Par SNP Hash Pipeline to parse SNP data














- Slides: 17

Par. SNP Hash Pipeline to parse SNP data and output summary statistics across sliding windows

Objective • Parse VCF files • Calculate summary statistics across sliding windows throughout the genome • Implement NTFreq module to calculate nucleotide frequencies for each population and combined population • Implement Tajimas. D module to calculate Tajima’s D • Implement GO module to annotate identified SNPs

Data set • Simulated data set for chromosome 2 R in Drosophila melanogaster • 1. 4 Mbp – 2 populations • Pooled individuals per population – 75 bp reads, error rate 1% – 10, 000 simulated SNPs • • • 100 x coverage per variant At least 100 bp apart Allelic Frequencies ranging from. 1 to. 9 per population

Fast. Q -> sai SAM -> BAM >. bcf -> VCF Data to->Variant Call -Format Index Reference Genome Only chromosome 2 R of D. melanogaster -Genome build Dmel 3 from Flybase Use BWA to Align Fast. Q to Reference Genome Gap open penalty = 1 Gap extension max = 12 Disallowing deletion within 12 bp of 3’UTR Maximum level of gap extensions = 12 Use SAMTools to Remove Ambiguously mapped Regions (MAPQ >= 20) Use BCFTools mpileup to Generate a Binary Code Format (BCF) BCF -> VCF

Formatting data: Parse VCF For each window: • Fetch the VCF rows from each BCF file • Convert the VCF rows into hashes of arrays • Compute the Theta, Pi, Tajima’s D for each population • Compute Fst for each window between each population

Sliding windows • Sliding window size is specified, and called modules are calculated across specified window size

Module 1: Calculate allele frequencies • Input is taken from parsed VCF file • Hashes are created for each population with the following structure – {SNP_location} {nucleotide} -> frequency; • Hashes created for full dataset – {SNP_location}{Population} -> {nucleotide} ->frequency

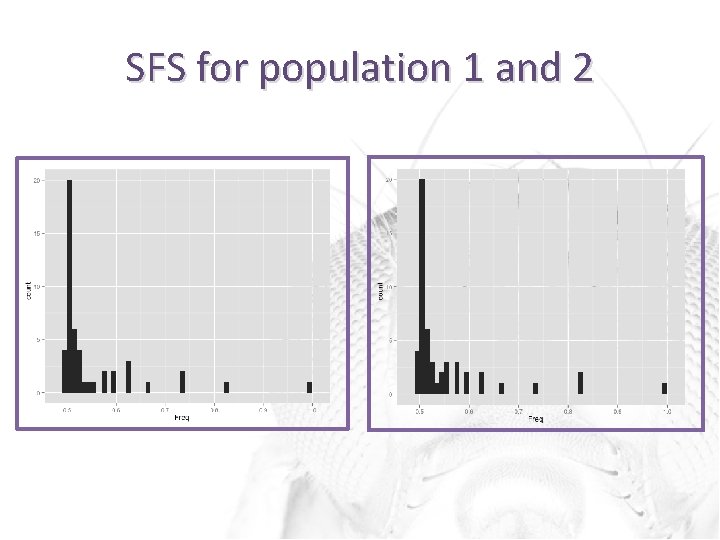
Output site frequency spectra • Site frequency spectrum (SFS) output as the following hash: – {nonref_allele}{frequency}->count; • Allows us to calculate a histogram for the nonreference allele frequencies • Send output to R to generate SFS graphs

Module 2: Calculate Summary Statistics and Tajima’s D • theta_pi (index of diversity) • theta_watterson (index of diversity)

Module 2: Calculate Summary Statistics and Tajima’s D • Tajima’s D (index of selection/population expansion)

Module 3: FST for DNA sequence • Calculate FST (index of differentiation) according to Hudson et al. 1992 1 – Hw/Hb Hw: average number of differences within each population Hb: average number of differences between the 2 populations

Module 4: GO annotations • Module takes SNP list as input • Outputs the following: – List of genes that have overlap with SNP positions – Gene Ontology (GO) IDs and terms associated with each SNP matched gene – List of genes for a selected window • Visualization using GOSlim

