packed sheets can form a “sandwich” or a barrel-like structure. ©Thomas Lengauer, MPI Informatik 43
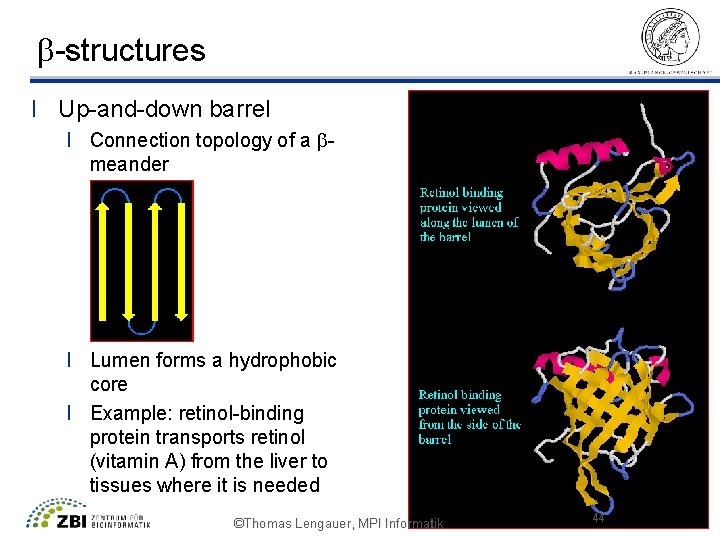
b-structures l Up-and-down barrel l Connection topology of a bmeander l Lumen forms a hydrophobic core l Example: retinol-binding protein transports retinol (vitamin A) from the liver to tissues where it is needed ©Thomas Lengauer, MPI Informatik 44
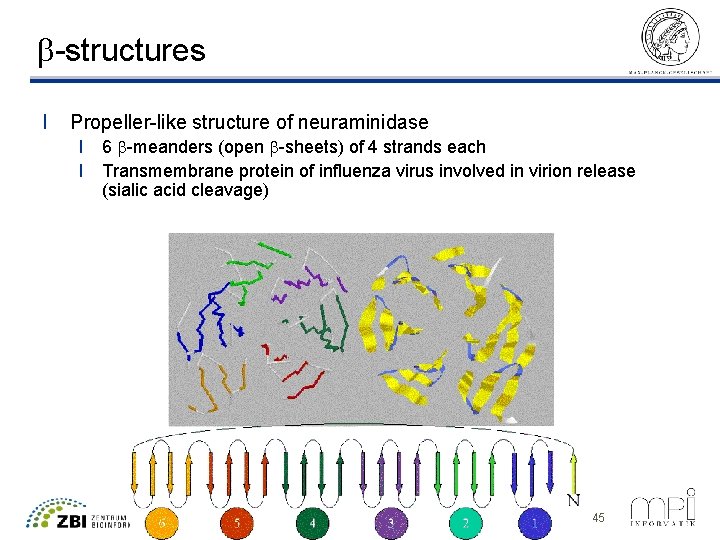
b-structures l Propeller-like structure of neuraminidase l 6 b-meanders (open b-sheets) of 4 strands each l Transmembrane protein of influenza virus involved in virion release (sialic acid cleavage) ©Thomas Lengauer, MPI Informatik 45
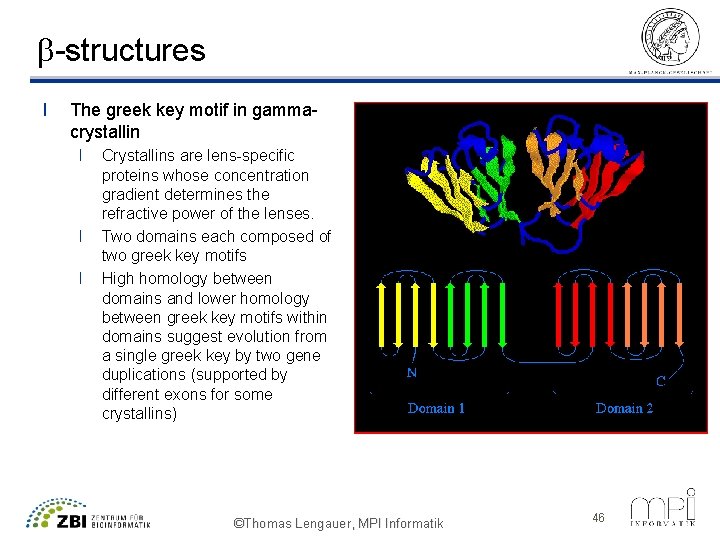
