Part 1 Theory MPRI Bioinformatique formelle LC Standard

Part 1 : Theory MPRI - Bio-informatique formelle - LC

Standard laws of biochemical kinetics applied to molecular networks MPRI - Bio-informatique formelle - LC
![Rate of Mass Action: forward reaction Xi Biocham model: present(Xi). absent(Xa). ka*[Xi] for Xi=>Xa. Rate of Mass Action: forward reaction Xi Biocham model: present(Xi). absent(Xa). ka*[Xi] for Xi=>Xa.](http://slidetodoc.com/presentation_image_h2/d0753e3b4ba0de76be34317fd0fdb904/image-3.jpg)
Rate of Mass Action: forward reaction Xi Biocham model: present(Xi). absent(Xa). ka*[Xi] for Xi=>Xa. parameter(ka, 0. 2). MPRI - Bio-informatique formelle - LC ka Xa
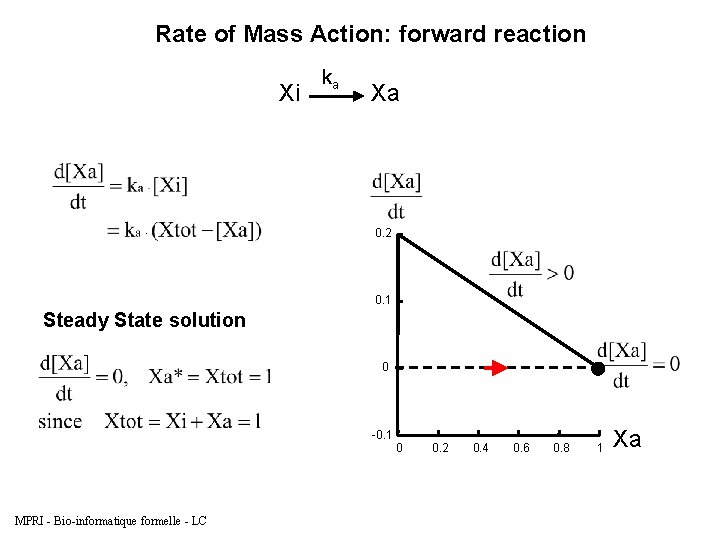
Rate of Mass Action: forward reaction Xi ka Xa 0. 2 0. 1 Steady State solution 0 -0. 1 0 MPRI - Bio-informatique formelle - LC 0. 2 0. 4 0. 6 0. 8 1 Xa
![Rate of Mass Action: reversible reaction Xi Biocham model: present(Xi). absent(Xa). ka*[Xi] for Xi=>Xa. Rate of Mass Action: reversible reaction Xi Biocham model: present(Xi). absent(Xa). ka*[Xi] for Xi=>Xa.](http://slidetodoc.com/presentation_image_h2/d0753e3b4ba0de76be34317fd0fdb904/image-5.jpg)
Rate of Mass Action: reversible reaction Xi Biocham model: present(Xi). absent(Xa). ka*[Xi] for Xi=>Xa. ki*[Xa] for Xa=>Xi. parameter(ka, 0. 2). parameter(ki, 0. 1). MPRI - Bio-informatique formelle - LC ka ki Xa
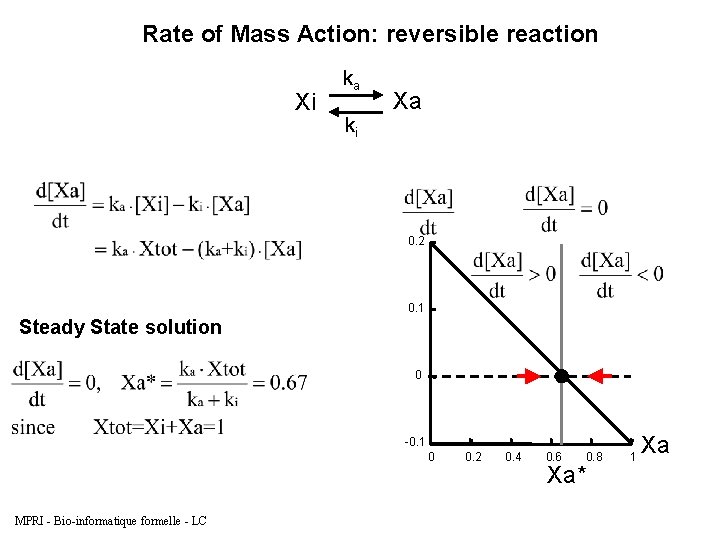
Rate of Mass Action: reversible reaction Xi ka ki Xa 0. 2 0. 1 Steady State solution 0 -0. 1 0 MPRI - Bio-informatique formelle - LC 0. 2 0. 4 0. 6 0. 8 Xa* 1 Xa
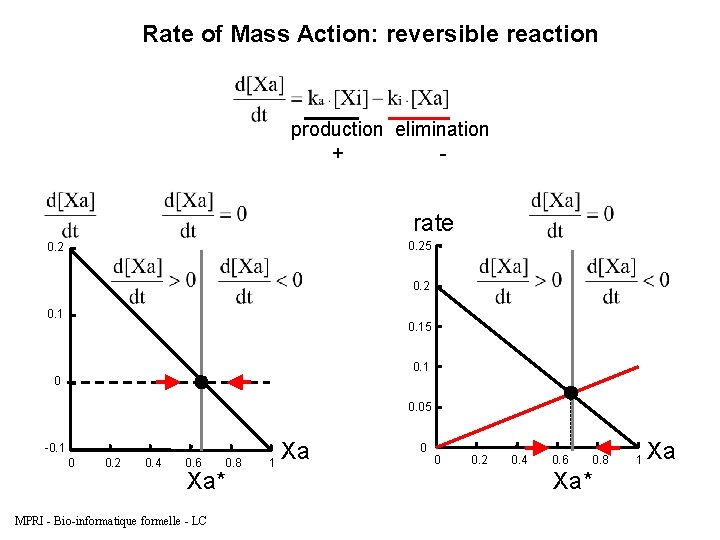
Rate of Mass Action: reversible reaction production elimination + - rate 0. 25 0. 2 0. 15 0. 1 0 0. 05 -0. 1 0 0. 2 0. 4 0. 6 0. 8 Xa* MPRI - Bio-informatique formelle - LC Xa 1 0 0 0. 2 0. 4 0. 6 0. 8 Xa* 1 Xa
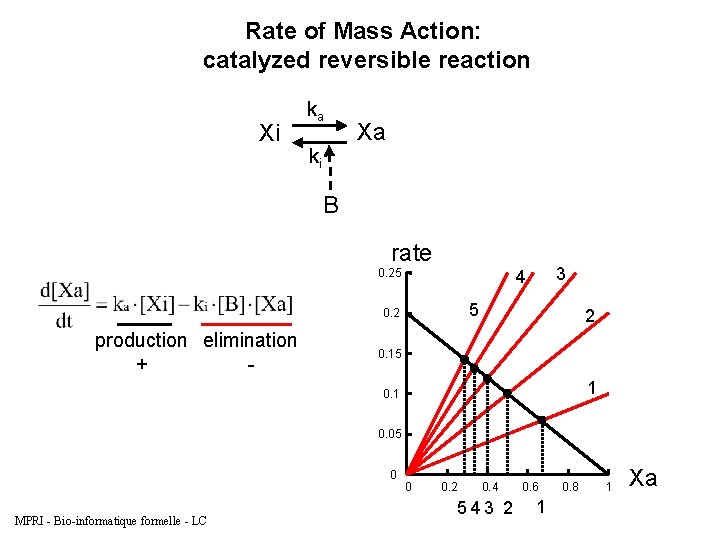
Rate of Mass Action: catalyzed reversible reaction Xi ka ki Xa B rate 0. 25 5 0. 2 production elimination + - 3 4 2 0. 15 1 0. 05 0 MPRI - Bio-informatique formelle - LC 0 0. 2 0. 4 543 2 0. 6 1 0. 8 1 Xa
