Parametric Bootstrap Efron 1985 Huelsenbeck et al 1996

Parametric Bootstrap (Efron 1985; Huelsenbeck et al. 1996) 1) build best tree 2) generate new simulated data sets using estimated branch lengths and other parameters (e. g. , alpha and substitution model) (from tree/data) 3) build new tree for each simulated data set 4) determine what fraction of the trees come out with each topology or generate majority rule consensus 5) assign p values
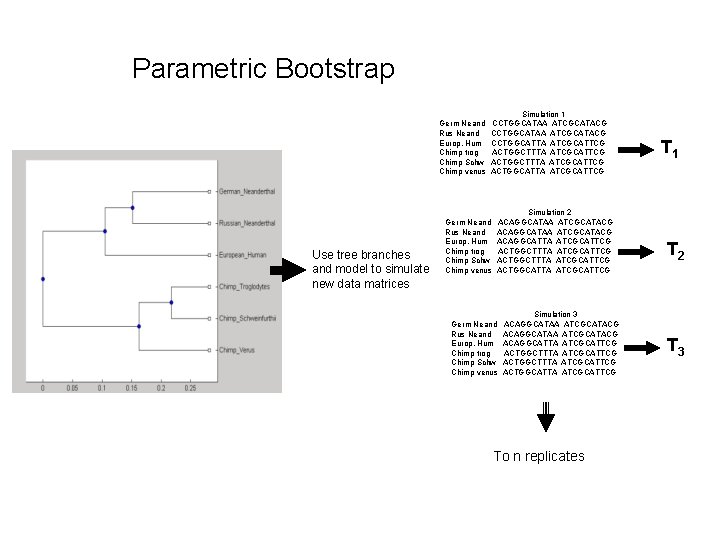
Parametric Bootstrap Germ Neand Rus Neand Europ. Hum Chimp trog Chimp Schw Chimp venus Use tree branches and model to simulate new data matrices Simulation 1 CCTGGCATAA ATCGCATACG CCTGGCATTA ATCGCATTCG ACTGGCTTTA ATCGCATTCG ACTGGCATTA ATCGCATTCG Germ Neand Rus Neand Europ. Hum Chimp trog Chimp Schw Chimp venus Simulation 2 ACAGGCATAA ATCGCATACG ACAGGCATTA ATCGCATTCG ACTGGCTTTA ATCGCATTCG ACTGGCATTA ATCGCATTCG Germ Neand Rus Neand Europ. Hum Chimp trog Chimp Schw Chimp venus Simulation 3 ACAGGCATAA ATCGCATACG ACAGGCATTA ATCGCATTCG ACTGGCTTTA ATCGCATTCG ACTGGCATTA ATCGCATTCG To n replicates T 1 T 2 T 3
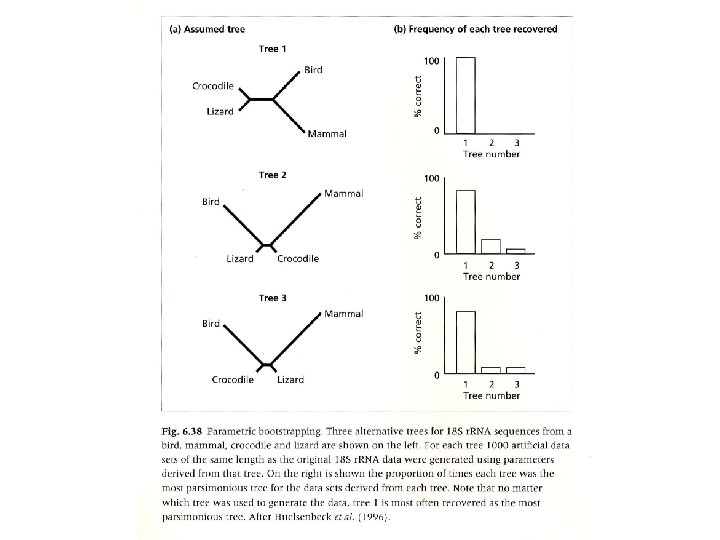

Paired-Sites Tests Examines the number of sites supporting tree 1 versus number of sites supporting tree 2 with the null model that the trees do not differ any more than would be expected by random error: “…if two trees have equal log-likelihoods, the differences in loglikelihoods at each site will be drawn independently from some distribution whose expectation is zero. If we do a statistical test of whether the mean differences is zero, we are then also testing whethere is significant statistical evidence that one tree is better than another. ” Felsenstein (2004) • Winning sites & Templeton Wilcoxon sign test • Kishino-Hasegawa (KH) • Shimodaira-Hasegawa (SH)

from Felsenstein. 2004. Inferring Phylogenies.
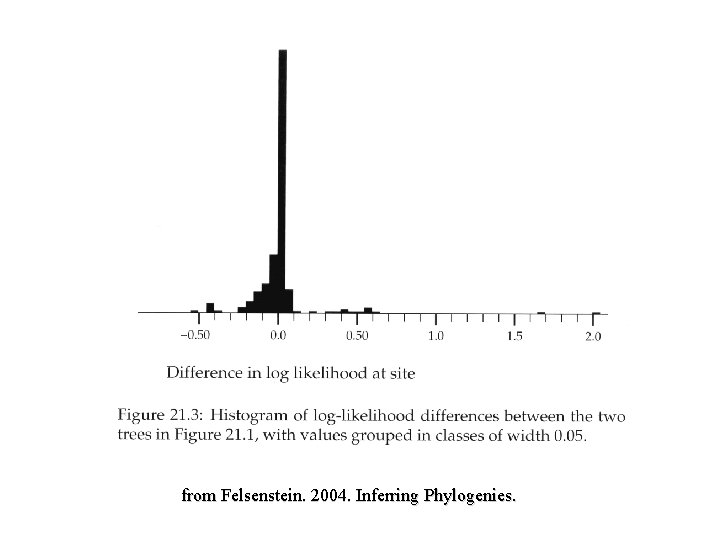
from Felsenstein. 2004. Inferring Phylogenies.