b-structures l The greek key motif in gammacrystallin l l l Crystallins are lens-specific proteins whose concentration gradient determines the refractive power of the lenses. Two domains each composed of two greek key motifs High homology between domains and lower homology between greek key motifs within domains suggest evolution from a single greek key by two gene duplications (supported by different exons for some crystallins) ©Thomas Lengauer, MPI Informatik 46
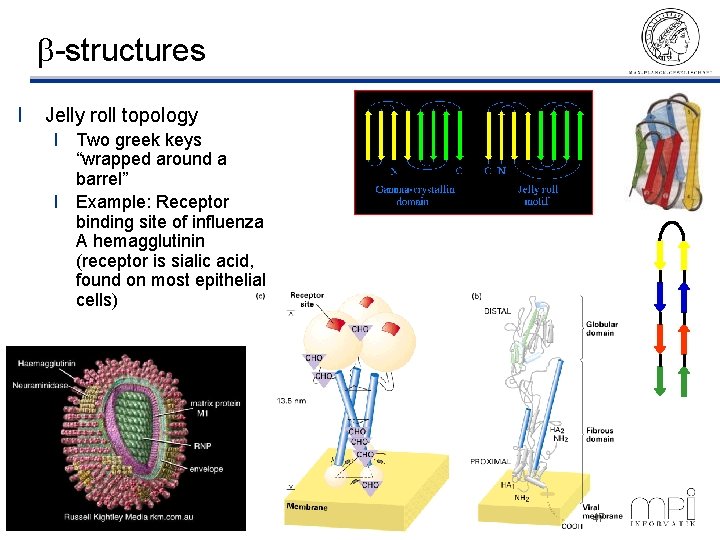
b-structures l Jelly roll topology l Two greek keys “wrapped around a barrel” l Example: Receptor binding site of influenza A hemagglutinin (receptor is sialic acid, found on most epithelial cells) ©Thomas Lengauer, MPI Informatik 47
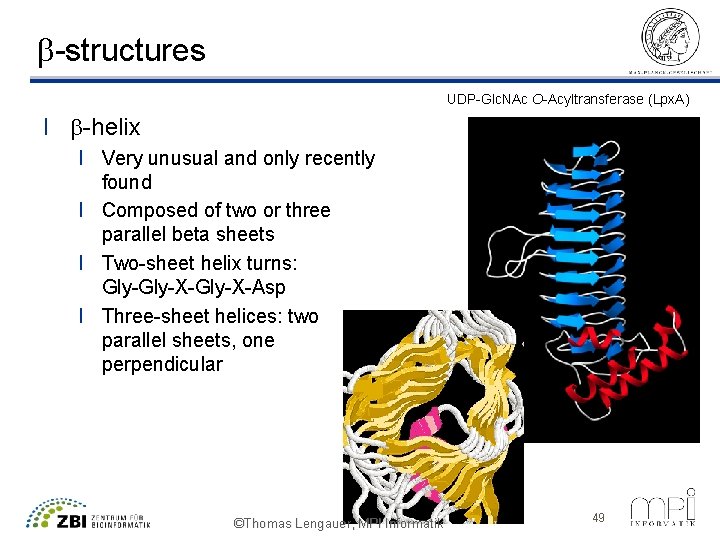