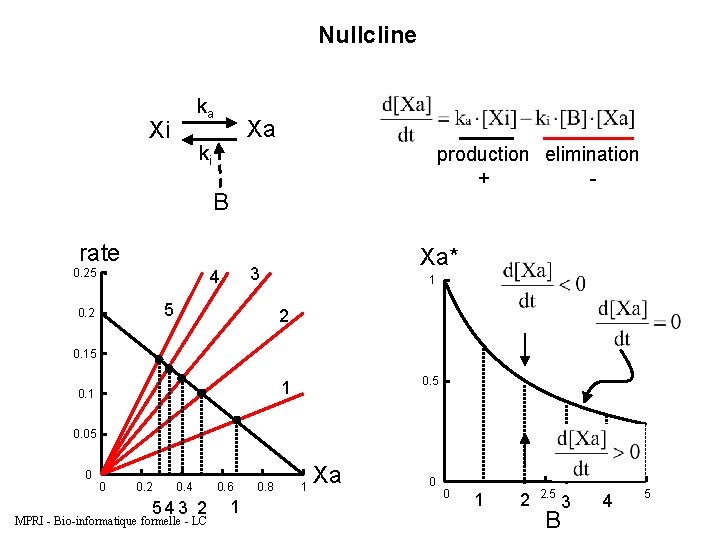
Nullcline ka Xi Xa ki production elimination + - B rate 0. 25 4 5 0. 2 Xa* 3 1 2 0. 15 0. 5 1 0. 05 0 0 0. 2 0. 4 543 2 MPRI - Bio-informatique formelle - LC 0. 6 1 0. 8 1 Xa 0 0 1 2 2. 5 B 3 4 5
![Michaelis-Menten: forward reaction Xi Biocham model: present(Xi). absent(Xa). ka*[Xi]/(Ja+[Xi]) for Xi=>Xa. parameter(ka, 0. 2). Michaelis-Menten: forward reaction Xi Biocham model: present(Xi). absent(Xa). ka*[Xi]/(Ja+[Xi]) for Xi=>Xa. parameter(ka, 0. 2).](http://slidetodoc.com/presentation_image_h2/d0753e3b4ba0de76be34317fd0fdb904/image-10.jpg)
Michaelis-Menten: forward reaction Xi Biocham model: present(Xi). absent(Xa). ka*[Xi]/(Ja+[Xi]) for Xi=>Xa. parameter(ka, 0. 2). parameter(Ja, 0. 05). MPRI - Bio-informatique formelle - LC ka Xa
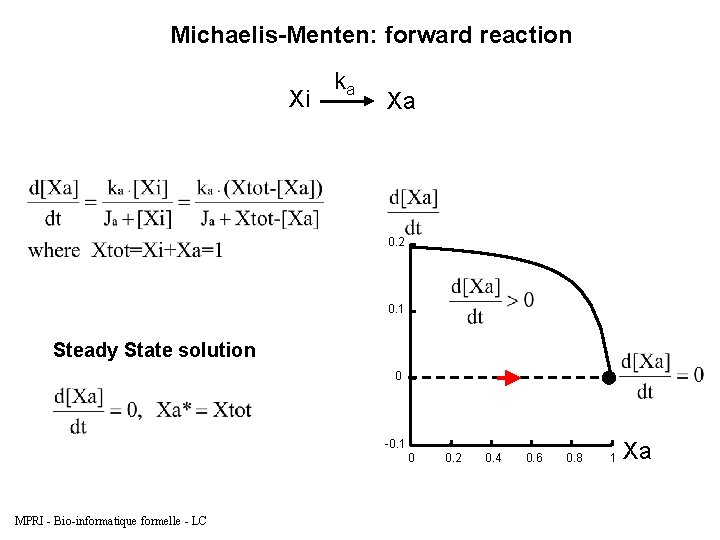
Michaelis-Menten: forward reaction Xi ka Xa 0. 2 0. 1 Steady State solution 0 -0. 1 0 MPRI - Bio-informatique formelle - LC 0. 2 0. 4 0. 6 0. 8 1 Xa
![Michaelis-Menten: reverse reaction Xi ka ki Xa Biocham model: present(Xi). absent(Xa). ka*[Xi]/(Ja+[Xi]) for Xi=>Xa. Michaelis-Menten: reverse reaction Xi ka ki Xa Biocham model: present(Xi). absent(Xa). ka*[Xi]/(Ja+[Xi]) for Xi=>Xa.](http://slidetodoc.com/presentation_image_h2/d0753e3b4ba0de76be34317fd0fdb904/image-12.jpg)
Michaelis-Menten: reverse reaction Xi ka ki Xa Biocham model: present(Xi). absent(Xa). ka*[Xi]/(Ja+[Xi]) for Xi=>Xa. ki*[Xa]/(Ji+[Xa]) for Xa=>Xi. parameter(ka, 0. 2). parameter(ki, 0. 1). parameter(Ja, 0. 05). parameter(Ji, 0. 05). MPRI - Bio-informatique formelle - LC Þ Goldbeter-Koshland switch
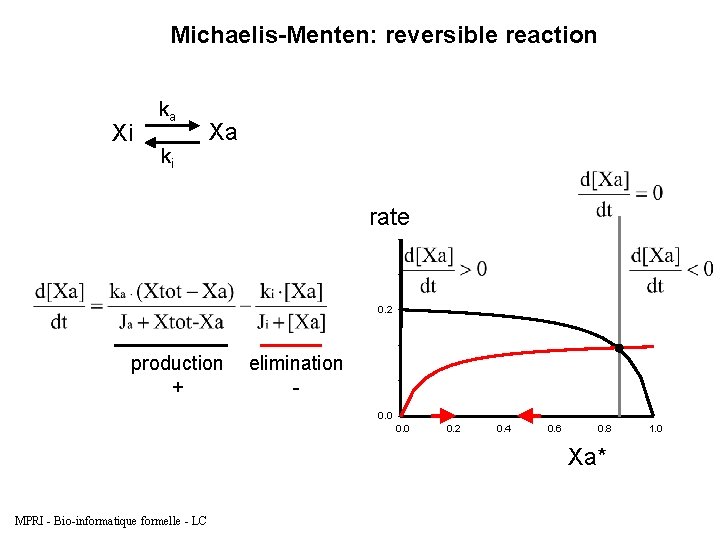
Michaelis-Menten: reversible reaction Xi ka ki Xa rate 0. 2 production + elimination 0. 0 0. 2 0. 4 0. 6 0. 8 Xa* MPRI - Bio-informatique formelle - LC 1. 0
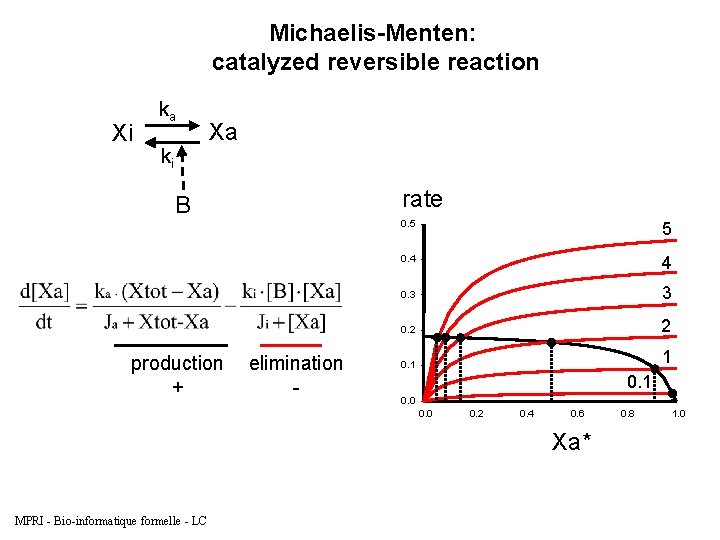
Michaelis-Menten: catalyzed reversible reaction Xi ka ki Xa rate B production + elimination - 0. 5 5 0. 4 4 0. 3 3 0. 2 2 0. 1 1 0. 0 0. 2 0. 4 0. 6 Xa* MPRI - Bio-informatique formelle - LC 0. 8 1. 0