Data visualization • Integrated Genomics Viewer (IGV) • Broad Institute • http: //www. broadinstitute. org/igv/

SFS for population 1 and 2

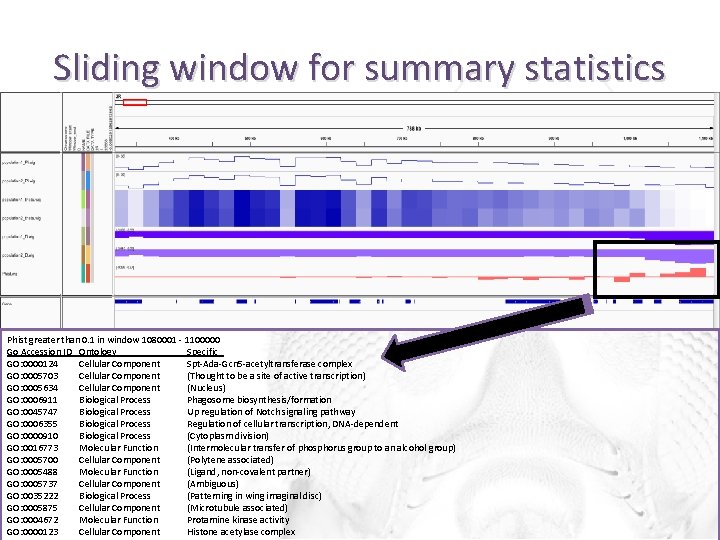
Sliding window for summary statistics Phist greater than 0. 1 in window 1080001 - 1100000 Go Accession ID Ontology Specific GO: 0000124 Cellular Component Spt-Ada-Gcn 5 -acetyltransferase complex GO: 0005703 Cellular Component (Thought to be a site of active transcription) GO: 0005634 Cellular Component (Nucleus) GO: 0006911 Biological Process Phagosome biosynthesis/formation GO: 0045747 Biological Process Up regulation of Notch signaling pathway GO: 0006355 Biological Process Regulation of cellular transcription, DNA-dependent GO: 0000910 Biological Process (Cytoplasm division) GO: 0016773 Molecular Function (Intermolecular transfer of phosphorus group to an alcohol group) GO: 0005700 Cellular Component (Polytene associated) GO: 0005488 Molecular Function (Ligand, non-covalent partner) GO: 0005737 Cellular Component (Ambiguous) GO: 0035222 Biological Process (Patterning in wing imaginal disc) GO: 0005875 Cellular Component (Microtubule associated) GO: 0004672 Molecular Function Protamine kinase activity GO: 0000123 Cellular Component Histone acetylase complex

Identify differentiated genomic regions • For each window with a Fst > 0. 1, print the name of the SNP and associated GO term Phist (Fst) greater than 0. 1 in window 1080001 - 1100000 Go Accession ID Ontology Specific GO: 0000124 Cellular Component Spt-Ada-Gcn 5 -acetyltransferase complex GO: 0005703 Cellular Component (Thought to be a site of active transcription) GO: 0005634 Cellular Component (Nucleus) GO: 0006911 Biological Process Phagosome biosynthesis/formation GO: 0045747 Biological Process Regulation of cellular transcription, DNA-dependent GO: 0000910 Biological Process (Cytoplasm division) GO: 0016773 Molecular Function (Intermolecular transfer of phosphorus group to an alcohol group)GO: 0005700 Cellular Component (Polytene associated) GO: 0005488 Molecular Function (Ligand, non-covalent partner) GO: 0005737 Cellular Component (Ambiguous) GO: 0035222 Biological Process (Patterning in wing imaginal disc) GO: 0005875 Cellular Component (Microtubule associated) GO: 0004672 Molecular Function Protamine kinase activity GO: 0000123 Cellular Component Histone acetylase complex

Thank You Use PERL or die , print “ (X_x)”; ##Hashes to Hashes## Print “ % 2 %”;