b-structures UDP-Glc. NAc O-Acyltransferase (Lpx. A) l b-helix l Very unusual and only recently found l Composed of two or three parallel beta sheets l Two-sheet helix turns: Gly-X-Gly-X-Asp l Three-sheet helices: two parallel sheets, one perpendicular ©Thomas Lengauer, MPI Informatik 49
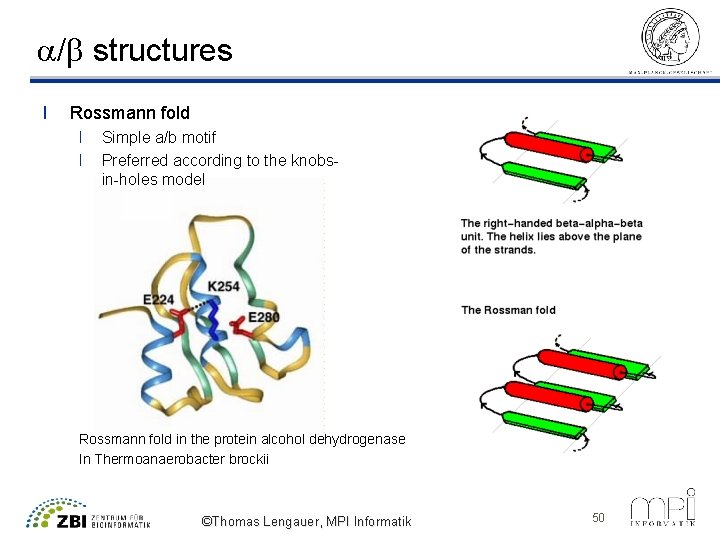
a/b structures l Rossmann fold l l Simple a/b motif Preferred according to the knobsin-holes model Rossmann fold in the protein alcohol dehydrogenase In Thermoanaerobacter brockii ©Thomas Lengauer, MPI Informatik 50
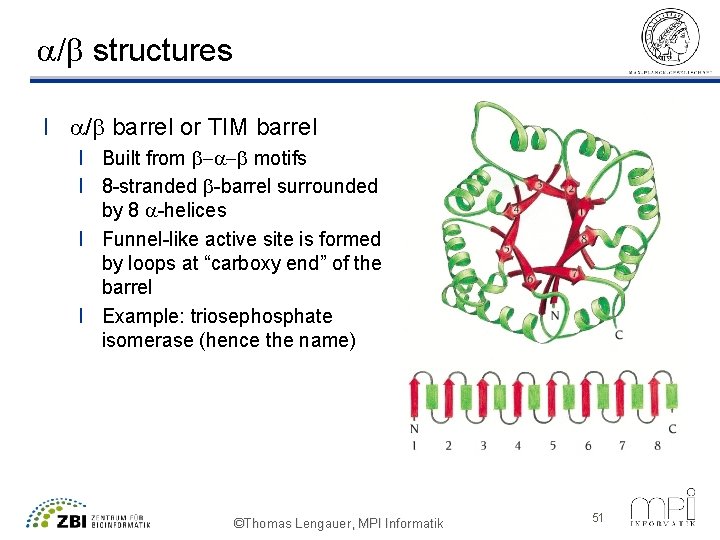
a/b structures l a/b barrel or TIM barrel l Built from b-a-b motifs l 8 -stranded b-barrel surrounded by 8 a-helices l Funnel-like active site is formed by loops at “carboxy end” of the barrel l Example: triosephosphate isomerase (hence the name) ©Thomas Lengauer, MPI Informatik 51