Nullcline Xi ka Xa ki production + B Xa* rate 1 0. 5 5 0. 4 4 0. 3 3 0. 6 0. 2 2 0. 4 0. 1 1 0. 8 0. 2 . 1 0. 0 elimination - 0. 2 0. 4 0. 6 MPRI - Bio-informatique formelle - LC 0. 8 1. 0 Xa* 0 0 1 2 3 [B] 4 5
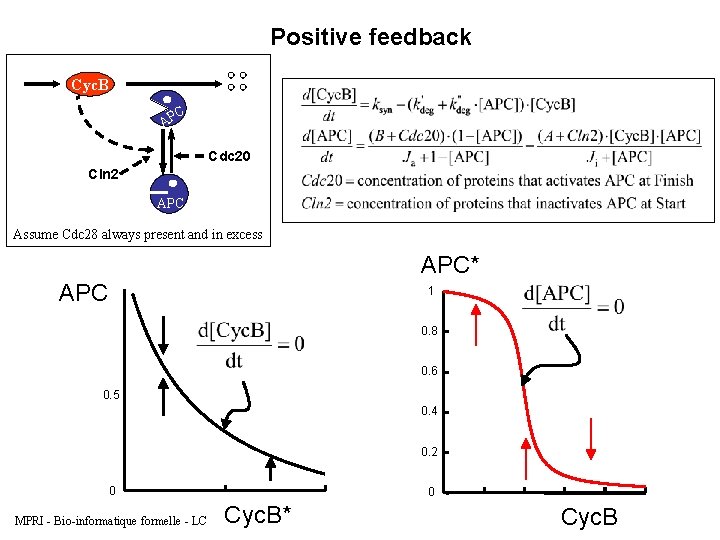
Positive feedback Cyc. B C AP Cdc 20 Cln 2 APC Assume Cdc 28 always present and in excess APC* APC 1 0. 8 0. 6 0. 5 0. 4 0. 2 0 MPRI - Bio-informatique formelle - LC 0 Cyc. B* Cyc. B
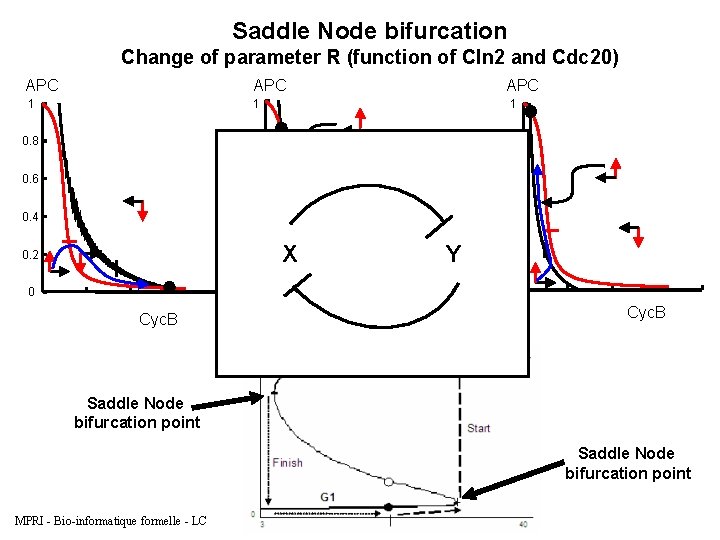
Saddle Node bifurcation Change of parameter R (function of Cln 2 and Cdc 20) APC 1 1 1 0. 8 0. 6 0. 4 0. 2 0 0 Cyc. B APC X Y 0. 2 0 Cyc. B Saddle Node bifurcation point MPRI - Bio-informatique formelle - LC