Sign Tests Work with parsimony or maximum likelihood scores Records treelengths (steps or likelihoods) for each character Winning sites model sums the number of sites supporting tree A versus number of sites supporting tree B and vice versa (those having better fit, fewer steps, on alternative tree). Test against a binomial distribution: determine what fraction of winning sites significantly different from expectation of 0. 5 Wilcoxon signed ranks test Templeton (1983) replaces character differences with their rank and then uses Wilcoxon rank sum to test between two trees
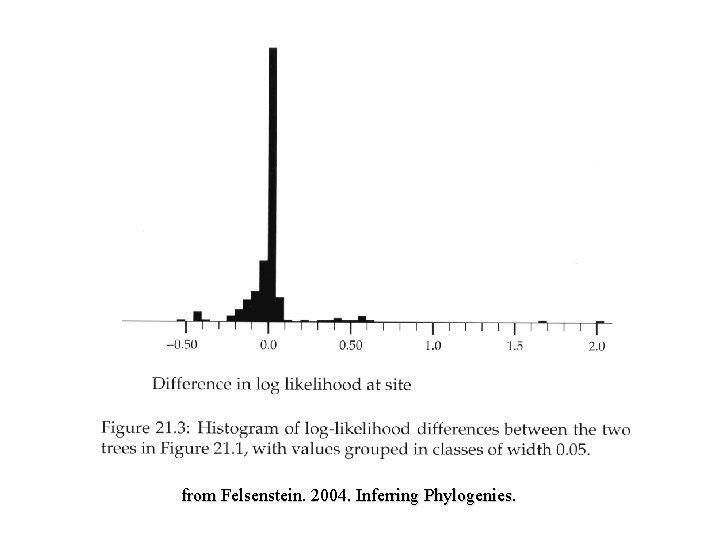
from Felsenstein. 2004. Inferring Phylogenies.

Kishino-Hasegawa Test (1989) * carry ML analysis for two (or more) trees * obtain difference (δ) in likelihood scores for each site (with expectation of zero if trees are not different) * calculate variance among sites in likelihood differences (σ2) * if δ /σ > 1. 96 reject the null hypothesis (that trees are equivalent)

from Felsenstein. 2004. Inferring Phylogenies.
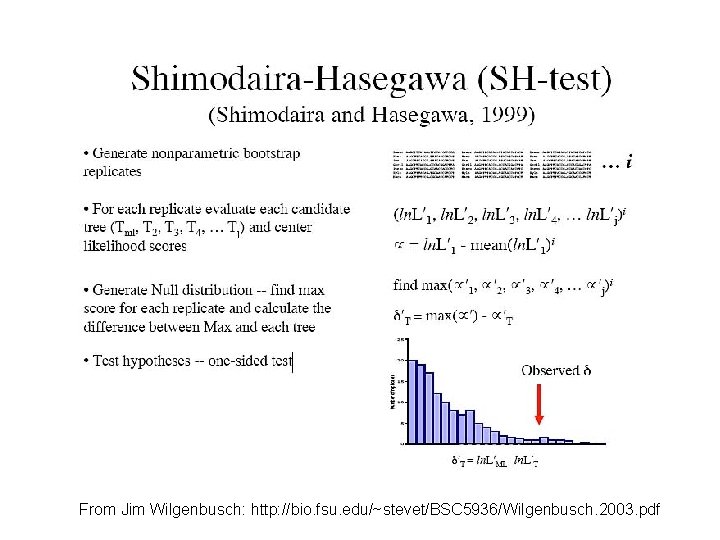
From Jim Wilgenbusch: http: //bio. fsu. edu/~stevet/BSC 5936/Wilgenbusch. 2003. pdf
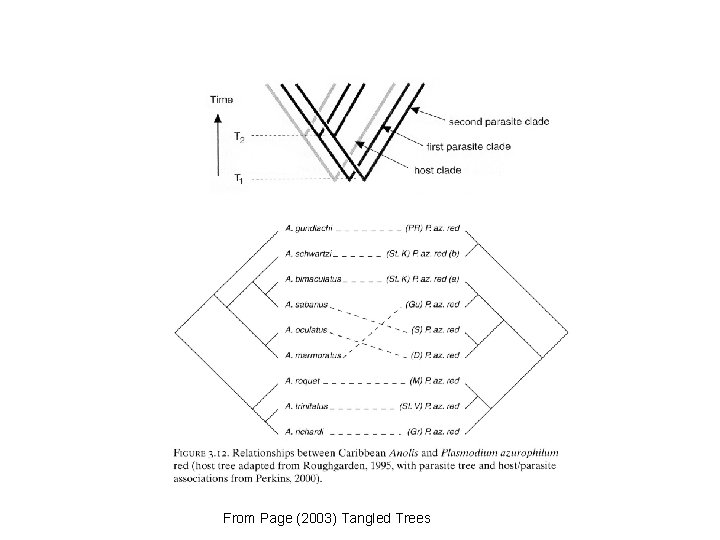
From Page (2003) Tangled Trees
- Slides: 12