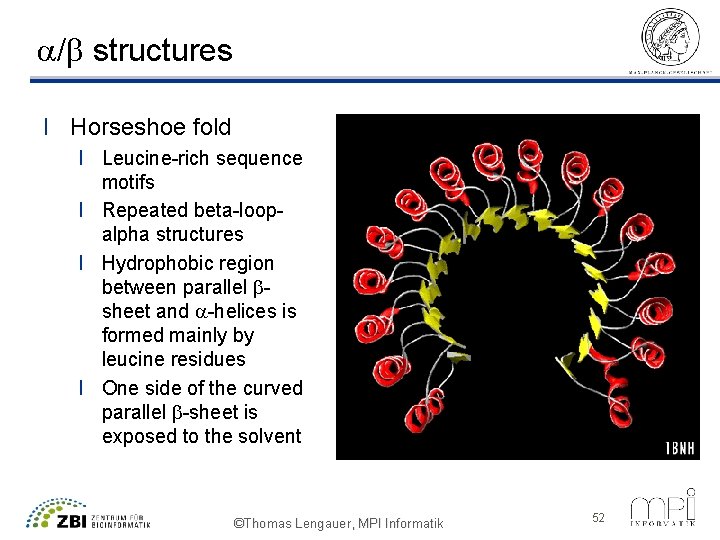
a/b structures l Horseshoe fold l Leucine-rich sequence motifs l Repeated beta-loopalpha structures l Hydrophobic region between parallel bsheet and a-helices is formed mainly by leucine residues l One side of the curved parallel b-sheet is exposed to the solvent ©Thomas Lengauer, MPI Informatik 52
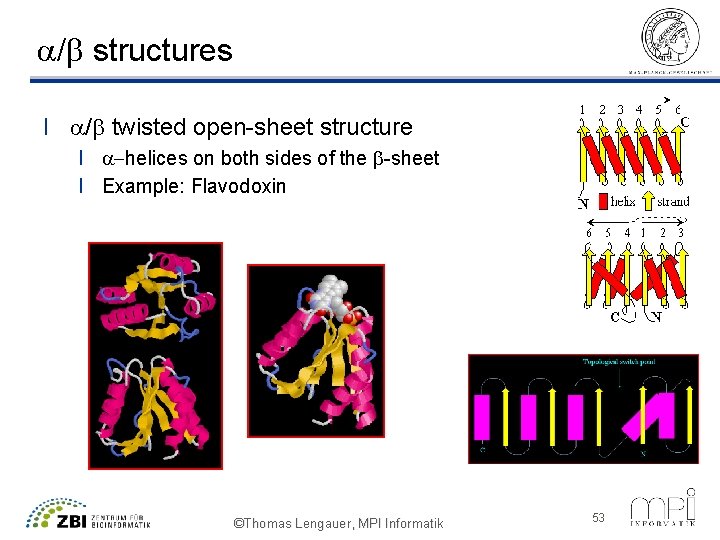
a/b structures l a/b twisted open-sheet structure l a-helices on both sides of the b-sheet l Example: Flavodoxin ©Thomas Lengauer, MPI Informatik 53

a+b structures l l a-domains, b-structures and the “wound” a/b structures do not cover all folds. The class of a+b structures collects all remaining topologies that are not easily described or classified, e. g. ©Thomas Lengauer, MPI Informatik 54