Negative feedback E T A L L I X N N T O A C MPRI - Bio-informatique formelle - LC O C S Y
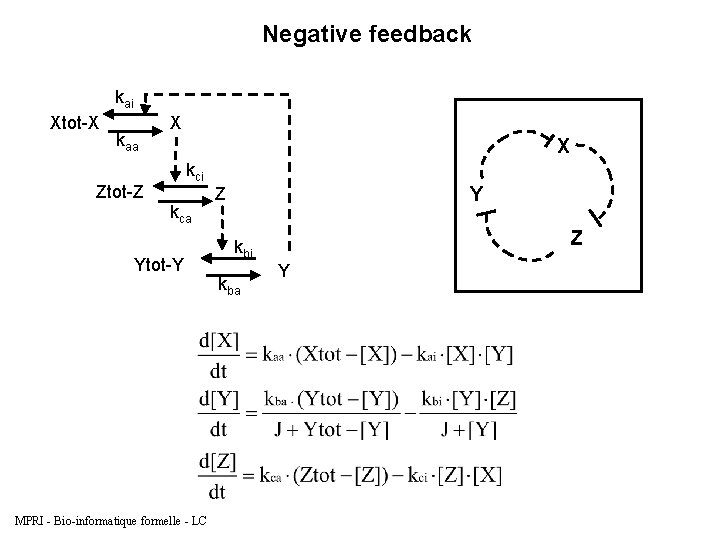
Negative feedback kai Xtot-X kaa X X kci Ztot-Z kca Ytot-Y MPRI - Bio-informatique formelle - LC Y Z Z kbi kba Y
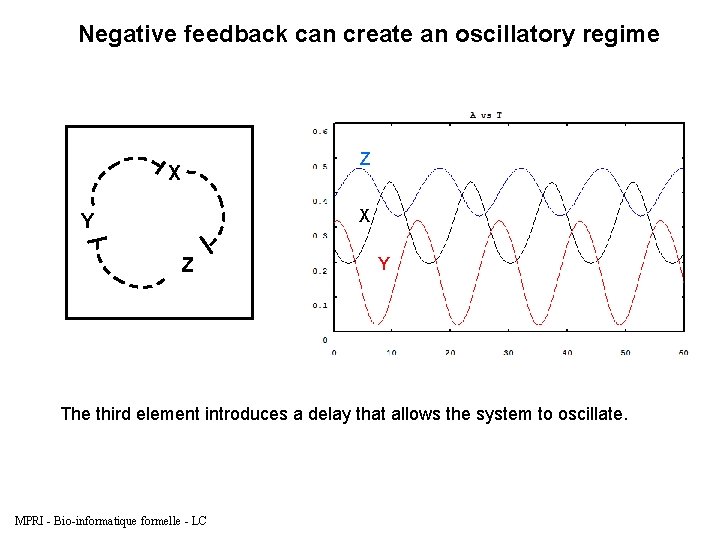
Negative feedback can create an oscillatory regime Z X X Y Z Y The third element introduces a delay that allows the system to oscillate. MPRI - Bio-informatique formelle - LC
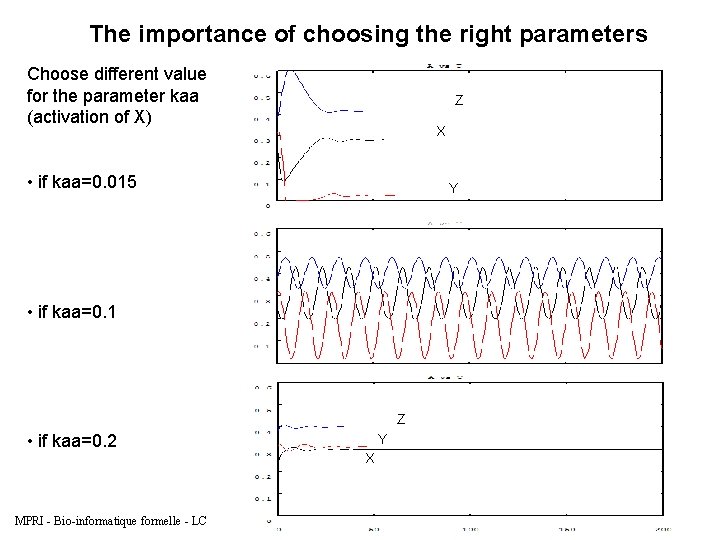
The importance of choosing the right parameters Choose different values for the parameter kaa (activation of X) Z X • if kaa=0. 015 Y • if kaa=0. 1 Z • if kaa=0. 2 MPRI - Bio-informatique formelle - LC Y X
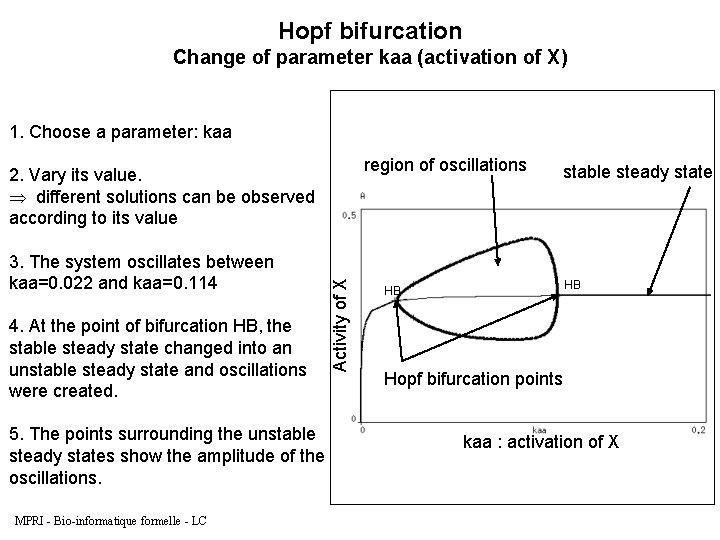
Hopf bifurcation Change of parameter kaa (activation of X) 1. Choose a parameter: kaa region of oscillations 3. The system oscillates between kaa=0. 022 and kaa=0. 114 4. At the point of bifurcation HB, the stable steady state changed into an unstable steady state and oscillations were created. 5. The points surrounding the unstable steady states show the amplitude of the oscillations. MPRI - Bio-informatique formelle - LC Activity of X 2. Vary its value. Þ different solutions can be observed according to its value stable steady state HB HB Hopf bifurcation points kaa : activation of X

Introduction to bifurcation theory 1. Saddle Node (SN) bifurcation 2. Hopf (H) bifurcation 3. SNIC bifurcation : when SN meets H 4. Numerical Bifurcation theory 5. Signature of bifurcations MPRI - Bio-informatique formelle - LC

Basic Definitions Bifurcation : Qualitative change in dynamics of the solutions of a system Bifurcation point : Border line between two behaviours of solutions MPRI - Bio-informatique formelle - LC
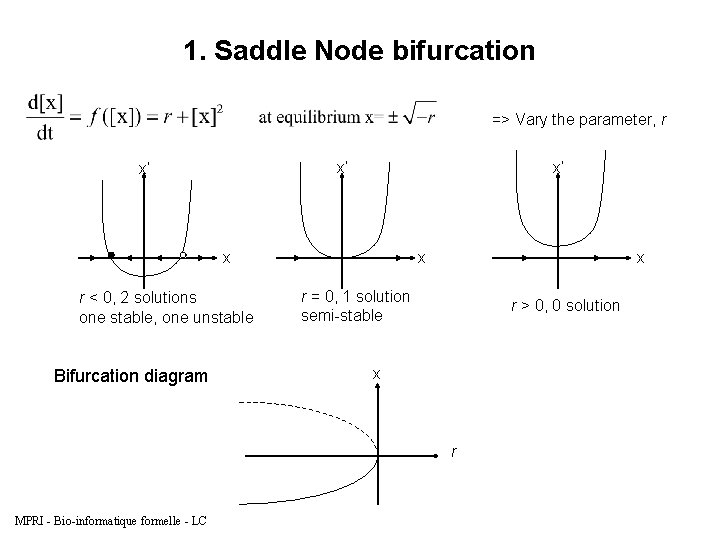
1. Saddle Node bifurcation => Vary the parameter, r x’ x’ x’ x r < 0, 2 solutions one stable, one unstable Bifurcation diagram x x r = 0, 1 solution semi-stable r > 0, 0 solution x r MPRI - Bio-informatique formelle - LC
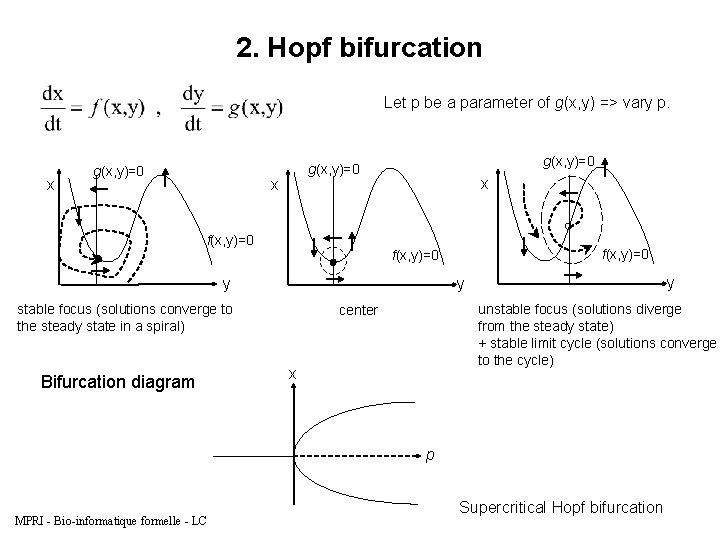
2. Hopf bifurcation Let p be a parameter of g(x, y) => vary p. x g(x, y)=0 x f(x, y)=0 x y y y stable focus (solutions converge to the steady state in a spiral) Bifurcation diagram f(x, y)=0 unstable focus (solutions diverge from the steady state) + stable limit cycle (solutions converge to the cycle) center x p MPRI - Bio-informatique formelle - LC Supercritical Hopf bifurcation
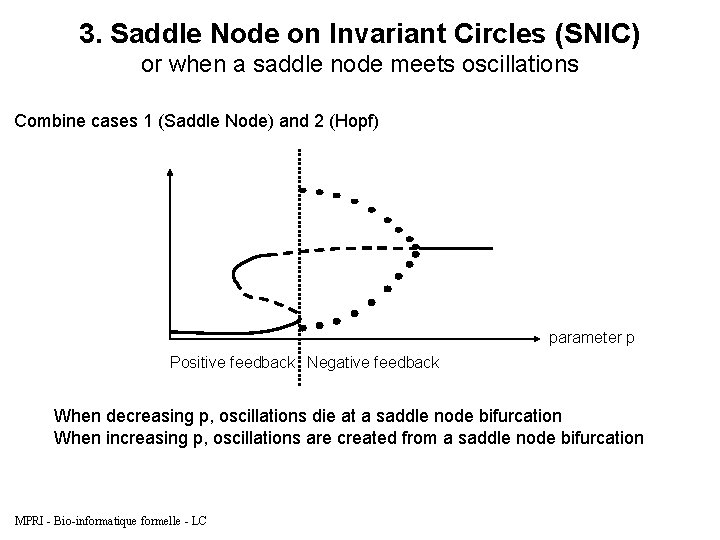
3. Saddle Node on Invariant Circles (SNIC) or when a saddle node meets oscillations Combine cases 1 (Saddle Node) and 2 (Hopf) parameter p Positive feedback Negative feedback When decreasing p, oscillations die at a saddle node bifurcation When increasing p, oscillations are created from a saddle node bifurcation MPRI - Bio-informatique formelle - LC

4. Numerical bifurcation theory How to solve numerically a system of n ODEs : the case of n=2 1. Consider the following system of ODEs: 2. Solve at the equilibrium and determine the fixed points: 3. Determine the stability of the fixed points by computing the Jacobian A at these values (Jacobian is the matrix of the partial derivatives of the functions with respect to the components computed at the fixed points) MPRI - Bio-informatique formelle - LC