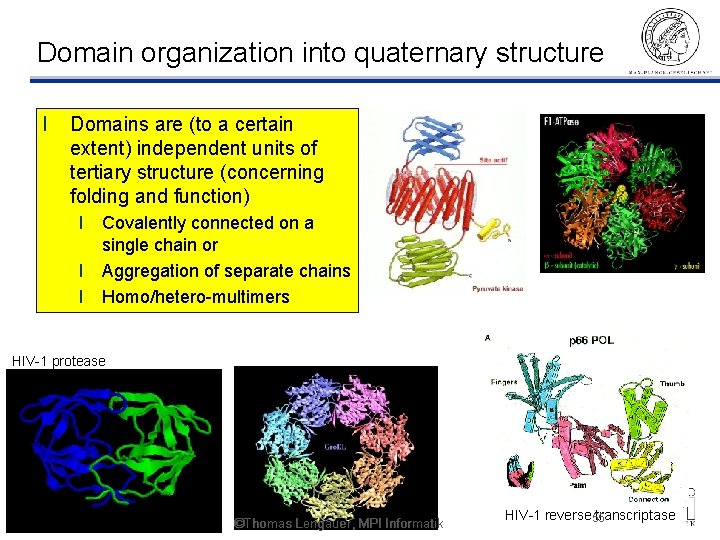
Domain organization into quaternary structure l Domains are (to a certain extent) independent units of tertiary structure (concerning folding and function) l Covalently connected on a single chain or l Aggregation of separate chains l Homo/hetero-multimers HIV-1 protease ©Thomas Lengauer, MPI Informatik HIV-1 reverse 55 transcriptase
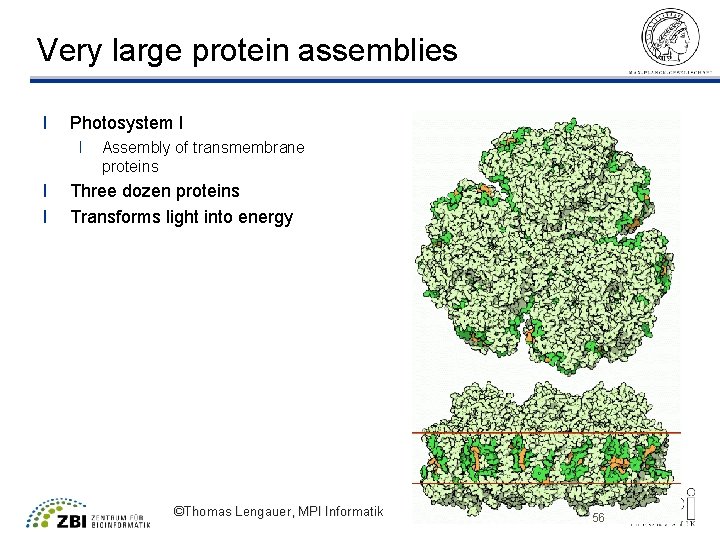
Very large protein assemblies l Photosystem I l l l Assembly of transmembrane proteins Three dozen proteins Transforms light into energy ©Thomas Lengauer, MPI Informatik 56
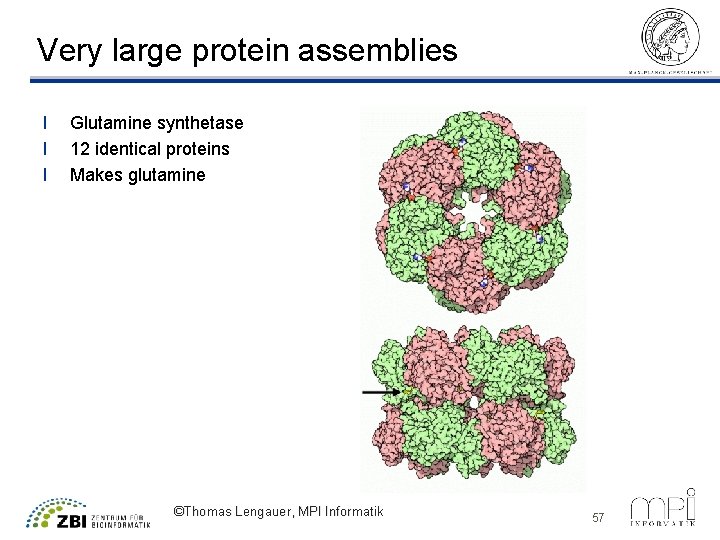