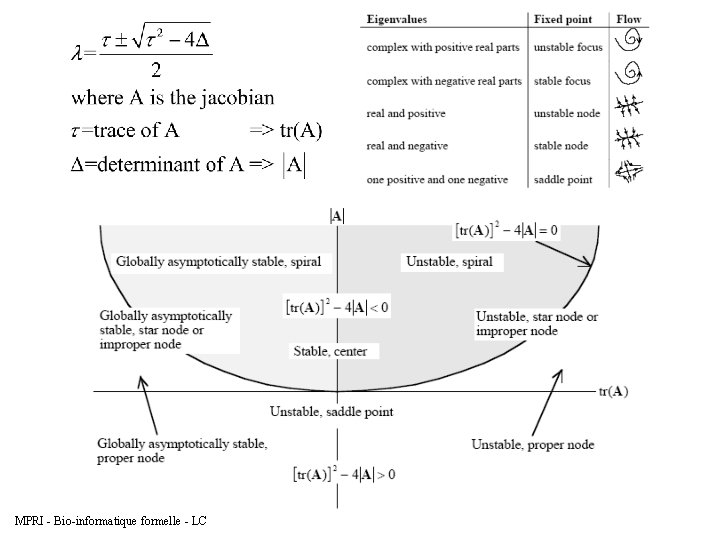
4. Compute the characteristic equation in terms of the eigenvalues λ and where the equation is determined as follows: The solution of the equation is the following: 5. The eigenvalues can inform on the stability of the fixed points MPRI - Bio-informatique formelle - LC
MPRI - Bio-informatique formelle - LC

5. Signature of bifurcations MPRI - Bio-informatique formelle - LC

Continuation of a saddle node in one-parameter – One parameter bifurcation graph Example of a system of 2 ODEs Þ 2 equations, 3 unknowns. Fix p=p* and solve for the steady state (x 1, x 2). We seek an equation of x (either 1 or 2) in terms of p. That way, we can follow a steady state as a parameter changes. MPRI - Bio-informatique formelle - LC
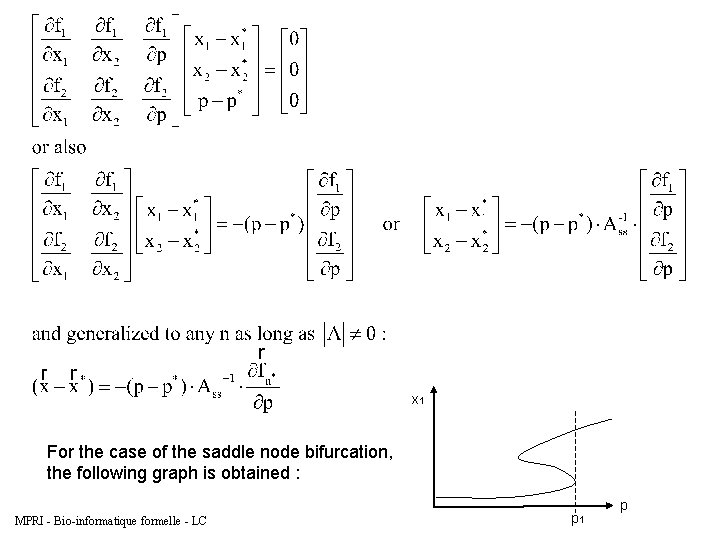
x 1 For the case of the saddle node bifurcation, the following graph is obtained : MPRI - Bio-informatique formelle - LC p 1 p

Part 2 : application to biology MPRI - Bio-informatique formelle - LC

Quelques faits - 13 cycles rapides et synchronisés juste après fécondation - Alternance entre les phases S et M (sans G 1 ni G 2) - 6000 noyaux partagent le même cytoplasme - Le niveau total des cyclines n’oscille qu’après le cycle 8 ou 9 - En interphase du cycle 14, arrêt en G 2 Quelquestions - Pourquoi ne voit-on pas le niveau des cyclines osciller plus tôt puisqu’il y a division nucléaire ? - Pourquoi les cycles s’arrêtent-ils au 14 e cycle ? MPRI - Bio-informatique formelle - LC
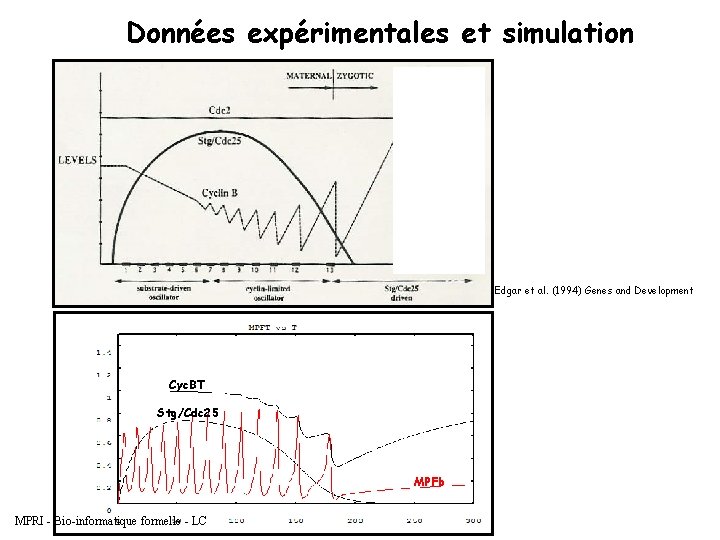
Données expérimentales et simulation Edgar et al. (1994) Genes and Development Cyc. BT Stg/Cdc 25 MPFb MPRI - Bio-informatique formelle - LC