Very large protein assemblies l l l Glutamine synthetase 12 identical proteins Makes glutamine ©Thomas Lengauer, MPI Informatik 57
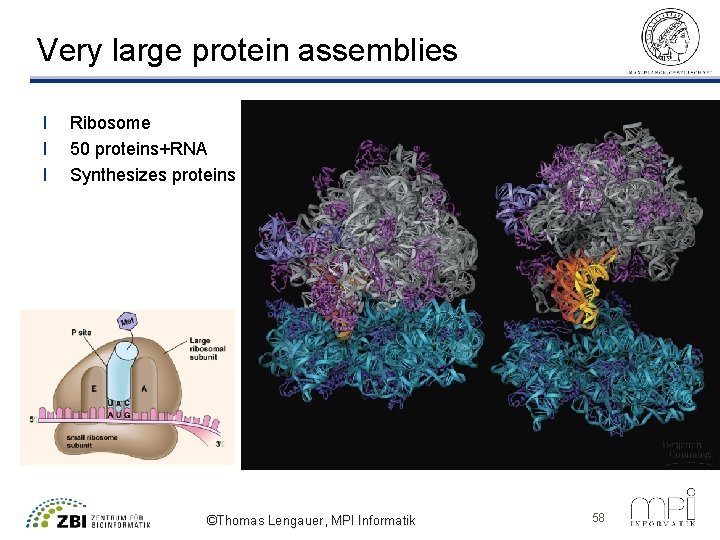
Very large protein assemblies l l l Ribosome 50 proteins+RNA Synthesizes proteins ©Thomas Lengauer, MPI Informatik 58
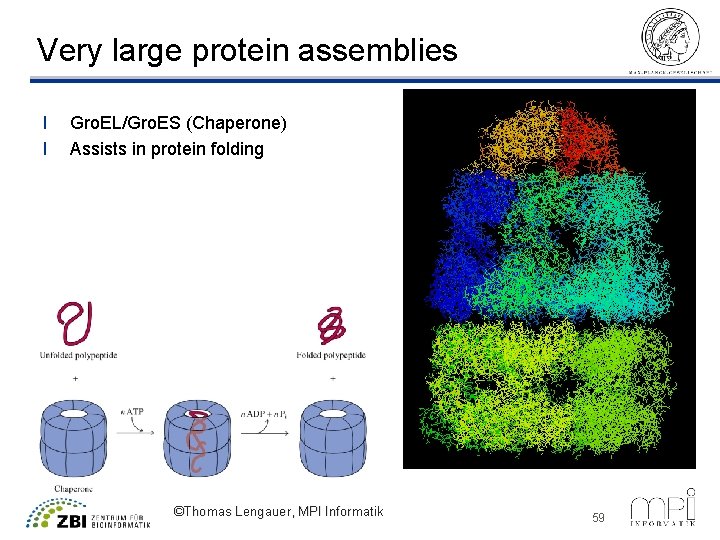
Very large protein assemblies l l Gro. EL/Gro. ES (Chaperone) Assists in protein folding ©Thomas Lengauer, MPI Informatik 59
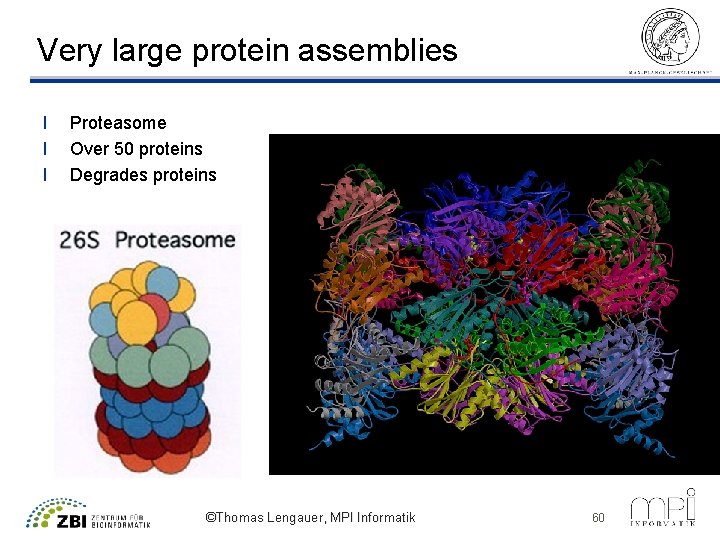
Very large protein assemblies l l l Proteasome Over 50 proteins Degrades proteins ©Thomas Lengauer, MPI Informatik 60
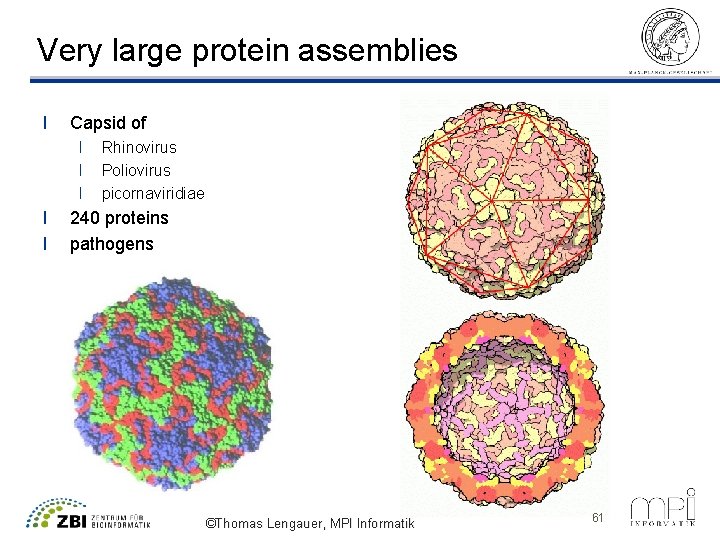
Very large protein assemblies l Capsid of l l l Rhinovirus Poliovirus picornaviridiae 240 proteins pathogens ©Thomas Lengauer, MPI Informatik 61
- Slides: 59