Pourquoi ne voit-on pas le niveau des cyclines osciller plus tôt puisqu’il y a division nucléaire ? MPRI - Bio-informatique formelle - LC
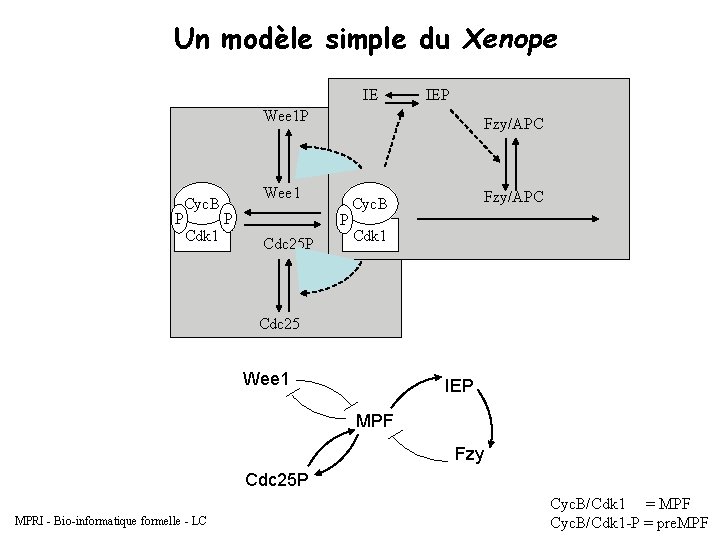
Un modèle simple du Xenope IE IEP Wee 1 P P Cyc. B Cdk 1 Fzy/APC Wee 1 P P Cdc 25 P Fzy/APC Cyc. B Cdk 1 Cdc 25 Wee 1 IEP MPF Fzy Cdc 25 P MPRI - Bio-informatique formelle - LC Cyc. B/Cdk 1 = MPF Cyc. B/Cdk 1 -P = pre. MPF
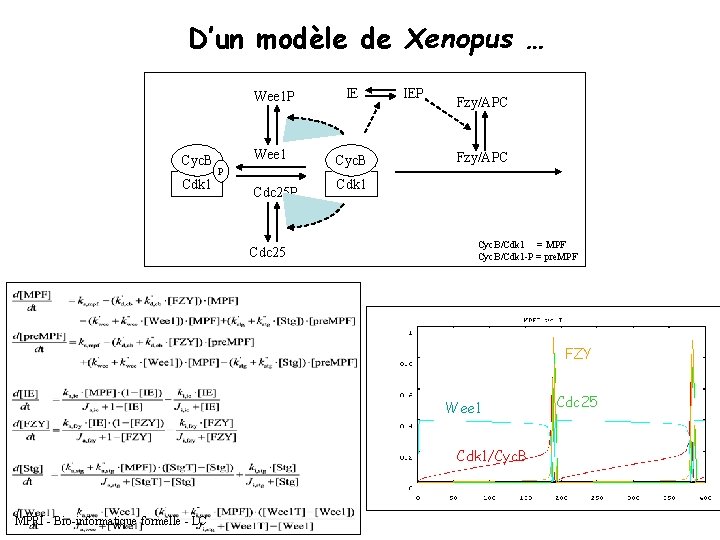
D’un modèle de Xenopus … Cyc. B Cdk 1 Wee 1 P IE Wee 1 Cyc. B P Cdc 25 IEP Fzy/APC Cdk 1 Cyc. B/Cdk 1 = MPF Cyc. B/Cdk 1 -P = pre. MPF FZY Wee 1 Cdk 1/Cyc. B MPRI - Bio-informatique formelle - LC Cdc 25
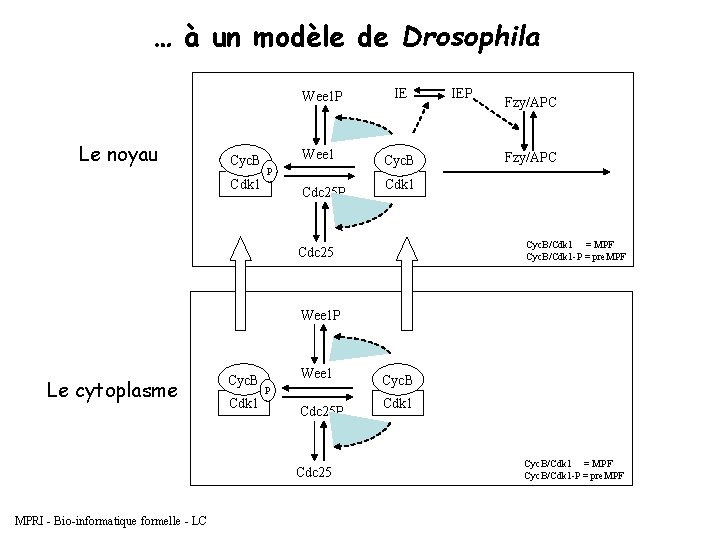
… à un modèle de Drosophila Le noyau Cyc. B Cdk 1 Wee 1 P IE Wee 1 Cyc. B P Cdc 25 P IEP Fzy/APC Cdk 1 Cyc. B/Cdk 1 = MPF Cyc. B/Cdk 1 -P = pre. MPF Cdc 25 Wee 1 P Le cytoplasme Cyc. B Cdk 1 Wee 1 P Cdc 25 MPRI - Bio-informatique formelle - LC Cyc. B Cdk 1 Cyc. B/Cdk 1 = MPF Cyc. B/Cdk 1 -P = pre. MPF
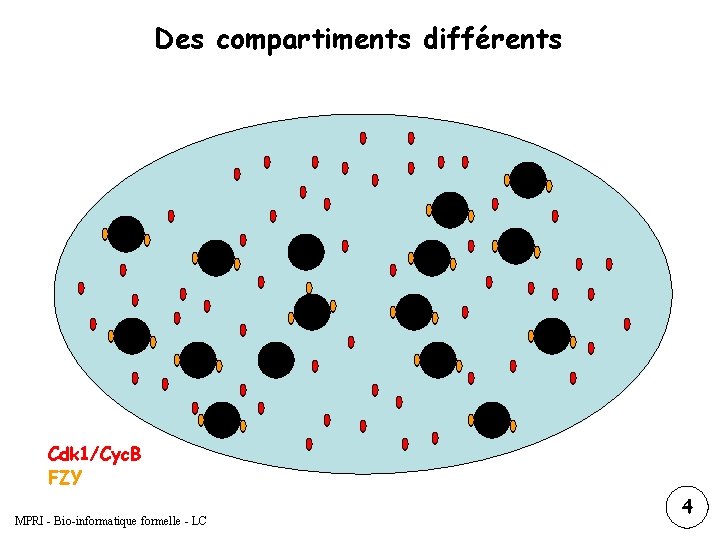
Des compartiments différents Cdk 1/Cyc. B FZY MPRI - Bio-informatique formelle - LC 1 4 3 2
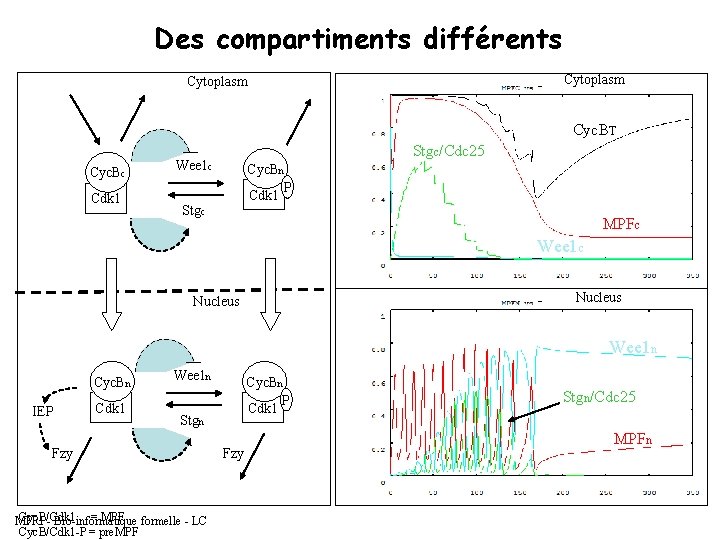
Des compartiments différents Cytoplasm Cyc. BT Cyc. Bc Cdk 1 Stgc/Cdc 25 Wee 1 c Cyc. Bn P Cdk 1 Stgc MPFc Wee 1 c Nucleus Wee 1 n Cyc. Bn IEP Cdk 1 Wee 1 n Cyc. Bn P Cdk 1 Stgn Fzy Cyc. B/Cdk 1 = MPF formelle - LC MPRI - Bio-informatique Cyc. B/Cdk 1 -P = pre. MPF Fzy Stgn/Cdc 25 MPFn

Pourquoi les cycles s’arrêtent-ils au 14 e cycle ? MPRI - Bio-informatique formelle - LC
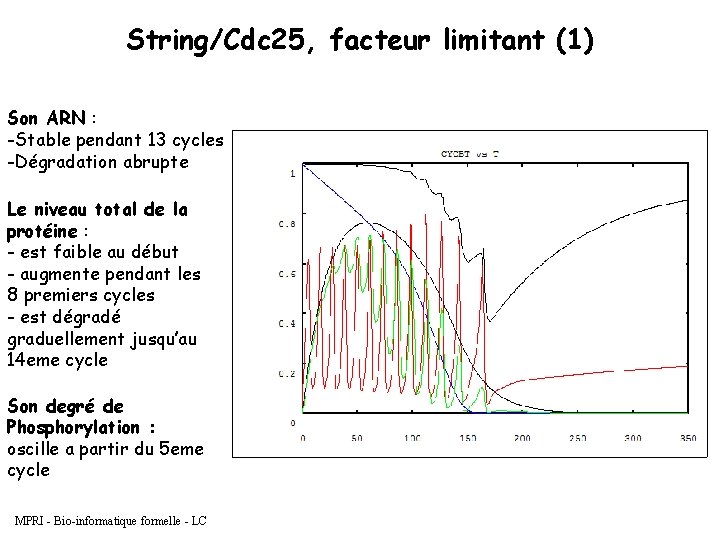
String/Cdc 25, facteur limitant (1) Son ARN : -Stable pendant 13 cycles -Dégradation abrupte Le niveau total de la protéine : - est faible au début - augmente pendant les 8 premiers cycles - est dégradé graduellement jusqu’au 14 eme cycle Son degré de Phosphorylation : oscille a partir du 5 eme cycle MPRI - Bio-informatique formelle - LC
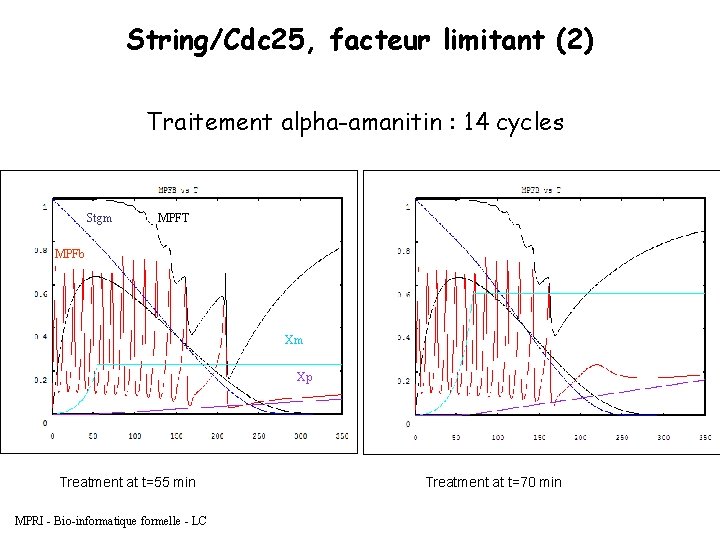
String/Cdc 25, facteur limitant (2) Traitement alpha-amanitin : 14 cycles Stgm MPFT MPFb Xm Xp Treatment at t=55 min MPRI - Bio-informatique formelle - LC Treatment at t=70 min
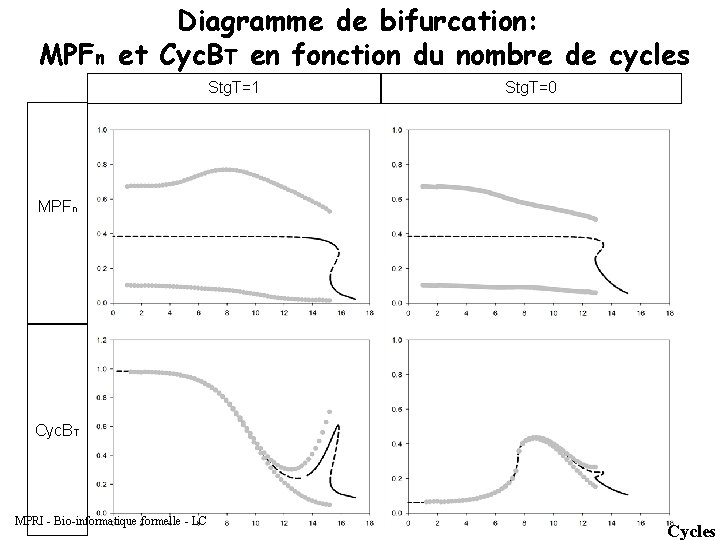
Diagramme de bifurcation: MPFn et Cyc. BT en fonction du nombre de cycles Stg. T=1 Stg. T=0 MPFn Cyc. BT MPRI - Bio-informatique formelle - LC Cycles

Ce que la théorie de la bifurcation nous permet de conclure : => String est responsable de l’endroit où se trouve le saddle node (feedback positif) Si on réduit la valeur de String, le saddle node va bouger. => Si on élimine le feedback négatif, on perd les oscillations (dans le cytoplasme, il n’y a pas de feedback negatif). MPRI - Bio-informatique formelle - LC
- Slides: 47